Home
Query Results
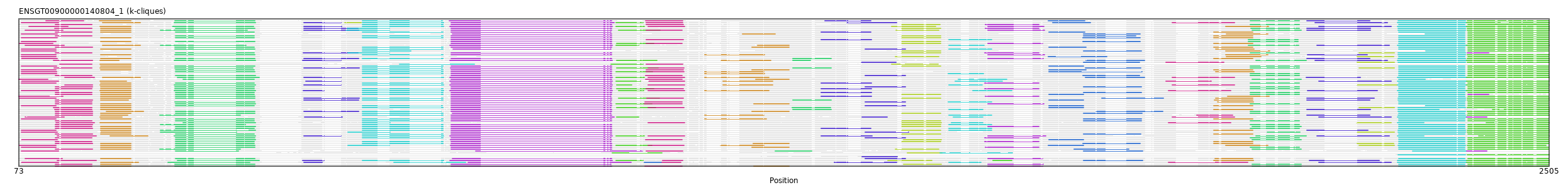
"ENSGT00900000140804_1"
Found:
ENSGT00900000140804_1
ENSSBOP00000011138 (view gene)
ENSPEMP00000017918 (view gene)
--------------MDRVK-FAASGPALRGWLLLATVTVGVLAQSVLAGVKKSDVPCGGR
DCS-GGCQCFPEKGARGQPGPVGTQGYTGPPGLQGFPGLQGRKGDKGERGAPGPTGPKGD
VGARGVSGFP
PF01391 // ENSGT00900000140804_1_Domain_0 (131, 189)
GADGIP---GHPGQGGPRGRPGYDGCNGTRGDAGPQGPSGPGGFPGLPGP
QGPKGQKGENSGT00900000140804_1_Linker_0_1 (189, 203)
EPYALSQEERDKYRPF01391 // ENSGT00900000140804_1_Domain_1 (203, 251)
GEPGEPGLVGYQGPPGRPGPIGQMGPVGAPGRPGPPGP
PGPKGQPGNRENSGT00900000140804_1_Linker_1_2 (251, 322)
GLGF--YGEKGEKGDVGQPGPNGIPSDT---TLFGPTTTTYHPDLYKGEK
GSRG---ETGIPGT----TLKPF01391 // ENSGT00900000140804_1_Domain_2 (322, 449)
GEEGIMGFPGARGLPGL---DGEKGIFGQ----------
---------------------------------------------------------KGS
RGLDGFQGPS-GPRGPKGERGEQGPPGPPAYSPHP-------------------------
-----------------------------------------ALAKGIRGDPGSPGAHGEP
GSRGEPGDPGPMGP---P--------------------------GPSVGDED-TKRGLPG
EMGPKGFSGEPGHPARYP
PF01391 // ENSGT00900000140804_1_Domain_5 (619, 747)
GSPGVDGKPGLQGATGPPGPPGPD------------------
-GFLFGLKGSEGRVGYPGPSGFPGTRGQKGWKG---------------------------
-----------------------EAGENSGT00900000140804_1_Linker_5_6 (747, 760)
DC-QCG---QFIGPF01391 // ENSGT00900000140804_1_Domain_6 (760, 1015)
GLPGLPGPKGFPGVNGELGKK
GDQGDPGVHGIPGFPGFKGAPGLTGI----------------------------------
------------------------------------------------------------
------------------------------------------------------------
-----------------------------------------PGP---KG--VKGDSRTVT
TKGERGQPGIPGVHGMKGDDGIPGRDGLDGFPGL-----PGP-PGDGIKGPPGDAGLPGL
PGTKGFPGEVGPPGQGLPGPKGERGFPGDAGFPGPPGLPGPPGPPGTPGQIDCD---TGV
KRPFGDGQQVAV--QPGCV--EGPT-----------------------GSPGQQGPP---
G----------------PTGIKGIRGTPGFPGGSGAPGLKGFPGDPGREGFPGPP-GFMG
PRGSKGATGLPGPSGLPGPIGLPGPAGPPGDRGIPGEVLGAQPGPRGDAGLPGHPGLKGP
PGERGPPGFRGS-----QGMPGMPGLKGQPGFPGPSGQPGLSGPPGQHGF-PGAPGREGP
LGLPGSP-GL--------------------GGLPGDRGEPGEPGVPGPVGMKGLSGDRGD
AGMSGERGHPGSPGFKGMAGMPGLPGQK-------
PF01391 // ENSGT00900000140804_1_Domain_13 (1476, 1538)
GDRGSPGMDGFQGMLGLKGRPGFPG
IKGEAGFFGV----PGLKGLPGEPGVKGNRGDMGPPGPPPIIQPG-MKDIKGEKGDEGPM
GLKGYLGVK-------------GVQGMPGVPGLSGIP-GLPGRPGYIKGV-K
PF01391 // ENSGT00900000140804_1_Domain_15 (1613, 1694)
GDIGVPGI
PGLPGQQGVIGPPGITGFPGFTGSRGEKGAPGR---------------------------
---AGFFGETGPTGDF---GDIGDTVNLPGSPGLKGERGVTGIP-GLKGFFGEKGAQGEV
GFPGITGIPGGQGSPGLKGQTGFPGLTGLQGPQGEPGRSGVPG--DKGDYGWPGVPGLP-
--------------------------------GFPGIRGISGLHGLPGTKGFPGSPGADV
HG----------D---PGFPGPAGDRGDRGEANTLPGPVGAAGQKGERGTP---------
--------------------------GERGPAGSPGLQGFPGI-SPPSNISGPPGDVGA-
---PGIFGLEGYRGPPGPPGSDALPGIKGDEGTSGAAGLPGEKGWVGDPGPQ
PF01391 // ENSGT00900000140804_1_Domain_20 (2033, 2111)
GQPGVVGL
PGEKG---------PRGEQGFMGNTGPS-----GNVGDRGPKGPKGDQGFPG-------A
PGSMGSPGIPGIPQKIATQPGTMGPQGRRGLPGAQGEMGPQGPPGDP-------------
---------------------------------------GFRGAPGKAGPQGRGGVSAVP
GFRGDQGPMGQQGPVGQE----GEPGRPGSPGLPGMPGRSVSIGY
PF01413 // ENSGT00900000140804_1_Domain_23 (2266, 2371)
LLVKHSQTDQEPMCP
VGMNKLWSGYSLLYFEGQEKAHNQDLGLAGSCLARFSTMPFLYCNPGDVCYYASRNDKSY
WLSTTAPLPMMPVAEEE--IKPYISRCSVCENSGT00900000140804_1_Linker_23_25 (2371, 2376)
EAPAVPF01413 // ENSGT00900000140804_1_Domain_25 (2376, 2504)
AIAVHSQ--DVTIPHCPTGWRSLWI
GYSFLMHTAAGDEGGGQS----LVSPGSCLEDFRAT---PFIECNG-GRGTCHYFANKYS
--FWLTTIPEQNFQGTPSA-----DTLKAGLIRTHISRCQVCMKNL--------------
--------------
ENSRBIP00000027815 (view gene)
--------------MGRDQ-RAVAGPALRRWLLLGTVTVGFLAQGVLAGVKKFDVPCGGR
DCS-GGCQCYPEKGGRGQPGPVGPQGYNGPPGLQGFPGLQGRKGDKGERGAPGITGPKGD
VGARGVSGFP
PF01391 // ENSGT00900000140804_1_Domain_0 (131, 189)
GADGIP---GHPGQGGPRGRPGYDGCNGTQGDSGPQGPPGSEGFTGPPGP
QGPKGQKGEPYALPREERDRYRGEPGEPGLVGFQGPPGRPGPVGQMGPVGAPGRPGPPGP
PGPKGQQGNRGLGF--YGVKGEKGDVGQPGPNGIPSDL-LHPIIAPTGVTFHPDQYKGEK
GS---EGEPGIRGI----SLK
PF01391 // ENSGT00900000140804_1_Domain_2 (322, 448)
GEEGIMGFPGLRGYPGL---SGEKGSSGQ----------
---------------------------------------------------------KGS
RGLDGYQGPD-GPRGPKGEAGDLGPPGLPAYSPHP-------------------------
-----------------------------------------SLAKGARGDPGFPGAQGEP
GSQGEPGDPGPRGP---P--------------------------GLSIGDED-QRRGLPG
EMGPKGFIGDPGIPALYP
PF01391 // ENSGT00900000140804_1_Domain_5 (619, 747)
GPPGPDGRPGPPGPPGLPGPPGPD------------------
-GFLFGLKGAEGRAGFPGLSGSPGARGPKGWKG---------------------------
-----------------------DAGENSGT00900000140804_1_Linker_5_6 (747, 760)
DC-RCAEGDEPVRPF01391 // ENSGT00900000140804_1_Domain_6 (760, 1014)
GPPGLPGPKGFAGINGEPGRK
GDKGDPGQHGLPGFPGLKGVPGNVGA----------------------------------
------------------------------------------------------------
------------------------------------------------------------
-----------------------------------------PGP---KG--AKGDSRTIT
TKGERGQPGVPGVPGMKGDDGGPGRDGLDGFPGL-----PGP-PGDGIKGPPGDAGYPGI
PGTKGTPGEVGPPGLGLPGFKGQRGFPGDAGLPGPPGFPGPPGPAGTPGQRDCD---TDV
KRPIGGDRQEAI--QPGCV--GGPK-----------------------GLPGLPGPP---
G----------------PSGAKGLRGIPGFSGADGGPGPKGLPGDPGREGFPGPP-GFIG
PRGSKGAVGLPGPDGPPGPIGLPGPDGPPGDRGLPGEVLGAQPGPRGDAGVPGQPGFKGL
PGDRGPPGFRGS-----QGMPGMPGPKGQPGLPGPSGQPGLSGPPGQHGF-PGAPGQEGP
LGLPGIP-GL--------------------EGLPGDR
PF01391 // ENSGT00900000140804_1_Domain_12 (1418, 1482)
GDPGDIGAPGPVGMKGLSGDRGD
AGLAGERGHPGSPGFKGIDGMPGTPGLK-------GERGSPGMDGFQGMPGLKGRPGFPG
SKGEAGFFGI----PGLKGVAGEPGFKGSRGDPGPPGPPPIILPG-MKDIKGEKGDEGPM
GLKGYLGAK-------------GIPGMPGIPGLSGIP-GLPGRPGHI
PF01391 // ENSGT00900000140804_1_Domain_15 (1608, 1694)
KGV-KGDIGAPGI
PGLPGFPGVAGPPGITGFPGFIGSRGDKGAPGR---------------------------
---AGLYGEIGPTGDF---GDIGDTINLPGRPGLKGEQGTAGIP-GLKGFFGEKGTEGDI
GFPGITGVTGVQGPPGLKGQTGFPGLTGPPGSQGQPGRIGLPG--GKGDDGWPGTPGLP-
--------------------------------GFPGLRGIRGLHGLPGTKGFPGSPGADI
HG----------D---PGFPGPPGDRGDPGDANTLPGPVGVPGQKGEQGAP---------
--------------------------GERGPPGSPGLQGFPGI-TPPSNISGVPGDKGA-
---PGIFGLKGYRGPPGPPGSAALPGSKGDTGNPGAPGTPGTKGWAGDS
PF01391 // ENSGT00900000140804_1_Domain_20 (2030, 2108)
GPQGRPGVFGL
PGEKG---------PRGEQGFMGNTGPA-----GAVGDRGPKGPKGDPGFPG-------A
PGIVGAPGIAGIPQKIAIPPGTVGPQGRRGPPGAQGEMGPQGPPGEP-------------
---------------------------------------GFRGAPGKAGPQGRGGVSAVP
GFRGDEGPIGHQGPIGQE----GAPGRPGSPGLPGMPGRSASIGY
PF01413 // ENSGT00900000140804_1_Domain_23 (2266, 2371)
LLVKHSQTDQEPMCP
VGMNKLWSGYSLLYFEGQEKAHNQDLGLAGSCLARFSTMPFLYCNPGDVCYYASRNDKSY
WLSTTAPLPMMPVAEDE--IKPYISRCSVCENSGT00900000140804_1_Linker_23_25 (2371, 2376)
EAPAVPF01413 // ENSGT00900000140804_1_Domain_25 (2376, 2504)
AIAVHSQ--DVSIPHCPAGWRSLWI
GYSFLMHTAAGDEGGGQS----LVSPGSCLEDFRAT---PFIECNG-GRGTCHYYANKYS
--FWLTTIPEQSFQGSPSA-----DTLKAGLIRTHISRCQVCMKNL--------------
--------------
ENSFCAP00000013714 (view gene)
--------------MDRHQ-CSVSSPALRRWLLLGAVTVGVLAQSVLAGVKKFDVPCGGR
DCS-GGCQCFPEKGGR
PF01391 // ENSGT00900000140804_1_Domain_0 (77, 133)
GQPGPVGPQGYTGPPGLQGFPGLQGRKGDKGERGSPGITGPKGD
VGARGVSGFPGADGIP---GHPGQGGPRGPPGHDGCNGTRGDVGRQGPVGPGGFAGPPGP
QGPKGQKGEPYALSSVDRDKYR
PF01391 // ENSGT00900000140804_1_Domain_1 (203, 251)
GEPGEPGLVGFQGPPGRHGPVGQMGPVGAPGRPGPPGP
PGPKGQPGNRENSGT00900000140804_1_Linker_1_2 (251, 321)
GLGF--YGQKGEKGDMGQPGPNGIPSDI-HHAIIGPMKEMFHPDQYKGEK
GS---EGEPGRKGI----ALPF01391 // ENSGT00900000140804_1_Domain_2 (321, 448)
KGEEGVMGFSGIRGSPGF---DGEKGSPGQ----------
---------------------------------------------------------KGS
KGLDGYQGPD-GPPGLKGEVGDRGPPGAPAYSPHP-------------------------
-----------------------------------------SLAKGVRGDPGFPGAHGEP
GSRGEPGDPGPPGP---P--------------------------GTSIGDED-GKRGLPG
EMGPKGFIGDPGIPALYP
PF01391 // ENSGT00900000140804_1_Domain_5 (619, 747)
GPPGTDGKPGPRGPPGPPGLPGPD------------------
-GFLFGLKGAEGNVGYPGPSGFPGARGQKGWKG---------------------------
-----------------------DAGENSGT00900000140804_1_Linker_5_6 (747, 760)
DC-KCAEGDQFIGPF01391 // ENSGT00900000140804_1_Domain_6 (760, 1015)
GLPGPPGPKGLPGVNGEPGKK
GSQGDPGQHGIPGFPGFKGAPGNVGP----------------------------------
------------------------------------------------------------
------------------------------------------------------------
-----------------------------------------PGP---KG--VKGDSRAIT
TKGERGQPGVPGVPGLKGDDGVPGRDGLDGFPGL-----PGP-PGDGITGPPGDIGYPGT
PGTKGAPGERGPPGLGLPGPKGERGFPGDVGPPGPPGFPGPAGPPGAPGQIDCD---TGV
KRPIGDDGQKTV--RPDCV--GGPK-----------------------GAPGLRGPP---
G----------------PTGAKGLPGIPGPSGADGVPGLKGLPGDPGREGFPGPP-GFVG
PRGSKGVVGLRGPDGHPGPTGLPGPVGPPGDRGLPGEVLGAQPGPRGDAGLPGHPGLKGP
PGERGPPGFRGS-----QGMPGMPGLKGQPGFPGPSGQPGLPGPPGLHGF-PGAPGQEGP
LGLPGAP-GR--------------------GGLPGDRGDPGDIGIPGPVGMKGLSGDRGD
PGLLGERGPPGGPGFKGMSGIPGVPGPK-------
PF01391 // ENSGT00900000140804_1_Domain_13 (1476, 1538)
GSRGSPGMDGFQGMVGLKGRAGLPG
NKGEAGFFGI----PGLKGLAGEPGVKGSRGDPGPPGPPPIIQPG-MKDIKGEKGDEGPM
GLKGYLGLK-------------GVPGMPGIPGLSGIP-GLPGKPGHIKGV-KGDIGVPGV
PGLPGFPGVPGPPGIIGFPGFTGSRGDKGAPGR---------------------------
---AGLYGEVGPTGDF---GDIGDTIDLP
PF01391 // ENSGT00900000140804_1_Domain_17 (1710, 1765)
GSPGLKGERGIAGAP-GLKGFFGEKGTEGEV
GFPGITGVPGVQGPPGPKGQAGFPGLTGLQGPQGEPGRAGAPG--DKGDYGWPGIPGSP-
--------------------------------GFPGIRGISGLHGLPGTKGFPGSPGVDA
HG----------D---PGFPGPAGDRGDPGEANTRPGPVGAPGQKGERGNP---------
--------------------------GERGPVGSPGLQGFPGI-TPPSNISGSPGDKGA-
---PGIFGPEGYRGPPGPPGPAALPGRKGDEGNPGAPGNPGTKGWGGDP
PF01391 // ENSGT00900000140804_1_Domain_20 (2030, 2108)
GPQGRPGVFGL
PGEKG---------PRGEPGLMGNTGAT-----GSVGDPGPKGPKGDRGLPG-------A
PGSVGSPGIVGIPQKITVQPGPVGPHGRRGPPGAQGEMGPQGPPGEP-------------
---------------------------------------GFRGAPGKAGPQGRGGVSAVP
GFRGDQGPMGQQGPIGQE----GEPGRPGTPGLPGMPGRSVSIGY
PF01413 // ENSGT00900000140804_1_Domain_23 (2266, 2371)
LLVKHSQTDQEPMCP
VGMNKLWSGYSLLYFEGQEKAHNQDLGLAGSCLARFSTMPFLYCNPGDVCYYASRNDKSY
WLSTTAPLPMMPVAEDE--IKPYISRCSVCENSGT00900000140804_1_Linker_23_25 (2371, 2376)
EAPAVPF01413 // ENSGT00900000140804_1_Domain_25 (2376, 2504)
AIAVHSQ--DVSIPHCPAGWRSLWI
GYSFLMHTAAGDEGGGQS----LVSPGSCLEDFRAT---PFIECNG-GRGTCHYYANKYS
--FWLTTIPERSFQGSPSA-----DTLKAGLIRTHISRCQVCMKNL--------------
--------------
ENSCJAP00000026472 (view gene)
--------------MGRNQ-RAAAGPALPRWLLLGTVTVGFLSQSVLAGVKKSDVPCGGR
DCS-GGCQCFPEKGGRGQPGPVGPQGYTGPPGLQGFPGLQGRKGDKGERGAPGITGPKGD
VGARGVSGFP
PF01391 // ENSGT00900000140804_1_Domain_0 (131, 190)
GADGIP---GHPGQGGPRGRPGYDGCNGTQGDSGPQGPPGSEGFTGPPGP
QGPKGQKGEPYALPKEERDRYRGEPGEPGLVGFQGPPGRPGPVGQMGPVGAPGRPGPPGP
PGPKGQQGNRGLGF--YGVKGEKGDVGQPGPNGIPPE--THPVIAPTRVTHHPEQDKGEK
GS---EGEPGIKGI----SL
PF01391 // ENSGT00900000140804_1_Domain_2 (321, 447)
KGEEGIMGFPGPRGYPGL---SGEKGSPGQ----------
---------------------------------------------------------KGS
RGLDGYQGSD-GPRGPKGETGDPGPPGLPVYSPHP-------------------------
-----------------------------------------SLAKGARGDPGFPGARGEP
GSQGEPGDPGPRGP---P--------------------------GISITAED-LGRGLPG
EMGPKGFIGGPGIPALYP
PF01391 // ENSGT00900000140804_1_Domain_5 (619, 747)
GPPGLDGKPGLPGPPGLPGPPGPD------------------
-GFLFGLKGAEGRGGFPGLSGSPGARGQKGRKG---------------------------
-----------------------DAGENSGT00900000140804_1_Linker_5_6 (747, 760)
DC-RCVEGDEAVRPF01391 // ENSGT00900000140804_1_Domain_6 (760, 1015)
GLPGLPGPKGFPGINGEPGRK
GDRGDPGQHGLPGFSGLKGVPGNLGA----------------------------------
------------------------------------------------------------
------------------------------------------------------------
-----------------------------------------PGP---KG--AKGDSRTIT
TKGERGQPGVPGVPGMKGDDGIPGREGLDGFPGL-----PGP-PGDGIKGPPGDPGYPGI
PGTKGSPGEMGPPGLGLPGFKGQRGFPGDAGLPGPRGFPGPPGPPGTPGQADCD---PDV
RRPIAGDRQEAV--QPGCA--GGPK-----------------------GLPGLPGPP---
G----------------PTGAKGLRGTPGFSGADGGA
PF01391 // ENSGT00900000140804_1_Domain_10 (1238, 1297)
GPKGLPGDPGREGFPGPP-GFIG
PRGSKGAAGLPGPDGLPGPVGLPGPVGPPGDKGLPGEVLGAQPGPRGDAGVPGHPGLKGL
PGDRGTPGFRGS-----QGMPGMPGLKGQLGFPGPSGQPGLSGPPGQHGF-PGAPGPEGP
LGLPGTL-GL--------------------GGLPGDRGEPGDVGAPGPVGMKGVSGDTGD
AGLAGERGRPGSPGFKGIDGMPGAPGLK-------GERGSPGMDGFQGMPGLKGRPGIPG
SKGEAGFFGI----PGLKGLAGEPGFKGSRGDPGPPGPPPIILPG-MKDIKGEKGDEGPM
GLKGYLGAK-------------GTQGMPGIPGLSGIP-GLPGRPGHIKGL-KGDIGAPGI
PGLPGFPGVAGPPGITGFPGFTGSRGDKGAPGR---------------------------
---AGLYGEAGQTGDF---GDIGDTINLP
PF01391 // ENSGT00900000140804_1_Domain_17 (1710, 1769)
GRPGLKGERGTAGIP-GLKGFFGEKGTEGDI
GFPGITGVTGVQGPPGLKGQAGFPGLTGPPGPQGEPGRIGLPG--GKGDDGWPGAPGLP-
--------------------------------GFPGPRGISGLHGLPGTKGFPGSPGADI
HG----------D---PGYSGPPGERGDPGEANTLPGPVGVPGQKGEQGAP---------
--------------------------GQRGPPGSPGLQGFPGI-TPPSNIS
PF01391 // ENSGT00900000140804_1_Domain_19 (1972, 2034)
GAPGDKGA-
---PGIFGLKGYRGPPGPPGSAALPGSKGDTGNPGAPGTPGTKGWVGDPGPQGRPGVLGL
PGEKG---------PRGEQGFMGNPGLP-----GPVGDRGPKGPKGDPGFPG-------A
PGIVGAPGIAGIPQKIATQPGTVGPQGRRGPPGAPGEMGPQGPPGEP-------------
---------------------------------------GFRGAPGKAGPQGRGGVSAVP
GFRGDEGPTGHQGPIGQE----GVPGRPGSPGLPGMPGRSVSIGY
PF01413 // ENSGT00900000140804_1_Domain_23 (2266, 2371)
LLVKHSQTDQEPMCP
VGMNKLWSGYSLLYFEGQEKAHNQDLGLAGSCLARFSTMPFLYCNPGDVCYYASRNDKSY
WLSTTAPLPMMPVAEDE--IKPYISRCSVCENSGT00900000140804_1_Linker_23_25 (2371, 2376)
EAPAVPF01413 // ENSGT00900000140804_1_Domain_25 (2376, 2504)
AIAVHSQ--DVSIPHCPAGWRSLWI
GYSFLMHTAAGDEGGGQS----LVSPGSCLEDFRAT---PFIECNG-GRGTCHYYANKYS
--FWLTTIPEQSFQGSPSA-----DTLKAGLIRTHISRCQVCMKNL--------------
--------------
ENSPPAP00000016802 (view gene)
--------------MGRDQ-RAVAGPALRRWL-LGTVTVGFLAQSVLAGVKKFDVPCGGR
DCS-GGCQCYPEKGGRGQPGPVGPQGYNGPPGLQGFPGLQGRKGDKGERGAPGITGPKGD
VGARGVSGFP
PF01391 // ENSGT00900000140804_1_Domain_0 (131, 190)
GADGIP---GHPGQGGPRGRPGYDGCNGTQGDSGPQGPPGSEGFTGPPGP
QGPKGQKGEENSGT00900000140804_1_Linker_0_1 (190, 203)
PYALPKEERDRYRPF01391 // ENSGT00900000140804_1_Domain_1 (203, 251)
GEPGEPGLVGFQGPPGRPGHVGQMGPVGAPGRPGPPGP
PGPKGQQGNRENSGT00900000140804_1_Linker_1_2 (251, 322)
GLGF--YGVKGEKGDVGQPGPNGIPSDL-LHPIIAPTGVTFHPDQYKGEK
GS---EGEPGIRGI----SLKPF01391 // ENSGT00900000140804_1_Domain_2 (322, 447)
GEEGIMGFPGLRGYPGL---SGEKGSPGQ----------
---------------------------------------------------------KGS
RGLDGYQGPD-GPRGPKGEAGDPGPPGLPAYSPHP-------------------------
-----------------------------------------SLAKGARGDPGFPGAQGEP
GSQGEPGDPGLPGA---P--------------------------GLSIGDGD-QRRGLPG
EMGPKGFIGDPGIPALYG
PF01391 // ENSGT00900000140804_1_Domain_5 (619, 747)
GPPGPDGKRGPPGPPGLPGPPGPD------------------
-GFLFGLKGAKGRAGFPGLPGSPGARGPKGWKG---------------------------
-----------------------DAGENSGT00900000140804_1_Linker_5_6 (747, 760)
EC-RCTEGDEAIKPF01391 // ENSGT00900000140804_1_Domain_6 (760, 1015)
GLPGLPGPKGFAGINGEPGRK
GDKGDPGQHGLPGFPGLKGVPGNVGA----------------------------------
------------------------------------------------------------
------------------------------------------------------------
-----------------------------------------PGP---KG--AKGDSRTIT
TKGERGQPGVPGVPGMKGDDGSPGRDGLDGFPGL-----PGP-PGDGIKGPPGDPGYPGI
PGTKGTPGEMGPPGLGLPGLKGQRGFPGDAGLPGPPGFLGPPGPAGTPGQIDCD---TDV
KRAVGGDRQEAI--QPGCV--GGLK-----------------------GLPGLPGPP---
G----------------PTGAKGLRGIPGFSGADGGPGPKGLPGDAGREGFPGPP-GFIG
PRGSKGAVGLPGPDGSPGPIGLPGPDGPPGERGLPGEVLGAQPGPRGDAGVPGQPGLKGL
PGDRGPPGFRGS-----QGMPGMPGLK
PF01391 // ENSGT00900000140804_1_Domain_12 (1348, 1427)
GQPGLPGPSGQPGLYGPPGLHGF-PGAPGQEGP
LGLPGIP-GR--------------------EGLPGDRGDPGDTGAPGPVGMKGLSGDRGD
AGFTGERGHPGSPGFKGIDGMPGTPGLK-------GDRGSPGMDGFQGMPGLKGRPGFPG
SKGEAGFFGI----PGLKGLAGEPGFKGSRGDPGPPGPPPVILPG-MKDI
PF01391 // ENSGT00900000140804_1_Domain_14 (1551, 1619)
KGEKGDEGPM
GLKGYLGAK-------------GIQGMPGIPGLSGIP-GLPGRPGHIKGV-KGDIGAPGI
PGLPGFPGVAGPPGITGFPGFIGSRGDKGAPGR---------------------------
---AGLYGEIGATGDF---GDIGDTINLPGRPGLKGERGTTGIP-GLKGFFGEKGTEGDI
GFPGITGVTGVQGPPGLKGQTGFPGLTGPPGSQGEPGRIGLPG--GKGDDGWPGAPGLP-
--------------------------------GFPGLRGIRGLHGLPGTKGFPGSPGSDI
HG----------D---PGFPGPPGERGDPGEANTLPGPVGVPGQKGDQGAP---------
--------------------------GERGPPGSPGLQGFPGI-TPPSNISGAPGDKGA-
---PGIFGLKGYRGPPGPPGSAALPGSKGDTGNPGAPGTPGTKGWAGDS
PF01391 // ENSGT00900000140804_1_Domain_20 (2030, 2108)
GPQGRPGVFGL
PGEKG---------PRGEQGFMGNTGPT-----GAVGDRGPKGPKGDPGFPG-------A
PGTVGAPGIAGIPQKIAVQPGTVGPQGRRGPPGAPGEMGPQGPPGEP-------------
---------------------------------------GFRGAPGKAGPQGRGGVSAVP
GFRGDEGPIGHQGPIGQE----GAPGRPGSPGLPGMPGRSVSIGY
PF01413 // ENSGT00900000140804_1_Domain_23 (2266, 2371)
LLVKHSQTDQEPMCP
VGMNKLWSGYSLLYFEGQEKAHNQDLGLAGSCLARFSTMPFLYCNPGDVCYYASRNDKSY
WLSTTAPLPMMPVAEDE--IKPYISRCSVCENSGT00900000140804_1_Linker_23_25 (2371, 2376)
EAPAVPF01413 // ENSGT00900000140804_1_Domain_25 (2376, 2504)
AIAVHSQ--DVSIPHCPAGWRSLWI
GYSFLMHTAAGDEGGGQS----LVSPGSCLEDFRAT---PFIECNG-GRGTCHYYANKYS
--FWLTTIPEQSFQGSPSA-----DTLKAGLIRTHISRCQVCMKNL--------------
--------------
ENSMODP00000005712 (view gene)
--------------MGSRM-YPTARLALRRSLLLF-VLAGVLTKRAHAAAKKYDVPCGGR
DCS-GGCQCFPEK
PF01391 // ENSGT00900000140804_1_Domain_0 (74, 132)
GARGQPGSVGPQGYNGPPGLQGIPGLQGRKGDKGERGAPGITGPKGD
VGQRGVSGFPGADGIP---GHPGQGGPRGRPGSDGCNGTTGDAGTQGPPGPGGLPGLPGP
QGPKGQKGEPYAFSQADRDRYR
PF01391 // ENSGT00900000140804_1_Domain_1 (203, 251)
GDQGEPGFIGLQGPPGHPGLPGKMGPAGAPGRPGPPGP
PGPKGQPGNRENSGT00900000140804_1_Linker_1_2 (251, 322)
GLGF--YGEKGEKGDIGRAGPTGLPSDG-DRATSITDNVVDQPDQYKGEK
GN---WGEPGWKGY----SEKPF01391 // ENSGT00900000140804_1_Domain_2 (322, 448)
GEEGIVGFPGQRGWPGL---NGEPGEPGQ----------
---------------------------------------------------------RGP
SGPSGFPGSD-GIKGPRGDIGDPGPPGENSGT00900000140804_1_Linker_2_3 (448, 526)
QPSYSHHP-------------------------
-----------------------------------------SIMKPF01391 // ENSGT00900000140804_1_Domain_3 (526, 614)
GVKGEPGTPGVRGPD
GRPGEPGEAGQPG--------------------------------TSGIDGD-ERRGLPG
EMGAKGFRGEPGFENSGT00900000140804_1_Linker_3_5 (614, 619)
PALYPPF01391 // ENSGT00900000140804_1_Domain_5 (619, 747)
GPPGRDGIPGPQGPQGPPGLPGPD------------------
-GFLFGLKGAEGSVGSPGPSGFPGARGQKGWKG---------------------------
-----------------------DPGENSGT00900000140804_1_Linker_5_6 (747, 760)
DC-KCEVS---SVPF01391 // ENSGT00900000140804_1_Domain_6 (760, 1015)
GLPGPPGPIGLPGTNGEPGRK
GEIGDPGPHGIPGFSGSKGSTGAPGA----------------------------------
------------------------------------------------------------
------------------------------------------------------------
-----------------------------------------AGL---KG--QKGENSGT00900000140804_1_Linker_6_7 (1015, 1026)
DSRTIT
TKGLRPF01391 // ENSGT00900000140804_1_Domain_7 (1026, 1091)
GDQGIPGIPGLEGEDGFPGQDGLDGFPGL-----PGL-PGDGIKGIPGDAGFPGL
PGLKGTPGELGPPGLGLPGPKGARGLPGDPGLDGSPGFPGPPGPPGTLGQTDCD---EGVKRPIGQEI---I--QPGCI--RAPK-----------------------GSPGLPGLP---G----------------SPGTKGLRGIPGFPGADGFVGLKGFQGDAGREGFPGPP-GFIGPRGSKGAIGLPGLDGNPGSTGVPGLVGPTGQKGLPGEVLGAQTGFPGDSGLPGVPGLKGEPGSRGPPGFRGS-----PGMPGMPGLKGQPGFPGPSGPPGLPGPPGTHGF-PGPKGQEGLPGLPGPS-GF--------------------AGLPGDRGDRGDIGVPGPVGMKGLDGDRGDTGFIGEQGRTGEPGFQGIPGKPGLPGPK-------GFKGSPGMDGFKGMLGIKGRSGIIGYKGEVGNFGI----PGFKGLPGEPGLKGIRGEPGPPGDPPKFLPG-MKDFKGEKGDEGPTGGKGFWGTK-------------GGQGMPGMPGQTGIP-GLPGIPGYIKGV-KGEIGIPGVPGLQGFPGQIGSPGIRGFPGFVGVRGDKGMPGR------------------------------EGLYGETGTIGDY---GDEGDTINLPGRPGVKGELGISGLG-GEKGFTGEKGLSGDHGFIGLMGLTGSQGLPGLKGQTGFPGLTGLQGPQGDPGRAGPPG--EKGDLGWPGIPGLS---------------------------------GFPGSRGISGLHGLPGTKGLPGSPGAEGYG----------N---PGFPGPFGDKGERGEPNTIPGPLGAPGQKGERGAQ-----------------------------------GGQGPSGSPGIQGHPGF-TAPSNISGLPGDKGA----PGLFGLQGYRGLPGTPGPSAPPGIKGEVGSPGLPGERGPKGWVGDPGTQGRPGVYGLPGEKG---------PRGQQGLMGYHGIP-----GSSGDRGPKGPKGDRGFPG------TP-GTMGSPGIPGIPLKIVTEPGVAGPHGRRGPPGIQGEMGPQGPPGDP----------------------------------------------------GLRGLPGKPGPQGRGGMSALPGFRGDQGLMGHQGPVGQE----GEPGRPGSPGLPGMPGRSVSIGYLLVKHSQTDQEPMCPVGMNKLWSGYSLLYFEGQEKAHNQDLGLAGSCLARFSTMPFLYCNPGDVCHYASRNDKSYWLSTTAPLPMMPVAEEE--IKPYISRCSVCEAPAVAIAVHSQ--DVSIPHCPAGWRSLWIGYSFLMHTAAGDEGGGQS----LVSPGSCLEDFRAT---PFIECNG-GRGTCHYFANKYS--FWLTTIPEQSFQGSPSA-----DTLKAGLIRTHISRCQVCMKNL--------
PF01391 // ENSGT00900000140804_1_Domain_9 (1071, 1129)
IPGDAGFPGLPGLKGTPGELGPPGLGLPGPKGARGLPGDPGLDGSPGFPGPPGPPGTLGQTDCD---EGV
KRPIGQEI---I--QPGCI--RAPK-----------------------GSPGLPGLP---
G----------------SPGTKGLRGIPGFPGADGFVGLKGFQGDAGREGFPGPP-GFIG
PRGSKGAIGLPGLDGNPGSTGVPGLVGPTGQKGLPGEVLGAQTGFPGDSGLPGVPGLKGE
PGSRGPPGFRGS-----PGMPGMPGLKGQPGFPGPSGPPGLPGPPGTHGF-PGPKGQEGL
PGLPGPS-GF--------------------AGLPGDRGDRGDIGVPGPVGMKGLDGDRGD
TGFIGEQGRTGEPGFQGIPGKPGLPGPK-------
PF01391 // ENSGT00900000140804_1_Domain_13 (1476, 1538)
GFKGSPGMDGFKGMLGIKGRSGIIG
YKGEVGNFGI----PGFKGLPGEPGLKGIRGEPGPPGDPPKFLPG-MKDFKGEKGDEGPT
GGKGFWGTK-------------GGQGMPGMPGQTGIP-GLPGIPGYIKGV-KGEIGIPGV
PGLQGFPGQIGSPGIRGFPGFVGVRGDKGMPGR---------------------------
---EGLYGETGTIGDY---GDEGDTINLPGRPGVKGELGISGLG-GEKGFTGEKGLSGDH
GFIGLMGLTGSQGLPGLKGQTGFPGLTGLQGPQGDPGRAGPPG--EKGDLGWP
PF01391 // ENSGT00900000140804_1_Domain_17 (1794, 1892)
GIPGLS-
--------------------------------GFPGSRGISGLHGLPGTKGLPGSPGAEG
YG----------N---PGFPGPFGDKGERGEPNTIPGPLGAPGQKGERGAQ---------
--------------------------GGQGPSGSPGIQGHPGF-TAPSNISGLP
PF01391 // ENSGT00900000140804_1_Domain_19 (1975, 2034)
GDKGA-
---PGLFGLQGYRGLPGTPGPSAPPGIKGEVGSPGLPGERGPKGWVGDPGTQGRPGVYGL
PGEKG---------PRGQQGLMGYHGIP-----GSSGDRGPKGPKGDRGFPG------TP
-GTMGSPGIPGIPLKIVTE
PF01391 // ENSGT00900000140804_1_Domain_21 (2120, 2227)
PGVAGPHGRRGPPGIQGEMGPQGPPGDP-------------
---------------------------------------GLRGLPGKPGPQGRGGMSALP
GFRGDQENSGT00900000140804_1_Linker_21_23 (2227, 2265)
GLMGHQGPVGQE----GEPGRPGSPGLPGMPGRSVSIGPF01413 // ENSGT00900000140804_1_Domain_23 (2265, 2372)
YLLVKHSQTDQEPMCP
VGMNKLWSGYSLLYFEGQEKAHNQDLGLAGSCLARFSTMPFLYCNPGDVCHYASRNDKSY
WLSTTAPLPMMPVAEEE--IKPYISRCSVCEENSGT00900000140804_1_Linker_23_25 (2372, 2375)
APAPF01413 // ENSGT00900000140804_1_Domain_25 (2375, 2505)
VAIAVHSQ--DVSIPHCPAGWRSLWI
GYSFLMHTAAGDEGGGQS----LVSPGSCLEDFRAT---PFIECNG-GRGTCHYFANKYS
--FWLTTIPEQSFQGSPSA-----DTLKAGLIRTHISRCQVCMKNL--------------
--------------
ENSGGOP00000012281 (view gene)
ENSDNOP00000013857 (view gene)
ENSRROP00000029286 (view gene)
--------------MGRDQ-RAVAGPALRRWLLLGTVTVGFLAQGVLAGVKKFDVPCGGR
DCS-GGCQCYPEKGGRGQPGPVGPQGYNGPPGLQGFPGLQGRKGDKGERGAPGITGPKGD
VGARGVSGFP
PF01391 // ENSGT00900000140804_1_Domain_0 (131, 189)
GADGIP---GHPGQGGPRGRPGYDGCNGTQGDSGPQGPPGSEGFTGPPGP
QGPKGQKGEPYALPREERDRYRGEPGEPGLVGFQGPPGRPGPVGQMGPVGAPGRPGPPGP
PGPKGQQGNRGLGF--YGVKGEKGDVGQPGPNGIPSDL-LHPIIAPTGVTFHPDQYKGEK
GS---EGEPGIRGI----SLK
PF01391 // ENSGT00900000140804_1_Domain_2 (322, 447)
GEEGIMGFPGLRGYPGL---SGAKGSPGQ----------
---------------------------------------------------------KGS
RGLDGYQGPD-GPRGPKGEAGDLGPPGLPAYSPHP-------------------------
-----------------------------------------SLAKGARGDPGFPGAQGEP
GSQGEPGDPGPRGP---P--------------------------GLSIGDED-QRRGLPG
EMGPKGFIGDPGIPALYP
PF01391 // ENSGT00900000140804_1_Domain_5 (619, 747)
GPPGPDGRPGPPGPPGLPGPPGPD------------------
-GFLFGLKGAEGRAGFPGLSGSPGARGPKGWKG---------------------------
-----------------------DAGENSGT00900000140804_1_Linker_5_6 (747, 760)
DC-RCAEGDEPVRPF01391 // ENSGT00900000140804_1_Domain_6 (760, 1014)
GPPGLPGPKGFAGINGEPGRK
GDKGDPGQHGLPGFPGLKGVPGNVGA----------------------------------
------------------------------------------------------------
------------------------------------------------------------
-----------------------------------------PGP---KG--AKGDSRTIT
TKGERGQPGVPGVPGMKGDDGGPGRDGLDGFPGL-----PGP-PGDGIKGPPGDAGYPGI
PGTKGTPGEVGPPGLGLPGFKGQRGFPGDAGLPGPPGFPGPPGPAGTPGQRDCD---TDV
KRPIGGDRQEAI--QPGCV--GGPK-----------------------GLPGLPGPP---
G----------------PSGAKGLRGIPGFSGADGGPGPKGLPGDPGREGFPGPP-GFIG
PRGSKGAVGLPGPDGPPGPIGLPGPDGPPGDRGLPGEVLGAQPGPRGDAGVPGQPGFKGL
PGDRGPPGFRGS-----QGMPGMPGPKGQPGLPGPSGQPGLSGPPGQHGF-PGAPGQEGP
LGLPGIP-GL--------------------EGLPGDRGDPGDIGAPGPVGMKGLSGDRGD
AGLAGERGHPGSPGFKGIDGMPGTPGLK-------
PF01391 // ENSGT00900000140804_1_Domain_13 (1476, 1538)
GERGSPGMDGFQGMPGLKGRPGFPG
SKGEAGFFGI----PGLKGVAGEPGFKGSRGDPGPPGPPPIILPG-MKDIKGEKGDEGPM
GLKGYLGAK-------------GIPGMPGIPGLSGIP-GLPGRPGHI
PF01391 // ENSGT00900000140804_1_Domain_15 (1608, 1694)
KGV-KGDIGAPGI
PGLPGFPGVAGPPGITGFPGFIGSRGDKGAPGR---------------------------
---AGLYGEIGPTGDF---GDIGDTINLPGRPGLKGEQGTAGIP-GLKGFFGEKGTEGDI
GFPGITGVTGVQGPPGLKGQTGFPGLTGPPGSQGQPGRIGLPG--GKGDDGWPGTPGLP-
--------------------------------GFPGLRGIRGLHGLPGTKGFPGSPGADI
HG----------D---PGFPGPPGDRGDPGDANTLPGPVGVPGQKGEQGAP---------
--------------------------GERGPPGSPGLQGFPGI-TPPSNIS
PF01391 // ENSGT00900000140804_1_Domain_19 (1972, 2034)
GVPGDKGA-
---PGIFGLKGYRGPPGPPGSAALPGSKGDTGNPGAPGTPGTKGWAGDSGPQGRPGVFGL
PGEKG---------PRGEQGFMGNTGPA-----GAVGDRGPKGPKGDPGFPG-------A
PGIVGAPGIAGIPQKIAIPPGTVGPQGRRGPPGAQGEMGPQGPPGEP-------------
---------------------------------------GFRGAPGKVGPQGRGGVSAVP
GFRGDEGPIGHQGPIGQE----GAPGRPGSPGLPGMPGRSASIGY
PF01413 // ENSGT00900000140804_1_Domain_23 (2266, 2371)
LLVKHSQTDQEPMCP
VGMNKLWSGYSLLYFEGQEKAHNQDLGLAGSCLARFSTMPFLYCNPGDVCYYASRNDKSY
WLSTTAPLPMMPVAEDE--IKPYISRCSVCENSGT00900000140804_1_Linker_23_25 (2371, 2376)
EAPAVPF01413 // ENSGT00900000140804_1_Domain_25 (2376, 2504)
AIAVHSQ--DVSIPHCPAGWRSLWI
GYSFLMHTAAGDEGGGQS----LVSPGSCLEDFRAT---PFIECNG-GRGTCHYYANKYS
--FWLTTIPEQSFQGSPSA-----DTLKAGLIRTHISRCQVCMKNL--------------
--------------
ENSNGAP00000013591 (view gene)
--------------MASER-CAASGPALRGWLLLGAVTVGFLAQSVSAGVKKFDVPCGGR
DCS-GGCQCYPEKGGRGQPGPVGPQGFSGPPGLQGFPGLQGRKGDKGERGAPGPTGPKGD
VGARGVSGFP
PF01391 // ENSGT00900000140804_1_Domain_0 (131, 189)
GADGIP---GHPGQGGPRGRPGYDGCNGTRGDAGPQGPSGSGGFPGLPGP
QGPKGQKGENSGT00900000140804_1_Linker_0_1 (189, 203)
EPYALSREERDKYRPF01391 // ENSGT00900000140804_1_Domain_1 (203, 251)
GEPGEPGLVGFQGPPGRPGPIGQMGPVGAPGRPGPPGP
PGPKGHPGNRENSGT00900000140804_1_Linker_1_2 (251, 322)
GLGF--YGEKGEKGDVGQPGPNGVPSDT---TLIGPTTTTYHPDLYKGEK
GSEG---DPGILGI----SLKPF01391 // ENSGT00900000140804_1_Domain_2 (322, 449)
GEEGIMGFPGPRGLPGM---DGEKGALGQ----------
---------------------------------------------------------KGS
RGLDGFQGPS-GPRGPKGERGERGPPGPPAYLPHP-------------------------
-----------------------------------------SLAKGARGDPGFQGAQGEP
GSRGLPGDPGPIGL---P--------------------------GLSIGDGD-EKRGLPG
EMGPKGFKGEQGLPARYPGPPGADGKPGLQGVPGPAGLPGPD------------------
-GFLFGLKGSEGRVGYPGPSGFPGARGQKGWKG---------------------------
-----------------------EAGDC-KCD---QFIR
PF01391 // ENSGT00900000140804_1_Domain_6 (760, 1015)
GLPGLQGAKGFPGVIGELGKK
GDQGEPGQHGIPGFPGFKGAPGLIGI----------------------------------
------------------------------------------------------------
------------------------------------------------------------
-----------------------------------------PGP---KG--IKGENSGT00900000140804_1_Linker_6_7 (1015, 1022)
DSRTIT
TPF01391 // ENSGT00900000140804_1_Domain_7 (1022, 1079)
KGERGQPGVPGVHGMKGDDGIPGRDGLDGFPGL-----PGP-PGDGIKGLPGDAGYPGIPGTKGFPGEVGPPGLGLPGPKGERGFPGDAGLPGPPGRPGLPGPPGTPGQIDCD---TGVKRPIGDGQQEAI--QPGCI--GGPP-----------------------GLPGQPGPP---G----------------PTGFKGLRGTPGFPGADGAPGLKGFPGDPGREGFPGPP-GFMGPRGSKGATGLPGPNGLPGPVGLPGPAGPPGDRGIPGEVLGAQPGSRGDAGLPGHPGLKGLPGERGPPGFRGS-----QGMPGMPGLKGQLGFPGPSGQPGLPGPPGQYGF-PGAPGREGPLGLPGSP-GL--------------------GGLPGDRGEPGDTGVPGPVGMKGLSGDKGDAGVSGERGHPGSPGFKGMAGMPGIPGQK-------GERGSPGMDGFQGMLGLKGRPGLPGTKGEAGFFGI----PGLKGLAGEPGVKGSRGDPGPPGPPPVILPG-MKDIKGEKGDEGPMGLKGYLGLK-------------GIQGMPGVPGLAGIP-GLPGRPGYIKGV-KGDIGVPGVPGLPGFPGVSGPPGIIGFPGFTGSRGEKGTPGR------------------------------AGLYGETGPTGDF---GDIGDTIDLPGSPGLKGERGITGTP-GLKGFFGEKGAEGEVGFPGITGMAGGQGSPGLKGQTGFPGLTGLQGPQGEPGRIGVPG--DKGDYGWPGAPGLP---------------------------------GFPGIRGISGLHGLPGTKGFPGSPGADLPG----------D---PGFPGPTGDRGDRGEANILPGPVGVPGQKGEQGAP-----------------------------------GERGPAGSPGLQGFPGV-SPPSNISGSPGDVGS----PGIFGLEGYRGPPGPPGPAALPGSKGDEGNPGASGTPGIKGWVGDPGPQGRPGVFGLPGVKG---------PKGEQGFMGNTGPS-----GAVGDRGPKGPKGDQGFPG-------APGSMGSPGIPGIPQRIAIQPGPMGPQGRRGLPGTLGEMGPQGPPGDP----------------------------------------------------GFRGAPGKAGPQGRGGVSAVPGFRGDQGPMGLQGPVGQE----GEPGRPGSPGLPGMPGRSVSIGYLLVKHSQTDQEPMCPVGMNKLWSGYSLLYFEGQEKAHNQDLGLAGSCLARFSTMPFLYCNPGDVCYYASRNDKSYWLSTTAPLPMMPVAEEE--IKPYISRCSVCEAPAVAIAVHSQ--DVSIPHCPAGWRSLWIGYSFLMHTAAGDEGGGQS----LVSPGSCLEDFRAT---PFIECNG-GRGTCHYFANKYS--FWLTTIPEQSFQGSPSA-----DTLKAGLIRTHISRCQVCMKNL----------------
PF01391 // ENSGT00900000140804_1_Domain_9 (1067, 1127)
GIKGLPGDAGYPGI
PGTKGFPGEVGPPGLGLPGPKGERGFPGDAGLPGPPGRPGLPGPPGENSGT00900000140804_1_Linker_9_10 (1127, 1241)
TPGQIDCD---TGV
KRPIGDGQQEAI--QPGCI--GGPP-----------------------GLPGQPGPP---
G----------------PTGFKGLRGTPGFPGADGAPGLKPF01391 // ENSGT00900000140804_1_Domain_10 (1241, 1297)
GFPGDPGREGFPGPP-GFMG
PRGSKGATGLPGPNGLPGPVGLPGPAGPPGDRGIPGEVLGAQPGSRGDAGLPGHPGLKGL
PGERGPPGFRGS-----QGMPGMPGLKGQLGFPGPSGQPGLPGPPGQYGF-PGAPGREGP
LGLPGSP-GL--------------------GGLPGDRGEPGDTGVPGPVGMKGLSGDKGD
AGVSGERGHPGSPGFKGMAGMPGIPGQK-------GERGSPGMDGFQGMLGLKGRPGLPG
TKGEAGFFGI----PGLKGLAGEPGVKGSRGDPGPPGPPPVILPG-MKDIKGEKGDEGPM
GLKGYLGLK-------------GIQGMPGVPGLAGIP-GLPGRPGYIKGV-K
PF01391 // ENSGT00900000140804_1_Domain_15 (1613, 1702)
GDIGVPGV
PGLPGFPGVSGPPGIIGFPGFTGSRGEKGTPGR---------------------------
---AGLYGETGPTGDF---GDIGDTIDLPGSPGLKGERGITGTP-GLKGFFGEKGAEGEV
GFPGITGMAGGQGSPGLKGQTGFPGLTGLQGPQGEPGRIGVPG--DKGDYGWPGAPGLP-
--------------------------------GFPGIRGISGLHGLPGTKGFPGSPGADL
PG----------D---PGFPGPTGDRGDRGEANILPGPVGVPGQKGEQGAP---------
--------------------------GERGPAGSPGLQGFPGV-SPPSNISGSPGDVGS-
---PGIFGLEGYRGPPGPPGPAALPGSKGDEGNPGASGTPGIKGWVGDP
PF01391 // ENSGT00900000140804_1_Domain_20 (2030, 2108)
GPQGRPGVFGL
PGVKG---------PKGEQGFMGNTGPS-----GAVGDRGPKGPKGDQGFPG-------A
PGSMGSPGIPGIPQRIAIQPGPMGPQGRRGLPGTLGEMGPQGPPGDP-------------
---------------------------------------GFRGAPGKAGPQGRGGVSAVP
GFRGDQGPMGLQGPVGQE----GEPGRPGSPGLPGMPGRSVSIGY
PF01413 // ENSGT00900000140804_1_Domain_23 (2266, 2371)
LLVKHSQTDQEPMCP
VGMNKLWSGYSLLYFEGQEKAHNQDLGLAGSCLARFSTMPFLYCNPGDVCYYASRNDKSY
WLSTTAPLPMMPVAEEE--IKPYISRCSVCENSGT00900000140804_1_Linker_23_25 (2371, 2376)
EAPAVPF01413 // ENSGT00900000140804_1_Domain_25 (2376, 2504)
AIAVHSQ--DVSIPHCPAGWRSLWI
GYSFLMHTAAGDEGGGQS----LVSPGSCLEDFRAT---PFIECNG-GRGTCHYFANKYS
--FWLTTIPEQSFQGSPSA-----DTLKAGLIRTHISRCQVCMKNL--------------
--------------
ENSTGUP00000009053 (view gene)
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
----------------------------------------VSMG
PF01413 // ENSGT00900000140804_1_Domain_23 (2265, 2372)
YLLVKHSQSDQEPMCP
VGMNKLWSGYSLLYFEGQEKAHNQDLGLAGSCLSRFSTMPFLYCNPGDICYYASRNDKSY
WLSTTAPLPMMPVAEED--IKPYISRCSVCEENSGT00900000140804_1_Linker_23_25 (2372, 2375)
APAPF01413 // ENSGT00900000140804_1_Domain_25 (2375, 2505)
VAIAVHSQ--EASIPHCPEGWRSLWI
GYSFLMHTAAGDEGGGQS----LVSPGSCLEDFRAT---PF-ECNG-ARGTCHYFANKYS
--FWLTTI-DQPFQSKPSG-----DTLKAGLIRSHISRCQVCMKNL--------------
--------------
ENSTSYP00000000129 (view gene)
-------------------------XXXXXWLLLGAVTVGFLARSASAGVKKFDVPCGGR
DCS-GGCQCYPEKGGRGQPGPV
PF01391 // ENSGT00900000140804_1_Domain_0 (83, 142)
GPQGYNGPPGLQGFPGLQGRKGDKGERGAPGITGPKGD
VGARGVSGFPGADGIP---GHPGQGGPRGRPGYDGCNGTRGDAGPQGPPGSGGFSGPPGP
QGPKGQKGEPYALSREDRDKYR
PF01391 // ENSGT00900000140804_1_Domain_1 (203, 251)
GEPGEPGLVGFQGPPGHPGPVGQMGPVGAPGRPGPPGP
PGPKGQPGNRENSGT00900000140804_1_Linker_1_2 (251, 322)
GLGF--YGEKGEKGDMGQPGPNGIPSDT-IHPIIGPTKTTIHLDHYKGEK
GS---EGEPGIKGF----PLKPF01391 // ENSGT00900000140804_1_Domain_2 (322, 448)
GEEGIMGFPGARGPPGF---DGEKGSPGQ----------
---------------------------------------------------------KGS
RGLDGYQGSD-GPRGPKGEAGDPGPPGPPAYSPHP-------------------------
-----------------------------------------SLAKGTRGDPGFPGGHGEP
GSRGEPGDQGPPGP---P--------------------------GLSIGDTD-EKRGLPG
EMGPKGFRGDVGVPARYPGPPGADGKPGLRGLPGPAGQPGPD------------------
-GFLFGLKGSEGGVGFPGPSGSPGARGHKGWKG---------------------------
-----------------------DAGDC-RCAEGDQFISRV
PF01391 // ENSGT00900000140804_1_Domain_6 (762, 1015)
PGLPGPKGFPGVNGEPGRK
GDQGDPGQHGIPGFSGFKGAPGSTGP----------------------------------
------------------------------------------------------------
------------------------------------------------------------
-----------------------------------------PGP---KG--AKGDSRTIT
TKGERGQPGIPGVPGVKGEDGVPGRVGLDGFPGL-----TGP-PGDGIQGPPGDRGYPGA
PGTKGLPGEMGPPGLGLPGPKGQRGHPGDAGSRGPPGFPGPPGLPGAPGQIDCD---TGV
KRPIGGDRQETV--PPGCV--GGPK-----------------------GLPGLPGPP---
G----------------LTGAKGPRGVPGFLGADGTPGLKGLPGDPGREGFPGPP-GFIG
PRGSKGTVGLPGPDGSQGSAGLPGPAGPPGDRGLPGKVLGAQPGPRGDAGLPGHPGLKGI
PGERGPPGFRGS-----QGMPGMPGLKGQLGFPGPSGQPGLPGPPGQHGF-PGGPGPEGP
LGLTGAP-GL--------------------GGLPGDR
PF01391 // ENSGT00900000140804_1_Domain_12 (1418, 1482)
GEPGDTGVPGPVGMKGVSGDRGD
AGMSGEQGHRGNPGFKGMLGMPGTPGLK-------GDRGSPGMDGFQGMPGLQGRPGFGG
SKGEAGFFGI----PGLKGLAGEPGVKGSRGEPGLPGPPPVILPG-MKDVKGEKGDEGPM
GLKGYLGLK-------------GLQGMPGIPGLSGVP-GLPGRPGYVKGV-K
PF01391 // ENSGT00900000140804_1_Domain_15 (1613, 1700)
GDIGVPGV
PGLPGFPGVAGPPGIIGFPGFTGSRGEKGALGR---------------------------
---EGLYGETGPAGDF---GDIGDAVNLPGRTGLKGDRGGTGVP-GLKGFFGEKGSEGNI
GFPGIPGVTGVQGPPGLKGQTGFPGLTGLQGPQGDPGRVGRPG--DKGTHGWPGVPGLP-
--------------------------------GFPGIRGIGGLDGLPGTKGFPGPPGTDV
HG----------D---PGFSGSPGDRGDPGEANTFPGPVGTPGQKGEQGAP---------
--------------------------GQRGRAGNPGLQGFPGI-TPPSNISGI
PF01391 // ENSGT00900000140804_1_Domain_19 (1974, 2034)
PGDKGA-
---PGIFGPKGYQGPPGQPGPAALPGSKGDAGSPGTPGAPGTKGWVGDPGSQGRPGVFGL
PGEKG---------PRGEQGLMGNTGAT-----GTVGDRGPKGPKGDRGFPGA------S
-GSVGAPGIAGIPQKIAALPGTAGPRGRGGPPGAPGEMGPQGPPGEP-------------
---------------------------------------GFRGAPGKAGPQGRGGVSAVP
GFRGGQGPLGHQGPIGQE----GEPGRPGSPGLPGMPGRSVSIGY
PF01413 // ENSGT00900000140804_1_Domain_23 (2266, 2371)
LLVKHSQTDQEPMCP
VGMNKLWSGYSLLYFEGQEKAHNQDLGLAGSCLARFSTMPFLYCNPGDVCYYASRNDKSY
WLSTTAPLPMMPVAEDN--IKPYISRCSVCENSGT00900000140804_1_Linker_23_25 (2371, 2376)
EAPAVPF01413 // ENSGT00900000140804_1_Domain_25 (2376, 2504)
AIAVHSQ--DVSIPHCPAGWRSLWI
GYSFLMHTAAGDEGGGQS----LVSPGSCLEDFRAT---PFIECNG-GRGTCHYYANKYS
--FWLTTIPEQSFQGSPSA-----DTLKAGLIRTHISRCQVCMKNL--------------
--------------
ENSCPOP00000009528 (view gene)
ENSHGLP00100016468 (view gene)
------------------------------WLLLGAVTVGLLAHSVLAGVKKLDVPCGGR
DCS-GGCRCYPEKGGR
PF01391 // ENSGT00900000140804_1_Domain_0 (77, 133)
GLPGPAGLPGFHGPPGLQGFPGLQGRKGDKGERGAPGTPGPKGD
VGPRGVSGFPGANGIP---GYPGQGGPRGRPGYDGCNGTRGDTGPQGPSGSEGFPGTFGP
PGPKGQKGEPYALSREDQDRYRGEPGEPGLVGFQGPPGRPGPMGPMGPVGAPGR------
------PGNRGLGF--YGEKGEKGDQGQPGPNGIPSDT-THPIVAPTGSTIHPDMYKGEK
GS---QGEPGFKGI----SYKGEEGIMGFPGTRGPPGL---DGERGFPGQ----------
---------------------------------------------------------KGS
KGRDGYQGPD-GPQGSKGELGEPGPPGPPAYSPHP-------------------------
-----------------------------------------SLAKGVRGDPGFPGAHGEP
GSRGEPGDPGLPGPQGPP--------------------------GLPSVDQG-DRTGRPG
ETGLKGFTGDRGIPARYP
PF01391 // ENSGT00900000140804_1_Domain_5 (619, 747)
GPPGTDGRPGPPGPPGPAGPPGPD------------------
-GFLFGLKGAQGSRGYAGLPGTPGVNGQKGRKG---------------------------
-----------------------DPGENSGT00900000140804_1_Linker_5_6 (747, 760)
NC-KCAESDQFPGPF01391 // ENSGT00900000140804_1_Domain_6 (760, 1015)
GLPGLPGLKGFPGANGKPGWK
GEPGDPGQHGIPGFPGFKGAPGSIGA----------------------------------
------------------------------------------------------------
------------------------------------------------------------
-----------------------------------------PGP---KG--AKGDSRTVT
TKGERGQPGIPGVHGMKGDDGIPGRDGLDGFPGL-----PGP-PGDGIRGPPGDSGPPGA
PGTKGPPGEVGPPGLGMLGREGEQGFPGDVGLPGSPGFPGPPGAPGSPGELDCE---TGV
KSPTGGDTQEPV--QPGCT--G
PF01391 // ENSGT00900000140804_1_Domain_10 (1163, 1258)
GPK-----------------------GSPGLPGPP---
G----------------PPGDKGPRGTPGFLGVDGPPGLKGFPGDPGREGFPGPP-GFMG
PRGSKGAVGLPGPDGLQGTSGLPGPAGPPGDRGIPGEVLGAQPGPRGDTGLPGHPGRKGL
PGEQGAPGFRGS-----QGMPGMPGPKGQLGFPGPSGLPGLPGRAGQHGF-PGAPGPEGP
LGLPGVS-GS--------------------EGLPGDRGDPGDTGTPGPVGMKGLAGDRGD
TGLVGDQGRPGDPGFKGMDGMPGIPGLK-------GDRGSPGMQGFQGMIGLRGQPGTPG
TKGQAGFFGL----PGPKGPAGQRGGKGNRGEPGPPGPPPAIMPG-MKF
PF01391 // ENSGT00900000140804_1_Domain_14 (1550, 1606)
IKGEKGDEGPM
GLKGHLGLK-------------GLQGMPGVPGQAGVP-GLPGRPGHIKGI-KGDIGVPGS
PGLPGFPGVTGPPGVIGFPGFTGSRGDKGAPGS---------------------------
---SGLYGDDGLTGDF---GDIGDTIDLPGSPGLKGERGIAGTP-GLKGFFGERGAHGEV
GFPGITGLAGPQGQPGLKGQTGFPGLTGLQGPQGDPGQIGFPG--EKGADGWPGVPGLP-
--------------------------------GFPGLRGISGLHGLPGPKGFTGSPGADM
PG----------D---PGFPGPSGDRGDPGEPNTFPGSTGAPGQKGEQGAP---------
--------------------------GERGPAGRPGMQGFPGI-SPPSNISGLPGDVGS-
---PGIFGLKGYQGPQGPPGPAALQGSKGDAGSGGAPGVVGTKGWIGDQ
PF01391 // ENSGT00900000140804_1_Domain_20 (2030, 2108)
GPQGRPGVFGV
PGEKG---------PRGEPGLMGSTGST-----GAVGDRGPKGPKGDQGFPG-------A
PGLVGSPGIAGIPQRITAPPGAMGPEGRRGRPGAQGEMGPQGPPGDP-------------
---------------------------------------GFRGVPGKPGPQGRGGVSALP
GFRGDPGAVGHQGPIGQE----GEPGRPGSPGLPGMPGRSVSIGY
PF01413 // ENSGT00900000140804_1_Domain_23 (2266, 2371)
LLVKHSQADQEPMCP
VGMNKLWSGYSLLYFEGQEKAHNQDLGLAGSCLARFSTMPFLYCNPGDVCYYASRNDKSY
WLSTTAPLPMMPVAEDD--IKPYISRCSVCENSGT00900000140804_1_Linker_23_25 (2371, 2376)
EAPATPF01413 // ENSGT00900000140804_1_Domain_25 (2376, 2504)
AIAVHSQ--DVSIPHGPAGWRSLWI
GYSFLMHTAAGDEGGGQS----LMSPGSCLEDFRAT---PFIECNG-GRGTCHYFANKYS
--FWLTTIPDTSFQGTPSA-----DTLKAGLIRTHISRCQVCMKNL--------------
--------------
ENSTGUP00000015565 (view gene)
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
---------------------------------------------------GP-------
-RGDRGPNGLPGLQGHPGLTGKSGAPGAPGPKGLP-----AAAGSRGDVGLPGFPGLKGA
PGDQGVPGTR-----------------GDPGPQGLPGLTGLPGTPGTHGF-PGPPGNRGP
DGGPGSQGPL---------------GEEGLNC
PF01391 // ENSGT00900000140804_1_Domain_12 (1413, 1469)
PPGARGEDGEQGFPGPVGLKGLSGDKGD
MGHTGLPGIRGVTGPPGISGMDGFPGDKG-------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
--------------
ENSFALP00000004596 (view gene)
ENSACAP00000013906 (view gene)
ENSAMEP00000017981 (view gene)
ENSGALP00000027157 (view gene)
------MDLSPLPGARVSV-CQD------VGLLLVLLEVVLSAGRVDAGGKSYTGPCGGR
DCS-GGCQCFPEKGARGQPGILGSQGFPGPPGLMGIPGLQGPKGHKGERGHPGISGPKGE
TGQRGVTG
PF01391 // ENSGT00900000140804_1_Domain_0 (129, 189)
FPGADGVP---GHPGQPGSRGKPGHDGCNGTVGDPGDPGTPGHSGFPGTIGV
QGPKGQKGENSGT00900000140804_1_Linker_0_1 (189, 202)
EPYVLPPDIASRHPF01391 // ENSGT00900000140804_1_Domain_1 (202, 251)
RGDPGDPGFTGFPGAPGTLGIQGPIGPRGVPGRPGPPGS
PGPQGPQGNRENSGT00900000140804_1_Linker_1_2 (251, 321)
GLGF--YGEKGEQGSPGPPGPPGLPTRE-LIGVPTDK-----HKGERGEP
GQ---KGEAGFPGVLLFAPEPF01391 // ENSGT00900000140804_1_Domain_2 (321, 445)
KGEEGVMGFPGQRGLPGN---DGFPGLSGE----------
---------------------------------------------------------RGF
PGFDGQPGQY-GPRGGKGEQGEMGENSGT00900000140804_1_Linker_2_3 (445, 526)
PPGPPAYVPYR-------------------------
-----------------------------------------IPRKPF01391 // ENSGT00900000140804_1_Domain_3 (526, 614)
GVRGDPGSPGASGLR
GEQGEQGDRGLPGI---P--------------------------GFSDGDA--DKPGLPG
EIGPKGEKGEEGSENSGT00900000140804_1_Linker_3_5 (614, 619)
PAYQAPF01391 // ENSGT00900000140804_1_Domain_5 (619, 746)
GPPGFPGKHGEPGIRGPPGPPGTP------------------
-GSLFGLKGQEGTPGRPGVQGFPGPRGQRGPKG---------------------------
-----------------------EEENSGT00900000140804_1_Linker_5_6 (746, 759)
GDCSKCLLSDELRPF01391 // ENSGT00900000140804_1_Domain_6 (759, 1015)
RGSTGPRGPPGFPGTPGQPGRK
GEPGDQGPHGIPGYAGAKGQSGQEGL----------------------------------
------------------------------------------------------------
------------------------------------------------------------
-----------------------------------------PGP---KG--EKGENSGT00900000140804_1_Linker_6_7 (1015, 1023)
DSIYIT
TKPF01391 // ENSGT00900000140804_1_Domain_7 (1023, 1089)
GTKGIRGDPGLPGIRGEDGFPGRDGLDGLPGL-----PGL-PGDGIRGLPGDPGYPGE
LGPKGFPGEIGLPGEGYPGPKGYRGLPGDRGTDGHPGPPGLPGPPGEPGQLDCG---QVIEDFSRGEATDPIWSGGGCV--RPPK-----------------------GSQGNPGLP---G----------------ATGTKGARGFPGDPGPVGFPGLNGTRGDPGREGYPGPP-GFIGPRGDRGPNGLPGLQGHPGLMGKSGAPGLAGQKGAPGDVLGAAAGPRGDDGLPGFPGLKGAPGDQGIPGIRGA-----DGNPGLPGPKGDPGLQGLPGLMGLPGTPGTHGF-PGPPGNRGPDGGPGSQ-GP--------------------LGPPGARGEDGEQGFPGPVGMKGLSGDKGETGFPGLQGIPGVTGPPGISGMDGFPGDK-------GSRGSPGIDGFKGMPGLKGRPGIKGIKGEFGLLGT----RGDKGAQGARGFKGDRGEQGPPGEPPKLKPSMMMEVKGEKGDAGETGTKGFFGIK-------------GSKGMPGLPGKTGIP-GSPGHPSYVPGV-KGDIGAKGLTGLKGYPGPTGSPGIRGFPGSTGGRGDKGAPGI------------------------------SGHFGTPGSHGEI---GEPGDTINLPGMPGLKGEVGVPGLT-GLRGGPGQKGEGGDPGLPGIEGLKGIQGVPGSLGQKGLPGLVGPPGQQGSPGTPGFQG--EKGAPGWPGLPGQA---------------------------------GLPGLRGISGLHGLPGTKGLPGSPGPDGYG----------S---AGFPGPVGDKGEAGEPSRVEGSQGPPGQKGDRGVP-----------------------------------GVQGPFGIPGQDGLPGP-PGISNISGYPGDTGS----PGLDGVPGYPGLHGQPGIPAPPGSKGESGRAGVSGQAGPKGTRGDPGLPGRPGIPGYPGPKGR---------KGEQGVIGFIGTV-----GFPGDLGPIGPKGDRGLTG------FQ-GPPGSPGLPPIPPRLVAEQGSPGPRGNAGPRGSPGDAGPQGPPGEP----------------------------------------------------GLRGLPGEPGLQGRGGIPAPPGSRGEQGAMGFQGPVGFE----GQPGRPGSPGLPGMPGRSVSIGYLLVKHSQSDQEPMCPIGMNKLWDGYSLLYFEGQEKAHNQDLGLAGSCLARFSTMPFLYCNPGDICYYANRNDKSYWLSTTAPLPMMPVAEEE--IRPYISRCSVCEAPAVAIAVHSQ--EASIPRCPEGWRSLWIGYSFLMHTAAGDEGGGQS----LVSPGSCLEDFRAT---PFIECNG-ARGTCHYFANKYS--FWLTTI-DQPFQSKPSA-----DTLKAGLIRSHISRCQVCMKNL------------
PF01391 // ENSGT00900000140804_1_Domain_9 (1073, 1130)
GDPGYPGELGPKGFPGEIGLPGEGYPGPKGYRGLPGDRGTDGHPGPPGLPGPPGEPGQLDCG---QVI
EDFSRGEATDPIWSGGGCV--RPPK-----------------------GSQGNPGLP---
G----------------ATGTKGARGFPGDPGPVGFPGLNGTRGDPGREGYPGPP-GFIG
PRGDRGPNGLPGLQGHPGLMGKSGAPGLAGQKGAPGDVLGAA
PF01391 // ENSGT00900000140804_1_Domain_11 (1303, 1364)
AGPRGDDGLPGFPGLKGA
PGDQGIPGIRGA-----DGNPGLPGPKGDPGLQGLPGLMGLPGTPGTHGF-PGPPGNRGP
DGGPGSQ-GP--------------------LGPPGARGEDGEQGFPGPVGMKGLSGDKGE
TGFPGLQGIPGVTGPPGISGMDGFPGDK-------GSRGSPGIDGFKGMPGLKGRPGIKG
IKGEFGLLGT----RGDKGAQGARGFKGDRGEQGPPGEPPKLKPSMMMEVKGEKGDAGET
GTKGFFGIK-------------GSKGMPGLPGKTGIP-GSPGHPSYVPGV-KGDIGAKGL
TGLKGYPGPTGSPGIRGFPGSTGGRGDKGAPGI---------------------------
---SGHFGTPGSHGEI---GEPGDTINLPGMPGLKGEVGVPGLT-GLRGGPGQKGEGGDP
GLPGIEGLKGIQGVPGSLGQKGLPGLV
PF01391 // ENSGT00900000140804_1_Domain_17 (1768, 1856)
GPPGQQGSPGTPGFQG--EKGAPGWPGLPGQA-
--------------------------------GLPGLRGISGLHGLPGTKGLPGSPGPDG
YG----------S---AGFPGPVGDKGEAGEPSRVEGSQGPPGQKGDRGVP---------
--------------------------GVQGPFGIPGQDGLPGP-PGISNISGYPGDTGS-
---PGLDGVPGYPGLHGQPGIPAPPGSKGESGRAGVSGQAGPKGTRGDP
PF01391 // ENSGT00900000140804_1_Domain_20 (2030, 2108)
GLPGRPGIPGY
PGPKGR---------KGEQGVIGFIGTV-----GFPGDLGPIGPKGDRGLTG------FQ
-GPPGSPENSGT00900000140804_1_Linker_20_22 (2108, 2203)
GLPPIPPRLVAEQGSPGPRGNAGPRGSPGDAGPQGPPGEP-------------
---------------------------------------GLRPF01391 // ENSGT00900000140804_1_Domain_22 (2203, 2259)
GLPGEPGLQGRGGIPAPP
GSRGEQGAMGFQGPVGFE----GQPGRPGSPGLPGMPGENSGT00900000140804_1_Linker_22_23 (2259, 2266)
RSVSIGYPF01413 // ENSGT00900000140804_1_Domain_23 (2266, 2371)
LLVKHSQSDQEPMCP
IGMNKLWDGYSLLYFEGQEKAHNQDLGLAGSCLARFSTMPFLYCNPGDICYYANRNDKSY
WLSTTAPLPMMPVAEEE--IRPYISRCSVCENSGT00900000140804_1_Linker_23_25 (2371, 2376)
EAPAVPF01413 // ENSGT00900000140804_1_Domain_25 (2376, 2504)
AIAVHSQ--EASIPRCPEGWRSLWI
GYSFLMHTAAGDEGGGQS----LVSPGSCLEDFRAT---PFIECNG-ARGTCHYFANKYS
--FWLTTI-DQPFQSKPSA-----DTLKAGLIRSHISRCQVCMKNL--------------
--------------
ENSMAUP00000023031 (view gene)
ENSMEUP00000012988 (view gene)
------------------------------------------------------------
----------------
PF01391 // ENSGT00900000140804_1_Domain_0 (77, 136)
GQPGPVGPQGYSGPPGLQGIPGLQGRKGDKGERGAPGITGPKGD
VGQRGVSGFPGADGIP---GHPGQGGPRGRPGSDGCNGTTGDAGTQGPPGSGGLPGLPGP
QGPKGQKGEPYAFSQAERDRY
PF01391 // ENSGT00900000140804_1_Domain_1 (202, 244)
RGESGEPGFIGLPGPPGYPGSPGKIGPVGAPGRPGPPGP
PGPKGQQXXXXXXX--XXXXXXXXXXXXXXXXXXXXXX-XX----XXXXXXXXXXXXXXX
XX---XXXXXXXXXXXGTSAKGEEGIVGYPGQRXXXXX---XXXXXXXXX----------
---------------------------------------------------------XXX
XXXXXXXXXX-XXXXXXGDIGDRGPPGAPSYSPHP-------------------------
-----------------------------------------SLIKXXX
PF01391 // ENSGT00900000140804_1_Domain_3 (529, 585)
GDPGDSGARGPD
GSPGEPGDPGLPGP---R--------------------------GTSEGDGV-ERRGLPG
EMGAKGFIGDPGYPALYPGPPGTDGKPGLPGPPGPAGPPGPD------------------
-XXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXX---------------------------
-----------------------DPGDC-KC----DI
PF01391 // ENSGT00900000140804_1_Domain_6 (758, 1016)
PSGLPGPPGPPGRPGANGEPGRK
GEIGDPGPHGIPGFSGFKGNPGIPGL----------------------------------
------------------------------------------------------------
------------------------------------------------------------
-----------------------------------------PGP---KG--QKGDENSGT00900000140804_1_Linker_6_7 (1016, 1022)
SREIT
TPF01391 // ENSGT00900000140804_1_Domain_7 (1022, 1061)
KGSRGDQGNPGIPGLQGEDGFPGQDGLDGFPGL-----PENSGT00900000140804_1_Linker_7_9 (1061, 1074)
GL-P---------PF01391 // ENSGT00900000140804_1_Domain_9 (1074, 1131)
DPGFPGV
PGLKGIPGESGPPGLGLPGPKGERGFPGDPGLSGSPGFPGPPGPPGAPGQENSGT00900000140804_1_Linker_9_10 (1131, 1192)
TXXX---XXX
XXXXX--XXXXXXXXXXCV--IAPK-----------------------GSPPF01391 // ENSGT00900000140804_1_Domain_10 (1192, 1271)
GLPGLP---
G----------------SPGAKGQRGIPGFPGADGFAGLKGFQGEAGREGFPGPP-GFVG
PRGSKGATGLPGLDGNPGSTGLPGAVGPAGQKGLPXXXXXXXXXXXXXXXXXXXXXXXXX
XXXXXXXXXXXX-----XXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXX-XXXXXXXXX
XXXXXXX-XX--------------------XXLPAKEGDRGDVGVPVPVGMKGLGGDRGD
TGFIGEQGRRGDPGLQGMPGMPGIPGQK-------XXXXXXXXXXXXXXXXXXXXXXXXX
XXXXXXXXXX----XXXXXXXXXXXXXXSRGEPGPPGEPPKFLPG-MIDFKGEKGDEGPV
PF01391 // ENSGT00900000140804_1_Domain_14 (1561, 1636)
GGKGYWGTK-------------GGQGMPGIPGQAGIP-GLPGIPGYIKGA-KGEIGVPGV
PGLQGFPGITGSPGIRGFPGFVGVRXXXXXXXX---------------------------
---XXXX---XXXXXXXXXXDEGDTINLPGRPGLKGELGISGLA-GEKGFAGEKGTPGDR
GFVGLMGIKGXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXX--XXXXXXXXXXXXXX-
--------------------------------XXXXXXXXXXXXXXXXXXXXXXXXXADG
YG----------N---PGFPGPFGDKGERGEPNII
PF01391 // ENSGT00900000140804_1_Domain_18 (1896, 1968)
PGPLGAPGQKGERGNL---------
--------------------------GQPGPSGSPGIEGLPGF-TAPSNISGLPGDKGA-
---PGLFGIQGYRGPPGAPGPSSSPGIKGDAGSPGLTGERGPKGWPGDPGPQGRPGVYGL
PGEKG---------PKGQQGLMGYHGIP-----GNIGDRGPKGPKGDRGLPG------PP
-GAIGSPGIPGIPLKIATEPGVAGPQGRRGPIGMQGEM
PF01391 // ENSGT00900000140804_1_Domain_21 (2139, 2254)
GPQGPPGDP-------------
---------------------------------------GLRGLPGNPGPQGRGGISALP
GFRGDQGPTGLQGPVGQE----GEPGRPGSPGLENSGT00900000140804_1_Linker_21_23 (2254, 2264)
PGMPGRSVSIPF01413 // ENSGT00900000140804_1_Domain_23 (2264, 2373)
GYLLVKHSQTDQEPMCP
VGMNKLWSGYSLLYFEGQEKAHNQDLGLAGSCLARFSTMPFLYCNPGDICHYASRNDKSY
WLSTTAPLPMMPVAEEE--IKPYISRCSVCEAENSGT00900000140804_1_Linker_23_25 (2373, 2374)
PPF01413 // ENSGT00900000140804_1_Domain_25 (2374, 2505)
AVAIAVHSQ--DVSIPHCPAGWRSLWI
GYSFLMHTAAGDEGGGQS----LVSPGSCLEDFRAT---PFIECNG-GRGTCHYFANKYS
--FWLTTIPEQNFQASPSA-----DTLKAGLIRTHISRCQVCMKNL--------------
--------------
ENSMGAP00000015755 (view gene)
ENSOARP00000006991 (view gene)
--------------MDREL-RAAARPALRRWLLLGAVTVGLLAQSVLAGVKKLDVPCGGR
DCS-GGCQCYPEKGGRGQPGPVGPQGYTGPPGLQGFPGLQGRKGDKGERGAPGITGPKGD
VGPRGVSGF
PF01391 // ENSGT00900000140804_1_Domain_0 (130, 189)
PGADGIP---GHPGQGGPRGPPGYDGCNGTMGDSGYAGPPGPGGFLGPRGP
QGPKGQKGENSGT00900000140804_1_Linker_0_1 (189, 203)
EPYALSSEDRDKYRPF01391 // ENSGT00900000140804_1_Domain_1 (203, 251)
GEPGEPGLVGLQGPPGRPGPVGQMGPVGAPGRPGPPGP
PGPKGQPGNRGLGF--YGEKGEKGDMGLQGPGGIPPDNGYV-EKPTPGYELLPEQYKGEK
GS---QGEPGRIGV----SLKGEEGVMGFSGPRGAPGF---DGEKGLPGQ----------
---------------------------------------------------------KXG
GGVAGWGGPLPAPSSAGGEMGNPGPPGLPAYSPHP-------------------------
-----------------------------------------SVAKGIRGEP----APGEP
GARGEPGDPGLPGR---P--------------------------GTTIGDED-EKRGLPG
EMGPKGFIGERGSPALYPGPPGAEWRVILGPPCPPPALGGPP---W-GRLEALPSGPPRA
GGYFPGI-----------CPGSSGHKNQKGWKG---------------------------
-----------------------DAGDC-KC--ADQFIR
PF01391 // ENSGT00900000140804_1_Domain_6 (760, 1015)
GPPGLPGPKGFAGANGQPGSK
GSQGDPGPHGLPGFSGFKGAPGNVGP----------------------------------
------------------------------------------------------------
------------------------------------------------------------
-----------------------------------------PGP---KG--MKGDSRTIT
TKGERGQPGVPGVPGLKGTDGIPGPPGLDGFHGL-----PGP-PGDGIKGPRGDAGQPGA
PGTKGLPGERGPPGLGLPGLKGERGFPGDAGLPGPPGFPGPPGLPGAPGQTDCD---SGV
KRPIGADGQETI--QPGCV--GGPK-----------------------GSPGQPGLP---
G----------------PPGAKGLRGVPGFSGADGVPGLKGLPGDPGREGFPGPP-GFMG
PRGSKGAVGPPGLDGLPGASGLPGPVGPPGDRGLPGEVLGAQ
PF01391 // ENSGT00900000140804_1_Domain_11 (1303, 1364)
PGPRGDSGLPGRPGLKGP
PGERGPPGFRGS-----QGMPGMPGQKGQPGSPGFSGQPGLPGAGGRAWVCPREPALQSR
RGSESAA-HG--------------------PGLPGDRGEPGDTGVPGPVGRKGGSGDRGD
PGQQGERGHPGLPGFKGVSGMPGTPGLS-------RARGSPGMDGFEGMLGLKGRPGLPG
IKGEAGFFGI----PGLKGLAGEPGVKGSRGDPGPPGPPPIILPG-MKDI
PF01391 // ENSGT00900000140804_1_Domain_14 (1551, 1605)
KGEKGDEGPM
GLKGYLGLK-------------GLPGMPGIPGLSGIP-GLPGRPGHIKGV-KGDTGVPGV
PGSPGFPGVPGSPGVMGFQGFTGSRGDKGAPGR---------------------------
---AGLFGEVGETGDF---GDIGDTIDLPGSPGLKGERGTTGIP-GQKGFLGERGTEGDV
GFPGITGLAGVQGPPGFQGQKGFPGLTGLQGPQGDPGRAGVPG--IKGDSGWPGNPGLP-
--------------------------------GLPGLRGISGLHGLPGNKGFPGSPGADV
HG----------D---PGFPGPAGDKGDPGEANTLPGPTGAPGQKGERGAP---------
--------------------------GERGPIGSPGLQGFPGI-TPPSNIS
PF01391 // ENSGT00900000140804_1_Domain_19 (1972, 2034)
GSPGDIGA-
---PGIFGLEGYRGPPGPPGPAALPGSKGDEGSPGTPGNPGIKGWVGDPGPQGRPGVFGL
PGEKG---------PKGEPGFMGNIGPT-----GSPGDRGPRGPKGDRGLPG-------A
PGAVGAPGIAGIPQRIAVE
PF01391 // ENSGT00900000140804_1_Domain_21 (2120, 2221)
RGPVGPQGRRGPPGAQGEMGPQGPPGEP-------------
---------------------------------------GFRGVQGKAGPQGRGGVSAVPENSGT00900000140804_1_Linker_21_23 (2221, 2265)
GFRGDQGPVGLQGPVGFE----GEPGRPGSPGLPGMPGRSISIGPF01413 // ENSGT00900000140804_1_Domain_23 (2265, 2330)
YLLVKHSQTEKEPMCP
VGMNKLWSGYSLLYFEGQEKAHNQDLGLAGSCPARVSTTPLKLCQSGQATFYLEGLLLSG
DLGTDGAAPVCPWGTEEEALDPL------CFNSHHEMRKKQANVLASLPP----------
-APPAQHTAAGDEG
PF01413 // ENSGT00900000140804_1_Domain_25 (2415, 2505)
GGQS----LVS-------LRAT---PFNECNG-ARGTCHYYANKYS
--FWLTTIPEQNFQGTPSA-----DTLKAGLIRTHISRCQVCMKNL--------------
--------------
ENSAPLP00000009420 (view gene)
ENSNLEP00000009690 (view gene)
--------------MGRDL--RAAGPALRRWLLLGTVTVGFLAQSVLAGVKKFDVPCGGR
DCS-GGCQCYPEKGGR
PF01391 // ENSGT00900000140804_1_Domain_0 (77, 132)
GQPGPVGPQGYNGPPGLQGFPGLQGRKGDKGERGAPGITGPKGD
VGARGVSGFPGADGIP---GHPGQGGPRGRPGYDGCNGTQGDSGPQGPPGSEGFTGPPGP
QGPKGQKGEPYALPKEERDRYR
PF01391 // ENSGT00900000140804_1_Domain_1 (203, 251)
GEPGEPGLVGFQGPPGRPGHVGQMGPVGAPGRPGPPGP
PGPKGQQGNRENSGT00900000140804_1_Linker_1_2 (251, 322)
GLGF--YGVKGEKGDVGQPGPNGIPSDT-LHPIIAPTGVTFHPDQYKGEK
GS---EGEPGVRGI----SLKPF01391 // ENSGT00900000140804_1_Domain_2 (322, 447)
GEEGIMGFPGPRGYPGL---SGEKGSPGQ----------
---------------------------------------------------------KGS
RGLDGYQGPD-GPRGPKGEAGDPGPPGLPAYSPHP-------------------------
-----------------------------------------SLAKGARGDPGFPGAQGEP
GSQGEPGDPGLPGP---P--------------------------GLSIGDGD-QRRGLPG
EMGPKGFIGEPGIPALYG
PF01391 // ENSGT00900000140804_1_Domain_5 (619, 746)
GPPGPDGKRGPPGPPGLPGPPGPD------------------
-GFLFGLKGAEGRAGFPGLPGSPGARGPKGWKG---------------------------
-----------------------DGENSGT00900000140804_1_Linker_5_6 (746, 760)
GDC-RCAEGDEAITPF01391 // ENSGT00900000140804_1_Domain_6 (760, 1015)
GLPGLPGPKGFAGINGEPGRK
GDKGDPGQHGLPGFPGLKGVPGNVGA----------------------------------
------------------------------------------------------------
------------------------------------------------------------
-----------------------------------------PGP---KG--AKGDSRTIT
TKGERGQPGVPGVPGMKGDDGSPGRDGLDGFPGL-----PGP-PGDGIKGPPGDPGYPGI
PGTKGTPGERGPPGLGLPGLKGQRGFPGDAGLPGPPGFLGPPGPAGTPGQIDCD---TDV
KIAIGGDRQETI--QPGCV--GGPK-----------------------GLPGLPGPP---
G----------------PTGAKGLRGIPGFSGADGGPGPKGLPGDAGREGFPGPP-GFIG
PRGSKGAVGLPGPDGSPGPIGLPGPDGPPGERGLPGEVLGAQPGPRGDAGVPGQPGLKGL
PGDRGPPGFRGS-----QGMPGMPGLKGQPGLPGPSGQPGPYGPPGLHGF-PGAPGQEGP
LGLPGIP-GL--------------------EGLPGDR
PF01391 // ENSGT00900000140804_1_Domain_12 (1418, 1482)
GDPGDTGAPGPVGMKGLSGDRGD
AGLAGERGHPGSPGFKGIDGMPGTPGLK-------GDRGSPGMDGFQGMPGLKGRPGFPG
SKGEAGFFGI----PGLKGLAGEPGFKGSRGDPGPPGPPPVILPG-MKDIKGEKGDEGPM
GLKGYLGAK-------------GIQGMPGIPGLSGIP-GLPGRPGHIKGV-KGDIGAPGI
PGLPGFPGVAGPPGITGFPGFIGSRGEKGAPGR---------------------------
---AGLYGEIGPTGDF---GDIGDTINLP
PF01391 // ENSGT00900000140804_1_Domain_17 (1710, 1765)
GRPGLKGERGTTGIP-GLKGFFGEKGTEGDI
GFPGITGVTGVQGPPGLKGQTGFPGLTGPPGSQGEPGRIGLPG--GKGDDGWPGAPGLP-
--------------------------------GFPGLRGIRGLHGLPGTKGFPGSPGSDI
HG----------D---PGFPGPPGERGDPGEANTLPGPVGVPGQKGDQGAP---------
--------------------------GERGPPGSPGLQGFPGI-TPPSNIS
PF01391 // ENSGT00900000140804_1_Domain_19 (1972, 2034)
GAPGDKGA-
---PGIFGLKGYRGPPGPPGSAALPGSKGDTGNPGAPGTPGTKGWAGDSGPQGRPGVFGL
PGEKG---------PRGEQGFMGNTGPT-----GAVGDRGPKGPKGDPGFPG-------A
PGTVGAPGIAGIPQKIAVQPGTVGPQGRRGPPGAPGEMGPQGPPGEP-------------
---------------------------------------GFRGAPGKAGPQGRGGVSAVP
GFRGDEGPIGHQGPIGQE----G-------------------------------------
---------------------------
PF01413 // ENSGT00900000140804_1_Domain_23 (2308, 2371)
LAGSCLARFSTMPFLYCNPGDVCYYASRNDKSY
WLSTTAPLPMMPVAEDE--IKPYISRCSVCENSGT00900000140804_1_Linker_23_25 (2371, 2376)
EAPAVPF01413 // ENSGT00900000140804_1_Domain_25 (2376, 2504)
AIAVHSQ--DVSIPHCPAGWRSLWI
GYSFLMHTAAGDEGGGQS----LVSPGSCLEDFRAT---PFIECNG-GRGTCHYYANKYS
--FWLTTIPEQSFQGSPSA-----DTLKAGLIRTHISRCQVCMKNL--------------
--------------
ENSSTOP00000019851 (view gene)
--------------MGQER-CAAAGPALRRWLLLGALAVGFLAQSVLAGVKKYDVPCGGR
DCS-GGCQCYPEKGGRGQPGPVGPQGYTGPPGLQGFPGLQGRKGDKGERGAPGTAGPKGD
VGARGVSGFP
PF01391 // ENSGT00900000140804_1_Domain_0 (131, 190)
GADGIP---GHPGQGGPRGRPGYDGCNGTRGDTGPQGPSGPAGFPGSPGP
QGPKGQKGEENSGT00900000140804_1_Linker_0_1 (190, 203)
PYALSKEDRDKYRPF01391 // ENSGT00900000140804_1_Domain_1 (203, 251)
GEPGEPGLVGFQGPPGRPGPMGPMGPVGALGRPGPPGP
PGPKGQPGNRENSGT00900000140804_1_Linker_1_2 (251, 297)
GLGF--YGQKGEKGDVGLPGPNGIPSDT-FHPIDGPTKTTIHPDQYPF01391 // ENSGT00900000140804_1_Domain_2 (297, 425)
KGEK
GS---RGEPGIKGI----SVKGEEGIVGFSGPRGPSGF---DGEKGSPGQ----------
---------------------------------------------------------KGS
RGLDGYQGTD-GPRGPKGEAGDQGPPGPPVYSPHP-------------------------
-----------------------------------------SLAKGARGDPGFPGPHGEP
GSRGEPGDPGPTGP---P--------------------------GLSFGDED-EKRGLPG
EMGPKGFQGEPGIPAFYPGPPGADGKPGLPGAPGPAGPPGPD------------------
-GYLFGLKGAEGRVGYPGPSGFPGARGQKGWKG---------------------------
-----------------------DAGDC-KCAESDQFIR
PF01391 // ENSGT00900000140804_1_Domain_6 (760, 1015)
GLPGLPGPKGFSGINGEPGKK
GDQGDPGQHGIPGFPGFKGVSGVVGA----------------------------------
------------------------------------------------------------
------------------------------------------------------------
-----------------------------------------PGP---KG--IKGDSRTIT
TKGERGQPGVPGVHGIKGDDGVPGRDGLDGFPGL-----PGP-PGDGIQGPPGDAGHPGI
PGTKGIPGEMGPPGLGLPGPKGERGFPGDAGLPGPPGFPGPPGPPGTPGQPDCD---TDV
KRPIGGDRQETL--QPGCV--RGPK-----------------------GLPGRPGPP---
G----------------PPGTKGLRGIPGFLGADGTPGLKGFPGDPGREGFPGPP-GFMG
PRGSKGAVGLPGPDGLPGPVGLPGPAGPPGERGLPGEVLGAQPGPRGDAGLPGHPGLKGL
PGERGPPGFRGS-----QGMPGMPGLKGQLGFPGPSGQPGRPGPPGQHGF-PGAPGPEGP
LGRPGAP-GL--------------------GGLPGDRGDPGDTGVPGPVGMKGLSGDRGD
AGFSGERGHPGTPGFKGMAGMPGIP
PF01391 // ENSGT00900000140804_1_Domain_13 (1466, 1535)
GQK-------GERGSPGMHGFQGMVGLKGRSGFPG
TKGEAGFFGV----PGLKGLAGEPGVKGSRGDPGPPGPPPIILPG-MKDIKGEKGDEGPM
GLKGYLGVK-------------GIQGMPGIPGLAGIP-GLPGRPGHIKGA-KGDIGVPGV
PGLPGFPGVTGPPGITGFPGFTGSRGDKGAPGR---------------------------
---AGFYGEIGPTGDF---GDIGDTIDLPGSPGLKGERGIAGTP-GLKGFFGEKGAEGEV
GFPGIAGMT
PF01391 // ENSGT00900000140804_1_Domain_17 (1750, 1837)
GTQGPPGPRGQTGFPGLTGLRGPQGDPGRIGVPG--DKGDSGWPGVPGLP-
--------------------------------GFPGIRGISGLHGLPGTKGFPGSPGADI
HG----------E---PGFPGPTGDRGDPGEANTLPGPTGVPGRKGEQGAP---------
--------------------------GERGPAGSPGLQGFPGI-TPPSNISGSPGDKGA-
---PGIFGLEGYRGPPGPPGPAAFPGSKGDDGSPGASGVPGNKGWVGDPGPQ
PF01391 // ENSGT00900000140804_1_Domain_20 (2033, 2111)
GRPGVFGL
PGEKG---------PRGEPGFMGNTGAT-----GAVGDRGPKGPKGDQGFPG------AP
-GSMGSPGIAGVPLTISVQPGTMGPQGRRGPPGAQGEMGPQGPPGEP-------------
---------------------------------------GFRGAPGKAGPQGRGGVSAIP
GFRGDQGPMGHQGPVGQE----GEPGRPGIPGLPGMPGRSVSIGY
PF01413 // ENSGT00900000140804_1_Domain_23 (2266, 2371)
LLVKHSQTDQEPMCP
VGMNKLWSGYSLLYFEGQEKAHNQDLGLAGSCLARFSTMPFLYCNPGDICYYASRNDKSY
WLSTTAPLPMMPVAEDD--IKPYISRCSVCENSGT00900000140804_1_Linker_23_25 (2371, 2376)
EAPAVPF01413 // ENSGT00900000140804_1_Domain_25 (2376, 2504)
AIAVHSQ--DVSIPHCPAGWRSLWI
GYSFLMHTAAGDEGGGQS----LVSPGSCLEDFRAT---PFIECNG-ARGTCHYYANKYS
--FWLTTIPEQSFQGTPSA-----DTLKAGLIRTHISRCQVCMKNL--------------
--------------
ENSJJAP00000009510 (view gene)
--------------MAGER-CAMPGPTLRRWLWLGTVTVAFLTQSVLAGVKKFDVPCGGR
DCS-GGCQCYPEKGGRGQPGPVGPQGYNGPPGLQGFPGLQGRKGDKGERGAPGPTGPKGD
VGARGVSGFP
PF01391 // ENSGT00900000140804_1_Domain_0 (131, 189)
GADGIP---GHPGQGGPRGRPGYDGCNGTRGDAGPQGPSGSGGFPGLPGP
QGPKGQKGEPYALSERDRDKYRGEPGEPGLVGYQGPPGRPGPIGQMGPVGAPGRPGPPGP
PGPKGQPGNRGLGF--YGEKGEKGDVGQPGPNGIPSDV---TLIGPTMTTFPLDQYKGEK
GSEG---EPGMPGI----SLK
PF01391 // ENSGT00900000140804_1_Domain_2 (322, 449)
GEEGVMGFPGARGFPGL---DGEKGASGQ----------
---------------------------------------------------------KGS
RGQDGFQGPS-GPRGPKGERGERGPPGAPAYSPHP-------------------------
-----------------------------------------SLAKGSRGDPGFPGTHGEP
GSRGEPGDPGPIGP---P--------------------------GLSIGDGD-DKRGLPG
EMGPKGYIGEPGLPALYPGPPGADGKPGLPGVPGPAGPPGPD------------------
-GYLFGLKGVEGRVGYPGPSGFPGTRGQKGWKG---------------------------
-----------------------EAGDC-KCA---DGLG
PF01391 // ENSGT00900000140804_1_Domain_6 (760, 1015)
GLPGLPGTKGFPGISGTPGRK
GDQGDPGQHGIPGFPGFKGAPGHVGV----------------------------------
------------------------------------------------------------
------------------------------------------------------------
-----------------------------------------PGP---KG--VKGDSRTIT
TKGERGQPGVPGVHGMKGDDGIPGRDGLDGFPGL-----PGP-PGDGI
PF01391 // ENSGT00900000140804_1_Domain_9 (1069, 1127)
KGLPGDSGYPGL
PGTKGFPGEAGPPGLGLPGPKGERGFPGDSGLPGPPGFPGPPGPPGTPGQIDCD---TGV
KRPIGGDQQEIV--PAGCA--GGPK-----------------------GLPGQPGPP---
G----------------SPGVKGIRGTPGFSGADGAPGLKGFPGEPGREGFPGPP-GFMG
PRGSKGATGRPGPDGLPGPIGLPGPAGPPGDRGIPGEVLGAQPGPRGDAGLPGHPGLKGV
PGERGPPGFRGS-----QGMPGMPGLKGQLGFPGPSGQPGLPGPPGQYGF-PGAPGREGP
LGLPGAP-GL--------------------GGLPGDR
PF01391 // ENSGT00900000140804_1_Domain_12 (1418, 1482)
GQPGDTGVPGPVGMKGLSGDRGD
AGVPGERGHPGSPGFKGMEGTPGVPGKK-------GERGSPGMDGFQGMVGLKGRPGIPG
FKGESGFFGI----PGLKGPAGEPGVKGNRGEPGPPGPPPIILPG-MKDIKGEKGDEGPM
GLKGYFGLK-------------GTQGMPGVPGLAGIP-GLPGRPGYIKGV-KGDIGVPGV
PGLPGFPGVAGPPGIIGFPGFTGSRGDKGAPGR---------------------------
---AGLYGETGPTGDF---GDIGDTIDLPGSPGLKGERGITGTP-GVKGFFGEKGAEGEV
GFPGITGLAGAQGTPGLKGQTGF
PF01391 // ENSGT00900000140804_1_Domain_17 (1764, 1854)
PGLTGLRGPQGEPGRTGAPG--DKGDYGWPGAPGLP-
--------------------------------GFPGIRGISGLHGLPGTKGFSGSPGADI
HG----------D---PGFPGPIGERGDPGEPNTLPGPVGVPGRKGEQGAP---------
--------------------------GERGPVGTPGLQGFPGV-SPPSNISGAPGDIGS-
---PGIFGPDGYQGPAGPPGPAALPGSKGDEGSPGAPGSPGVKGWVGYP
PF01391 // ENSGT00900000140804_1_Domain_20 (2030, 2108)
GPQGRPGVFGL
PGEKG---------ARGDPGFMGNTGAT-----GAVGDRGPKGPKGDQGFPG-------A
PGSMGSPGIAGIPQKIAAQPGSMGPQGRRGLPGAQGEMGPQGPPGDP-------------
---------------------------------------GLRGAPGKAGPQGRGGVSAVP
GFRGDQGPMGHQGPIGQE----GEPGRPGSPGLPGMPGRSVSIGY
PF01413 // ENSGT00900000140804_1_Domain_23 (2266, 2371)
LLVKHSQTDQEPMCP
VGMNKLWSGYSLLYFEGQEKAHNQDLGLAGSCLARFSTMPFLYCNPGDVCYYASRNDKSS
WLSSTAPLPMMPVAEEE--IKPYISRCSVCENSGT00900000140804_1_Linker_23_25 (2371, 2376)
EAPAMPF01413 // ENSGT00900000140804_1_Domain_25 (2376, 2504)
AIAVHSQ--DVSIPHCPAGWRSLWI
GYSFLMHTAAGDEGGGQS----LVSPGSCLEDFRAT---PFIECNG-GRGTCHYFANKYS
--FWLTTIPEQSFQGSPSA-----DTLKAGLIRTHISRCQVCMKNL--------------
--------------
ENSCCAP00000035349 (view gene)
--------------MGRNQ-RAAAGPALPRWLLLGTVTVGFLSQSVLAGVKKSDVPCGGR
DCS-GGCQCFPEKGGR
PF01391 // ENSGT00900000140804_1_Domain_0 (77, 134)
GQPGPVGPQGYTGPPGLQGFPGLQGRKGDKGERGAPGITGPKGD
VGARGVSGFPGADGIP---GHPGQGGPRGRPGYDGCNGTQGDSGPQGPPGSEGFTGPPGP
QGPKGQKGEPYALPKEERDRYR
PF01391 // ENSGT00900000140804_1_Domain_1 (203, 251)
GEPGEPGLVGFQGPPGRPGPVGQMGPLGPPGRPGPPGP
PGPKGQQGNRENSGT00900000140804_1_Linker_1_2 (251, 322)
GLGF--YGVKGEKGDVGQPGPNGIPSE--IHPVIAPTGATYQPDQDKGEK
GS---EGEPGIKGI----SLKPF01391 // ENSGT00900000140804_1_Domain_2 (322, 447)
GEEGIMGFPGPRGYPGL---SGEKGSPGQ----------
---------------------------------------------------------KGS
RGLDGYQGSD-GPRGPKGETGDPGPPGLPAYSPHP-------------------------
-----------------------------------------SLAKGARGDPGFPGARGEP
GSQGEPGDPGPLGP---P--------------------------GISIGAED-LGRGLPG
EMGPKGFIGDPGIPALYP
PF01391 // ENSGT00900000140804_1_Domain_5 (619, 747)
GPPGLDGKPGLPGPRGLPGPPGPD------------------
-GFLFGLKGAEGRVGFPGLPGSPGARGQKGRKG---------------------------
-----------------------DAGENSGT00900000140804_1_Linker_5_6 (747, 760)
DC-RCAEGDEPVRPF01391 // ENSGT00900000140804_1_Domain_6 (760, 1015)
GLPGLPGPKGFPGINGEPGRK
GDKGDPGQHGLPGFPGLKGVPGNVGA----------------------------------
------------------------------------------------------------
------------------------------------------------------------
-----------------------------------------TGP---KG--AKGDSRTIT
TKGERGQPGVPGVPGMKGDDGIPGREGLDGFPGL-----PGP-PGDGIKGPPGDPGYPGI
PGTKGSPGEMGPPGLGLPGLKGQHGFPGDAGLPGPPGFRGPPGPPGTPGQTDCD---ADV
KRPIAGDRQEAV--QPGCA--GGPK-----------------------GLPGLPGPP---
G----------------PTGAKGLRGTPGFSGADGGPGPKGLPGDPGREGFPGPP-GFIG
PRGSKGAAGLPGPDGLPGPVGLPGPDGAPGDKGLPGEVLGAQPGPRGDAGVPGHPGLKGL
PGDRGSPGFRGS-----QGMPGMPGLKGQLGFPGPSGQPGLSGPPGQHGF-PGAPGPEGP
LGLPGTL-GL--------------------EGLPGDRGEPGDIGVPGPVGMKGVSGDRGD
TGLAGERGHPGSPGFKGIDGMPGAPGLK-------
PF01391 // ENSGT00900000140804_1_Domain_13 (1476, 1538)
GERGSPGMDGFQGMPGLKGRPGFPG
SKGEAGFFGI----PGLKGLAGEPGFKGSRGDPGPPGPPPKFLPG-MKDIKGDKGDEGPM
GLKGYLGAK-------------GIQGMPGIPGLSGIP-GLPGRPGHIKGV-KGDIGTPGI
PGLPGFPGVTGPPGITGFPGFTGIRGDKGAPGR---------------------------
---AGLYGEIGRTGDF---GDIGDTINLPGRPGLKGERGVTGIP-GLKGFFGEKGTEGDI
GFPGITGVTGVQGPPGLKGQAGF
PF01391 // ENSGT00900000140804_1_Domain_17 (1764, 1855)
PGLTGPQGPQGEPGRIGLPG--GKGDDGWPGAPGLP-
--------------------------------GFPGPRGISGLHGLPGTKGFPGSPGADI
HG----------D---PGFPGPPGERGDPGEANTLPGPVGVPGQKGEQGAP---------
--------------------------GQRGPPGNPGLQGFPGI-TPPSNIS
PF01391 // ENSGT00900000140804_1_Domain_19 (1972, 2034)
GAPGDKGA-
---PGIFGLKGYRGPPGPPGSAALPGSQGDTGNPGAPGTPGTKGWVGDPGPQGRPGVLGL
PGEKG---------PRGEQGFMGNPGPP-----GPVGDRGPKGPKGDPGFPG-------A
PGIVGAPGIAGIPQRIATQPGTVGPQGRRGPPGAPGEMGPQGPPGEP-------------
---------------------------------------GFRGAPGKAGPQGRGGVSAVP
GFRGDEGPTGHQGPVGQE----GVPGRPGSPGLPGMPGRSVSIGY
PF01413 // ENSGT00900000140804_1_Domain_23 (2266, 2371)
LLVKHSQTDQEPMCP
VGMNKLWSGYSLLYFEGQEKAHNQDLGLAGSCLARFSTMPFLYCNPGDVCYYASRNDKSY
WLSTTAPLPMMPVAEDE--IKPYISRCSVCENSGT00900000140804_1_Linker_23_25 (2371, 2376)
EAPAVPF01413 // ENSGT00900000140804_1_Domain_25 (2376, 2504)
AIAVHSQ--DVSIPHCPAGWRSLWI
GYSFLMHTAAGDEGGGQS----LVSPGSCLEDFRAT---PFIECNG-GRGTCHYYANKYS
--FWLTTIPEQSFQGSPSA-----DTLKAGLIRTHISRCQVCMKNL--------------
--------------
MGP_CAROLIEiJ_P0083735 (view gene)
--------------MDRVR-FKASGPPLRGWLLLATVTVGLMAQSVLGGVKKLDVPCGGR
DCS-GGCQCYPEKGARGQPGAVGPQGYSGPPGLQGFPGLQGRKGDKGERGVPGPTGPKGD
VGARGVSGFP
PF01391 // ENSGT00900000140804_1_Domain_0 (131, 189)
GADGIP---GHPGQGGPRGRPGYDGCNGTRGDAGPQGPSGSGGFPGLPGP
QGPKGQKGENSGT00900000140804_1_Linker_0_1 (189, 203)
EPYALSKEDRDKYRPF01391 // ENSGT00900000140804_1_Domain_1 (203, 251)
GEPGEPGLVGYQGPPGRPGPIGQMGPMGAPGRPGPPGP
PGPKGQPGNRENSGT00900000140804_1_Linker_1_2 (251, 321)
GLGF--YGQKGEKGDIGQPGPNGIPSDI---TLVGPTTSTIHPDLYKGEK
GDEG---EQGIPGV----ISPF01391 // ENSGT00900000140804_1_Domain_2 (321, 448)
KGEEGIMGFPGIRGFPGL---DGEKGVLGQ----------
---------------------------------------------------------KGS
RGLDGFQGPS-GPRGPKGERGEQGPPGPSVYSPHP-------------------------
-----------------------------------------SLAKGARGDPGFQGAHGEP
GSRGEPGEPGTAGP---P--------------------------GPSIGDED-SMRGLPG
EMGPKGFSGEPGPPARYPGPPGADGRPGPQGVPGPAGPPGPD------------------
-GFLFGLKGSEGRVGYPGPSGFPGTRGQKGWKG---------------------------
-----------------------EAGDC-QCG---QVIG
PF01391 // ENSGT00900000140804_1_Domain_6 (760, 1014)
GLPGLPGPKGFPGVNGELGKK
GDQGDPGLHGIPGFPGFKGAPGIAGA----------------------------------
------------------------------------------------------------
------------------------------------------------------------
-----------------------------------------PGP---KG--IKGDSRTVT
TKGERGQPGIPGVHGMKGDDGVPGRDGLDGFPGL-----PGP-PGDGIKGPPGDAGLPGV
PGTKGFPGDVGPPGQGLPGPKGERGFPGDAGLPGPPGFPGPPGPPGTPGQRDCD---TGV
KRPIGGGQQVIV--QPGCI--EGPT-----------------------GSPGQPGPP---
G----------------PTGAKGIRGMPGFPGASGEQGLKGFPGDPGREGFPGPP-GFMG
PRGSKGTTGLPGPDGPPGPIGLPGPAGPPGDRGIPGEVLGAQPGTRGDAGLPGQPGLKGL
PGETGAPGFRGS-----QGMPGMPGLKGQPGFPGPSGQPGQSGPPGQHGF-PGTPGREGP
LGQPGSP-GL--------------------GGLPGDRGEPGDPGVPGPVGMKGLSGDRGD
AGMSGERGHPGSPGFKGMAGMPGIPGQK-------GDR
PF01391 // ENSGT00900000140804_1_Domain_13 (1479, 1539)
GSPGMDGFQGMLGLKGRPGFPG
TKGEAGFFGV----PGLKGLPGEPGVKGNRGDRGPPGPPPLILPG-MKDIKGEKGDEGPM
GLKGYLGLK-------------GIQGMPGVPGVSGFP-GLPGRPGFIKGV-K
PF01391 // ENSGT00900000140804_1_Domain_15 (1613, 1701)
GDIGVPGT
PGLPGFPGVSGPPGITGFPGFTGSRGEKGTPGV---------------------------
---AGVFGETGPTGDF---GDIGDTVDLPGSPGLKGERGITGIP-GLKGFFGEKGAEGDI
GFPGITGMAGAQGSPGLKGQTGFPGLTGLQGPQGEPGRVGIPG--DKGDFGWPGVPGLP-
--------------------------------GFPGIRGISGLHGLPGTKGFPGSPGVDA
NG----------D---PGYPGPTGDRGDRGEANTLPGPVGVPGQKGERGTP---------
--------------------------GERGPAGSPGLQGFPGI-SPPSNISGSPGDVGA-
---PGIFGLQGYQGPPGPPGPNALPGIKGDEGSSGAAGFPGEKGWVGDPGPQ
PF01391 // ENSGT00900000140804_1_Domain_20 (2033, 2111)
GQPGVLGL
PGEKG---------PKGEQGFMGNTGPS-----GAVGDRGPKGPKGDQGFPG-------A
PGSMGSPGIPGIPQKIAAQPGTLGPQGRRGLPGALGEIGPQGPPGDP-------------
---------------------------------------GFRGAPGKAGPQGRGGVSAVP
GFRGDQGPMGHQGPVGQE----GEPGRPGSPGLPGMPGRSVSIGY
PF01413 // ENSGT00900000140804_1_Domain_23 (2266, 2371)
LLVKHSQTDQEPMCP
VGMNKLWSGYSLLYFEGQEKAHNQDLGLAGSCLARFSTMPFLYCNPGDVCYYASRNDKSY
WLSTTAPLPMMPVAEEE--IKPYISRCSVCENSGT00900000140804_1_Linker_23_25 (2371, 2376)
EAPAVPF01413 // ENSGT00900000140804_1_Domain_25 (2376, 2504)
AIAVHSQ--DTSIPHCPAGWRSLWI
GYSFLMHTAAGDEGGGQS----LVSPGSCLEDFRAT---PFIECNG-GRGTCHYFANKYS
--FWLTTIPEQNFQSTPSA-----DTLKAGLIRTHISRCQVCMKNL--------------
--------------
ENSMPUP00000010555 (view gene)
--------------MDRHQ-CSASSPALRRWLLLGAVTVGLLAQSVLAGVKKFDAPCGGR
DCS-GGCQCYPEKGGRGQPGPVGPQGYTGPPGLQGFPGLQGRKGDKGERGSPGTTGPKGD
VGARGVSGFP
PF01391 // ENSGT00900000140804_1_Domain_0 (131, 190)
GADGIP---GHPGQGGPRGPPGHDGCNGTRGDVGQQGPSGPPGFHGPPGP
QGPKGQKGEENSGT00900000140804_1_Linker_0_1 (190, 203)
PYALSSAERDRYRPF01391 // ENSGT00900000140804_1_Domain_1 (203, 247)
GEPGEPGLVGFQGSPGRPGPVGQMGPVGAPGRPGPPGP
PGPKGQPRLSGHGRGRAGCCGEKGDIGVPGPNGIVWDF--HPVAGPTKESFQPELYKGEK
GS---AGEPGVKGV----SLKGEEGVMGFSGPRGHMRLR--PNALSPAPL----------
---------------------------------------------------------LGS
RGPNGYQGPS-GPPGPKVSSVSLWLASRPAPNPHP-------------------------
-----------------------------------------VLVA
PF01391 // ENSGT00900000140804_1_Domain_3 (526, 613)
GARGDPGFPGTHGEP
GSRGEPGDPGPPGL---P--------------------------GTSVRDED-GKRGLPG
EMGPKGFIGDPGENSGT00900000140804_1_Linker_3_5 (613, 619)
VPALYPPF01391 // ENSGT00900000140804_1_Domain_5 (619, 747)
GTPGTDGRPGLQGPPGPAGPPGPD------------------
-GFLFGLKGTEGQAGFPGPSGFPGARGQKGWKG---------------------------
-----------------------DAGENSGT00900000140804_1_Linker_5_6 (747, 760)
DC-KCAEGDQFVPPF01391 // ENSGT00900000140804_1_Domain_6 (760, 1015)
GLPGFPGPKGLPGTNGEPGRK
GDQGDPGQHGIPGFPGFKGAPGHMGP----------------------------------
------------------------------------------------------------
------------------------------------------------------------
-----------------------------------------PGP---KG--LKGDSRAVT
TKGERGHPGVPGVPGLKGDDGVPGRDGLDGFPGL-----PGP-PGDGIKGPPGDSGYPGA
PGTKGAAGDRGPQGLGLPGPKGERGFPGDAGLPGPPGFPGPPGPPGTPGQIDCD---TGV
KRPIGGDGQETV--QPGCV--GGPK-----------------------GSPGLPGPPGPT
G----------------YLGAKGLQGTPGLSGADGAPGLKGLPGDPGREGFPGPP-GFVG
PRGSKGTAGLPGADGLPGPSGLPGPIGPPGERGLPGEVLGAQ
PF01391 // ENSGT00900000140804_1_Domain_11 (1303, 1364)
PGPRGDAGVPGQPGLRGP
PGERGPPGFRGS-----QGMPGMPGLKGQPGFPGPSGQPGLPGPPGQHGF-PGAPGQEGP
LGRPGAP-GH--------------------GGLPGDRGDPGDTGVPGPMGMKGLSGDRGD
PGLLGERGPRGSPGFKGMSGMPGVPGQK-------GGRGSPGMDGFQGMVGLKGRPGLPG
NKGEAGFYGV----PGLKGLAGEPGVKGSRGDPGPPGPPPIIQPG-MKDIKGEKGDEGPM
GLKGYLGLK-------------GVPGMPGVPGLSGIP-GLPGKPGYIKGV-KGDTGVPGV
PGLPGFPGVPGPPGIIGFPGFTGSRGDKGAPGR---------------------------
---AGLYGEVGPTGDF---GDIGDTINLP
PF01391 // ENSGT00900000140804_1_Domain_17 (1710, 1765)
GRPGLKGERGVAGAP-GLKGFFGEKGTVGEV
GFPGITGMPGVQGPPGAKGQTGFPGLTGLQGPQGEPGRAGRPG--DKGDFGWPGVPGSP-
--------------------------------GFPGIHGVSGLHGLPGTKGFPGSPGVDI
HG----------D---PGFPGPAGDRGDPGEPNTRPGPFGAPGQKGERGDP---------
--------------------------GERGPIGSPGRQGFPGV-TPPSNVSG
PF01391 // ENSGT00900000140804_1_Domain_19 (1973, 2034)
LPGDKGA-
---PGIFGLEGYRGPPGPPGPAALPGSKGDEGNPGAPGNPGTKGWGGDPGPQGRPGVFGL
PGEKG---------PRGDPGFMGNTGAT-----GSVGDRGPKGPKGDRGLPG-------A
PGSVGAPGIAGIPQRIAIQ
PF01391 // ENSGT00900000140804_1_Domain_21 (2120, 2220)
PGPVGPQGRRGPPGAQGEMGPQGPPGEP-------------
---------------------------------------GFRGDPGKAGPQGRGGVSAVENSGT00900000140804_1_Linker_21_23 (2220, 2265)
P
GFRGDQGPMGQQGPVGQD---PGEPGRPGTPGLPGMPGRSVSIGPF01413 // ENSGT00900000140804_1_Domain_23 (2265, 2372)
YLLVKHSQTDREPMCP
VGMNKLWGGYSLLYFEGQEKAHNQDLGLAGSCLARFSTMPFLYCNPGDVCYYASRNDKSY
WLSTTAPLPMMPVAEDD--IKPYISRCSVCEENSGT00900000140804_1_Linker_23_25 (2372, 2375)
APAPF01413 // ENSGT00900000140804_1_Domain_25 (2375, 2505)
VAIAVHSQ--DVSIPHCPAGWRSLWI
GYSFLMHTAAGDEGGGQS----LVSPGSCLEDFRAT---PFIECNG-GRGTCHYYANKYS
--FWLTTIPEPSFQGAPSA-----DTLKAGLIRTHISRCQVCMKNL--------------
--------------
ENSOANP00000024611 (view gene)
------------------------------------------------------------
---------------QGQPGPVGLQGSPGPSGLHGIPGLQGRKGDKGERGPPGISGPRGD
PGQRGVP
PF01391 // ENSGT00900000140804_1_Domain_0 (128, 189)
GFRGADGIP---GHPGQGGPRGRPGYDGCNGTIGDPGQPGTPGIYGSLGFPGP
QGPKGQKGENSGT00900000140804_1_Linker_0_1 (189, 203)
EPFALPWEEQNKYRPF01391 // ENSGT00900000140804_1_Domain_1 (203, 251)
GEPGEPGLVGMPGSPGRPGVEGPIGPVGAPGQPGPPGP
PGPKGLQGNRGLGF--YGQKGDKGEIGHPGPTGLPSENEHQ----STIEIVNLEQYKGEK
GRE---GFPVSEG----TPGRGEGGIMGFAGQRGFPGH---SGPPGHDGN----------
---------------------------------------------------------KGI
FGLNSIRAGS-GSVGPKGEMGDPGLPGPLPPYPPP-------------------------
-----------------------------------------HFRKGAPGDPGFPGLKGPS
GDPGDQGYPGPPGY---P--------------------------GSSDGYKD-NRTGWPG
EMGLKGDPGEPGIPAAFP
PF01391 // ENSGT00900000140804_1_Domain_5 (619, 747)
GPPGINGNIGLPGLPGPAGPRGVT------------------
-DIFHGIKGSKGMPGPKGSQGFPGLKGEKGLKG---------------------------
-----------------------EPGENSGT00900000140804_1_Linker_5_6 (747, 760)
EC-KCLDED--GEPF01391 // ENSGT00900000140804_1_Domain_6 (760, 1014)
GLPGFQGPPGIPGDQGEPGRK
GDSGDPGQYGTPGQQGLKGSQGNEGE----------------------------------
------------------------------------------------------------
------------------------------------------------------------
-----------------------------------------IGL---KG--EKENSGT00900000140804_1_Linker_6_7 (1014, 1023)
GDLQTVT
TKPF01391 // ENSGT00900000140804_1_Domain_7 (1023, 1085)
GAPGDQGIPGNSGPRGADGPPGQDGREGVLGI-----PGPLGADGVKGLSGDTGLQGE
LGPKGFPGIVGPPGFGFPGPKGARGSPGHSGYWGSPGHPGPRGPPGLPGRTDCE---DGVKKPIG-DGQEGIW--PGCV--TPPK-----------------------GPPGPPGPP---G----------------LQGSKGIKGHPGKTGADGYVGLKGLPGEPGREGIPGPP-GFIGPRGQKGAPGFPGPKGSPGAKGSPGHPGLAGEKGRPGEVLSAQPGLPGDSGLPGIPGLKGQPGSIGPPGFRGG-----QGMPGMPGLKGRLGFPGPSGQRGPPGLTGTHGF-PGFPGHPGPPGLPGLL-GF--------------------KGHPGDIGDRGDPGFPGPIGMKGLAGDPGENGFVGMRGAQGSPGFQGMDGMPGNPGSK-------GPKGSPGMDGFKGMAGLKGRLGMPGYKGVTGNLGL----PGLKGLAGDRGFKGSRGDPGPPGEPPKLIPG-MKHFKGEKGDEGPTGLKGFLGLK-------------GGQGMPGVPGASGVP-GLPGQPSQVKGD-KGETGVRGHPGEQGYPGAAGIPGIRGFPGFPGIRGDKGNIGI------------------------------KGFEGESGSVGDL---GDKGDTINLPGSRGVKGAPGLAGVS-GVKGYVGQKGHEGDSGLQGISGMKGTQGPPGIRGQSGSPGMAGSPGHHGDPGRNGFQG--EKGDTGRPGLPGFS--------------------------------AGVSGLRGISGLDGLPGTKGFPGSPGGDDFG----------D---PGFLGPSGDKGEPGDHNPKPGPSGAVGPKGERGLT-----------------------------------GIRGPMGSRGLPGIPGP-FSVTNISGFPGDRGD----PGIFGTKGYPGQRGPPGVPASPGPKGEDGNPGLFGESGPKGWGGDPGPQGKPGVYGIPGTKG---------PKGEQGLMGFKGMF-----GNIGDQGPVGPKGDRGYPG-------TLGPLGSPGIPGIPQEVVLDRGIPGPFGKQGPPGIQGEMGPQGPPGHP----------------------------------------------------GFRGSPGNPGPQGRGGVSAVPGFRGDQGPVGHQGATGQE----GNPGRPGNPGMPGMPGRSVSIGYLLVKHSQTEQEPMCPLGMNKLWSGYSLLYFEGQEKAHNQDLGLAGSCLARFSTMPFLYCNPGDVCYYASRNDKSYWLSTTAPLPMMPVAEDE--IKPYISRCSVCEAPAVAIAVHSQ--DVTIPHCPNGWRSLWIGYSFLMHTAAGDEGGGQS----LVSPGSCLEDFRAT---PFIECNG-GRGNCHYFANKYS--FWLTTIPEPNFQSSPSP-----DTLKAGLIRTHISRCQVCMKNL----------------
PF01391 // ENSGT00900000140804_1_Domain_9 (1073, 1130)
GDTGLQGELGPKGFPGIVGPPGFGFPGPKGARGSPGHSGYWGSPGHPGPRGPPGLPGRTDCE---DGV
KKPIG-DGQEGIW--PGCV--TPPK-----------------------GPPGPPGPP---
G----------------LQGSKGIKGHPGKTGADGYVGLKGLPGEPGREGIPGPP-GFIG
PRGQKGAPGFPGPKGSPGAKGSPGHPGLAGEKGRPGEVLSA
PF01391 // ENSGT00900000140804_1_Domain_11 (1302, 1363)
QPGLPGDSGLPGIPGLKGQ
PGSIGPPGFRGG-----QGMPGMPGLKGRLGFPGPSGQRGPPGLTGTHGF-PGFPGHPGP
PGLPGLL-GF--------------------KGHPGDIGDRGDPGFPGPIGMKGLAGDPGE
NGFVGMRGAQGSPGFQGMDGMPGNPGSK-------GPKGSPGMDGFKGMAGLKGRLGMPG
YKGVTGNLGL----PGLKGLAGDRGFKGSRGDPGPPGEPPKLIPG-MKHFKGEKGDEGPT
GLKGFLGLK-------------GGQGMPGVPGASGVP-GLPGQPSQVKGD-KGETGVRGH
PGEQGYPGAAGIPGIRGFPGFPGIRGDKGNIGI---------------------------
---KGFEGESGSVGDL---GDKGDTINLP
PF01391 // ENSGT00900000140804_1_Domain_17 (1710, 1765)
GSRGVKGAPGLAGVS-GVKGYVGQKGHEGDS
GLQGISGMKGTQGPPGIRGQSGSPGMAGSPGHHGDPGRNGFQG--EKGDTGRPGLPGFS-
-------------------------------AGVSGLRGISGLDGLPGTKGFPGSPGGDD
FG----------D---PGFLGPSGDKGEPGDHNPKPGPSGAVGPKGERGLT---------
--------------------------GIRGPMGSRGLPGIPGP-FSVTNISGFPGDRGD-
---PGIFGTKGYPGQRGPPGVPASPGPKGEDGNPGLFGESGPKGWGGDPGPQGKPGVYGI
PGTKG---------PKGEQGLMGFKGMF-----GNIGDQGPVGPKGDRGYPG-------T
LGPLGSPGIPGIPQEVVLDRGIPGPFGKQGPPGIQGEMGPQGPPGHP-------------
---------------------------------------GFR
PF01391 // ENSGT00900000140804_1_Domain_22 (2203, 2259)
GSPGNPGPQGRGGVSAVP
GFRGDQGPVGHQGATGQE----GNPGRPGNPGMPGMPGENSGT00900000140804_1_Linker_22_23 (2259, 2265)
RSVSIGPF01413 // ENSGT00900000140804_1_Domain_23 (2265, 2372)
YLLVKHSQTEQEPMCP
LGMNKLWSGYSLLYFEGQEKAHNQDLGLAGSCLARFSTMPFLYCNPGDVCYYASRNDKSY
WLSTTAPLPMMPVAEDE--IKPYISRCSVCEENSGT00900000140804_1_Linker_23_25 (2372, 2375)
APAPF01413 // ENSGT00900000140804_1_Domain_25 (2375, 2505)
VAIAVHSQ--DVTIPHCPNGWRSLWI
GYSFLMHTAAGDEGGGQS----LVSPGSCLEDFRAT---PFIECNG-GRGNCHYFANKYS
--FWLTTIPEPNFQSSPSP-----DTLKAGLIRTHISRCQVCMKNL--------------
--------------
ENSHGLP00000002972 (view gene)
--------------------ANSPGPVLRRWLLLGAVTVGLLAHSVLAGVKKLDVPCGGR
DCS-GGCRCYPEKGGR
PF01391 // ENSGT00900000140804_1_Domain_0 (77, 133)
GLPGPAGLPGFHGPPGLQGFPGLQGRKGDKGERGAPGTPGPKGD
VGPRGVSGFPGANGIP---GYPGQGGPRGRPGYDGCNGTRGDTGPQGPSGSEGFPGTFGP
PGPKGQKGEPYALSREDQDRYRGEPGEPGLVGFQGPPGRPGPMGPMGPVGAPGR------
------PGNRGLGF--YGEKGEKGDQGQPGPNGIPSDT-THPIVAPTGSTIHPDMYKGEK
GS---QGEPGFKGI----SYK
PF01391 // ENSGT00900000140804_1_Domain_2 (322, 448)
GEEGIMGFPGTRGPPGL---DGERGFPGQ----------
---------------------------------------------------------KGS
KGRDGYQGPD-GPQGSKGELGEPGPPGPPAYSPHP-------------------------
-----------------------------------------SLAKGVRGDPGFPGAHGEP
GSRGEPGDPGLPGPQGPP--------------------------GLPSVDQG-DRTGRPG
ETGLKGFTGDRGIPARYP
PF01391 // ENSGT00900000140804_1_Domain_5 (619, 747)
GPPGADGRPGPPGPPGPAGPPGPD------------------
-GFLFGLKGAQGSRGYAGLPGTPGVNGQKGRKG---------------------------
-----------------------DPGENSGT00900000140804_1_Linker_5_6 (747, 760)
NC-KCAESDQFPGPF01391 // ENSGT00900000140804_1_Domain_6 (760, 1015)
GLPGLPGLKGFPGANGKPGWK
GEPGDPGQHGIPGFPGFKGAPGSIGA----------------------------------
------------------------------------------------------------
------------------------------------------------------------
-----------------------------------------PGP---KG--AKGENSGT00900000140804_1_Linker_6_7 (1015, 1022)
DSRTVT
TPF01391 // ENSGT00900000140804_1_Domain_7 (1022, 1080)
KGERGQPGIPGVHGMKGDDGIPGRDGLDGFPGL-----PGP-PGDGIRGPPGDSGPPGAPGTKGPPGEVGPPGLGMLGREGEQGFPGDVGLPGSPGFPGPPGAPGSPGELDCE---TGVKSPTGGDTQEPV--QPGCT--GGPK-----------------------GSPGLPGPP---G----------------PPGDKGPRGTPGFLGVDGPPGLKGFPGDPGREGFPGPP-GFMGPRGSKGAVGLPGPDGLQGTSGLPGPAGPPGDRGIPGEVLGAQPGPRGDTGLPGHPGRKGLPGEQGAPGFRGS-----QGMPGMPGPKGQLGFPGPSGLPGLPGRAGQHGF-PGAPGPEGPLGLPGVS-GS--------------------EGLPGDRGDPGDTGTPGPVGMKGLAGDRGDTGLVGDQGRPGDPGFKGMDGMPGIPG-----------------------------------------------------------LKGNRGEPGPPGPPPAIMPG-MKFIKGEKGDEGPMGLKGHLGLK-------------GLQGMPGVPGQAGVP-GLPGRPGHIKGI-KGDIGVPGSPGLPGFPGVTGPPGVIGFPGFTGSRGDKGAPGS------------------------------SGLYGDDGLTGDF---GDIGDTIDLPGSPGLKGERGIAGTP-GLKGFFGERGAHGEVGFPGITGLAGPQGQPGLKGQT-----------------------------------------------------------------------------------------------GADMPG----------D---PGFPGPSGDRGDPGEPNTFPGSTGAPGQKGEQGAP-----------------------------------GERGPAGRPGMQGFPGI-SPPSNISGLPGDVGS----PGIFGLKGYQGPQGPPGPAALQGSKGDAGSGGAPGVVGTKGWIGDQGPQGRPGVFGVPGEKG---------PRGEPGLMGSTGST-----GAVGDRGPKGPKGDQGFPG------AP-GLVGSPGIAGIPQRITAPPGAMGPEGRRGRPGAQGEMGPQGPPGDP----------------------------------------------------GFRGVPGKPGPQGRGGVSALPGFRGDPGAVGHQGPIGQE----GEPGRPGSPGLPGMPGRSVSIGYLLVKHSQADQEPMCPVGMNKLWSGYSLLYFEGQEKAHNQDLGLAGSCLARFSTMPFLYCNPGDVCYYASRNDKSYWLSTTAPLPMMPVAEDD--IKPYISRCSVCEAPATAIAVHSQ--DVSIPHCPAGWRSLWIGYSFLMHTAAGDEGGGQS----LMSPGSCLEDFRAT---PFIECNG-GRGTCHYFANKYS--FWLTTIPDTSFQGTPSA-----DTLKAGLIRTHISRCQVCMKNL------------------
PF01391 // ENSGT00900000140804_1_Domain_9 (1070, 1130)
GPPGDSGPPGA
PGTKGPPGEVGPPGLGMLGREGEQGFPGDVGLPGSPGFPGPPGAPGSPGENSGT00900000140804_1_Linker_9_10 (1130, 1163)
ELDCE---TGV
KSPTGGDTQEPV--QPGCT--GPF01391 // ENSGT00900000140804_1_Domain_10 (1163, 1257)
GPK-----------------------GSPGLPGPP---
G----------------PPGDKGPRGTPGFLGVDGPPGLKGFPGDPGREGFPGPP-GFMG
PRGSKGAVGLPGPDGLQGTSGLPGPAGPPGDRGIPGEVLGAQPGPRGDTGLPGHPGRKGL
PGEQGAPGFRGS-----QGMPGMPGPKGQLGFPGPSGLPGLPGRAGQHGF-PGAPGPEGP
LGLPGVS-GS--------------------EGLPGDRGDPGDTGTPGPVGMKGLAGDRGD
TGLVGDQGRPGDPGFKGMDGMPGIPG----------------------------------
-------------------------LKGNRGEPGPPGPPPAIMPG-MKFIKGEKGDEGPM
GLKGHLGLK-------------GLQGMPGVPGQAGVP-GLPGRPGHIKGI-KGDIGVPGS
PGLPGFPGVTGPPGVIGFPGFTGSRGDKGAPGS---------------------------
---SGLYGDDGLTGDF---GDIGDTIDLP
PF01391 // ENSGT00900000140804_1_Domain_17 (1710, 1858)
GSPGLKGERGIAGTP-GLKGFFGERGAHGEV
GFPGITGLAGPQGQPGLKGQT---------------------------------------
--------------------------------------------------------GADM
PG----------D---PGFPGPSGDRGDPGEPNTFPGSTGAPGQKGEQGAP---------
--------------------------GERGPAGRPGMQGFPGI-SPPSNISGLP
PF01391 // ENSGT00900000140804_1_Domain_19 (1975, 2036)
GDVGS-
---PGIFGLKGYQGPQGPPGPAALQGSKGDAGSGGAPGVVGTKGWIGDQGPQGRPGVFGV
PGEKG---------PRGEPGLMGSTGST-----GAVGDRGPKGPKGDQGFPG------AP
-GLVGSPGIAGIPQRITAPPGAMGPEGRRGRPGAQGEMGPQGPPGDP-------------
---------------------------------------GFRGVPGKPGPQGRGGVSALP
GFRGDPGAVGHQGPIGQE----GEPGRPGSPGLPGMPGRSVSIGY
PF01413 // ENSGT00900000140804_1_Domain_23 (2266, 2371)
LLVKHSQADQEPMCP
VGMNKLWSGYSLLYFEGQEKAHNQDLGLAGSCLARFSTMPFLYCNPGDVCYYASRNDKSY
WLSTTAPLPMMPVAEDD--IKPYISRCSVCENSGT00900000140804_1_Linker_23_25 (2371, 2376)
EAPATPF01413 // ENSGT00900000140804_1_Domain_25 (2376, 2504)
AIAVHSQ--DVSIPHCPAGWRSLWI
GYSFLMHTAAGDEGGGQS----LMSPGSCLEDFRAT---PFIECNG-GRGTCHYFANKYS
--FWLTTIPDTSFQGTPSA-----DTLKAGLIRTHISRCQVCMKNL--------------
--------------
ENSPVAP00000005802 (view gene)
ENSBTAP00000005916 (view gene)
------------------------------------------------------------
---------------QGQPGPVGPQGYTGPPGLQGFPGLQGRKGDKGQRGAPGITGPKGD
VGPRGVSGF
PF01391 // ENSGT00900000140804_1_Domain_0 (130, 189)
PGADGIP---GHPGQGGPRGPPGYDGCNGTVGDSGYAGPPGPGGFLGPRGP
QGPKGQKGENSGT00900000140804_1_Linker_0_1 (189, 203)
EPYALSSEDRDKYRPF01391 // ENSGT00900000140804_1_Domain_1 (203, 251)
GEPGEPGLVGLQGPPGRPGPVGQMGPVGAPGRPGPPGP
PGPKGQPGNRENSGT00900000140804_1_Linker_1_2 (251, 322)
GLGF--YGEKGEKGDMGLQGPGGIPPDN-GYVEKPTPVYELLPEQYKGEK
GS---QGEPGRIGV----SLKPF01391 // ENSGT00900000140804_1_Domain_2 (322, 448)
GEEGVVGFSGPRGAPGI---DSEKGLPGQ----------
---------------------------------------------------------KGS
RGLDGYQGPD-GLPGPKGERGDPGPPGENSGT00900000140804_1_Linker_2_3 (448, 526)
PPAYSPYP-------------------------
-----------------------------------------SVAKPF01391 // ENSGT00900000140804_1_Domain_3 (526, 612)
GVRGEPGSPGAPGEP
GDRGVPGDPGLPGR---P--------------------------GTTIGDED-EKRGLPG
EMGPKGFTGEQGFPALYPGPPGTDGQPGLRGPPGLPGPPGPD-------------------GFLFGLKGEKGSGGFPGSSGFPGARGQKGWKG--------------------------------------------------DAGDC-KC--ADQFIRGPPGLPGPKGFAGANGQPGSKGSQGDPGQHGVPGFAGFKGAPGNVGP---------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------PGP---KG--MKGDSRTITTKGERGQPGVPGAPGLKGSDGIPGPPGLDGFHGL-----PGP-PGDGIKGPRGDTGQPGAPGTKGLPGG-GPPGLGLPGPKGERGFPGDAGLPGPPGFPGPPGLPGSPGQIDCD---SGVKRPIGADGQETI--QPGCV--GGPK-----------------------GSPGQPGLP---G----------------PPGAKGLRGIPGFSGADGAPGLKGLPGDPGREGFPGPP-GFMGPRGSKGAVGPPGLDGLPGASGLPGPVGPPGDRGLPGEVLGAQPGPRGDSGLPGRPGPKGPPGERGPPGFRGS-----QGMPGMPGQKGQPGSPGLSGQPGLPGPPGQHGF-PGAPGREGPLGPPGAP-GF--------------------QGLPGDRGEPGDTGVPGPVGMKGGSGDRGDPGQQGERGHPGHPGFKGVSGMPGAPGLK-------GARGSPGMDGFEGMLGLKGRPGLPGIKGEAGFFGI----PGLKGLAGEPGVKGSRGDPGPPGPPPLIEPG-MKDIKGEKGDEGPMGLKGYLGLK-------------GLPGMPGIPGLSGIP-GLPGRPGQIKGV-KGDTGIPGVPGSPGFPGVPGSPGIMGFQGFTGSRGDKGVPGR------------------------------AGLFGEVGPTGDF---GDIGDTIDLPGSPGLKGERGTTGIP-GQKGFFGERGTEGDIGFPGITGLAGVQGPPGFKGQKGFPGLTGLQGPQGDPGRAGAPG--IKGDSGWPGNPGLP---------------------------------GLPGLRGISGLHGLPGSKGFPGSPGADVHG----------D---PGFPGPAGDKGDPGEPNTLPGPTGAPGQKGERGAP-----------------------------------GERGPIGSPGLQGFPGI-TPLSNISGSPGDVGA----PGIFGLEGYRGPPGPPGPAALPGSKGDEGSPGTPGSPGTKGWIGDPGPQGRPGVFGLPGEKG---------PKGEPGFMGNIGPT-----GSPGDRGPKGPKGDRGLPG------AP-GAVGTPGITGIPQRIAIERGPVGPQGRRGPPGAQGEMGPQGPPGEP----------------------------------------------------GFRGAQGKAGPQGRGGVSAIPGFRGDQGPVGLQGPVGFE----GEPGRPGSPGLPGMPGRSISIGYLLVKHSQTDKEPMCPVGMNKLWSGYSLLYFEGQEKAHNQDLGLAGSCLARFSTMPFLYCNPGDVCYYASRNDKSYWLSTTAPLPMMPVAEED--IRPYISRCSVCEAPAVAIAVHSQ--DVSIPHCPAGWRSLWIGYSFLMHTAAGDEGGGQS----LVSPGSCLEDFRAT---PFIECNG-ARGTCHYYANKYS--FWLTTIPEQSFQGTPSA-----DTLKAGLIRTHISRCQVCMKNL-----------
PF01391 // ENSGT00900000140804_1_Domain_5 (595, 662)
KRGLPGEMGPKGFTGEQGFPALYPGPPGTDGQPGLRGPPGLPGPPGPD------------------
-ENSGT00900000140804_1_Linker_5_6 (662, 760)
GFLFGLKGEKGSGGFPGSSGFPGARGQKGWKG---------------------------
-----------------------DAGDC-KC--ADQFIRPF01391 // ENSGT00900000140804_1_Domain_6 (760, 1015)
GPPGLPGPKGFAGANGQPGSK
GSQGDPGQHGVPGFAGFKGAPGNVGP----------------------------------
------------------------------------------------------------
------------------------------------------------------------
-----------------------------------------PGP---KG--MKGDSRTIT
TKGERGQPGVPGAPGLKGSDGIPGPPGLDGFHGL-----PGP-PGDGIKGPRGDTGQPGA
PGTKGLPGG-GPPGLGLPGPKGERGFPGDAGLPGPPGFPGPPGLPGSPGQIDCD---SGV
KRPIGADGQETI--QPGCV--GGPK-----------------------GSPGQPGLP---
G----------------PPGAKGLRGIPGFSGADGAPGLKGLPGDPGREGFPGPP-GFMG
PRGSKGAVGPPGLDGLPGASGLPGPVGPPGDRGLPGEVLGAQPGPRGDSGLPGRPGPKGP
PGERGPPGFRGS-----QGMPGMPGQKGQPGSPGLSGQPGLPGPPGQHGF-PGAPGREGP
LGPPGAP-GF--------------------QGLPGDRGEPGDTGVPGPVGMKGGSGDRGD
PGQQGERGHPGHPGFKGVSGMPGAPGLK-------
PF01391 // ENSGT00900000140804_1_Domain_13 (1476, 1538)
GARGSPGMDGFEGMLGLKGRPGLPG
IKGEAGFFGI----PGLKGLAGEPGVKGSRGDPGPPGPPPLIEPG-MKDIKGEKGDEGPM
GLKGYLGLK-------------GLPGMPGIPGLSGIP-GLPGRPGQIK
PF01391 // ENSGT00900000140804_1_Domain_15 (1609, 1694)
GV-KGDTGIPGV
PGSPGFPGVPGSPGIMGFQGFTGSRGDKGVPGR---------------------------
---AGLFGEVGPTGDF---GDIGDTIDLPGSPGLKGERGTTGIP-GQKGFFGERGTEGDI
GFPGITGLAGVQGPPGFKGQKGFPGLTGLQGPQGDPGRAGAPG--IKGDSGWPGNPGLP-
--------------------------------GLPGLRGISGLHGLPGSKGFPGSPGADV
HG----------D---PGFPGPAGDKGDPGEPNTLPGPTGAPGQKGERGAP---------
--------------------------GERGPIGSPGLQGFPGI-TPLSNISGSPGDVGA-
---PGIFGLEGYRGPPGPPGPAALPGSKGDEGSPGTPGSPGTKGWIGDPG
PF01391 // ENSGT00900000140804_1_Domain_20 (2031, 2108)
PQGRPGVFGL
PGEKG---------PKGEPGFMGNIGPT-----GSPGDRGPKGPKGDRGLPG------AP
-GAVGTPENSGT00900000140804_1_Linker_20_21 (2108, 2120)
GITGIPQRIAIEPF01391 // ENSGT00900000140804_1_Domain_21 (2120, 2221)
RGPVGPQGRRGPPGAQGEMGPQGPPGEP-------------
---------------------------------------GFRGAQGKAGPQGRGGVSAIPENSGT00900000140804_1_Linker_21_23 (2221, 2265)
GFRGDQGPVGLQGPVGFE----GEPGRPGSPGLPGMPGRSISIGPF01413 // ENSGT00900000140804_1_Domain_23 (2265, 2372)
YLLVKHSQTDKEPMCP
VGMNKLWSGYSLLYFEGQEKAHNQDLGLAGSCLARFSTMPFLYCNPGDVCYYASRNDKSY
WLSTTAPLPMMPVAEED--IRPYISRCSVCEENSGT00900000140804_1_Linker_23_25 (2372, 2375)
APAPF01413 // ENSGT00900000140804_1_Domain_25 (2375, 2505)
VAIAVHSQ--DVSIPHCPAGWRSLWI
GYSFLMHTAAGDEGGGQS----LVSPGSCLEDFRAT---PFIECNG-ARGTCHYYANKYS
--FWLTTIPEQSFQGTPSA-----DTLKAGLIRTHISRCQVCMKNL--------------
--------------
ENSLAFP00000007563 (view gene)
ENSCHIP00000032096 (view gene)
-----------------------------RSLCLGTLTLTF-SASFLQGVKKLDVPCGGR
DCS-GGCQCYPEK
PF01391 // ENSGT00900000140804_1_Domain_0 (74, 127)
GGRGQPGPVGPQGYTGPPGLQGFPGLQGRKGDKGERGAPGITGPKGD
VGPRGVSGFPGADGIP---GHPGQGGPRGPPGYDGCNGTMGDSGYAGPPGPGGFLGPRVS
TSS---------------------TQPAPLVGFKGPLPAPSSAGVSGW------------
-QPGGWTDQRGRGV--WAAAS--------------------------ARLVVLSDLDA--
-------------------------AFRCRAPRG---------------Q----------
---------------------------------------------------------LTV
SC-------F-PVFLLQGEMGDPGPPGLPAYSPHP-------------------------
-----------------------------------------SVAKGVRGEPGSPGAPGEP
GARGEPGDPGLPGQ---P--------------------------GTTIGDED-EKRGLPG
EMGPKGFTGERGFPALYPGPPGADGKPGLRGPPGRPGPPGPD------------------
-GFLFGLKGATGSGGFPGSSGFPGARGQKGWKG---------------------------
-----------------------DAGDC-KC--ADQFIR
PF01391 // ENSGT00900000140804_1_Domain_6 (760, 1015)
GPPGLPGPKGFAGANGQPGSK
GSQGDPGPHGLPGFSGFKGAPGNVGP----------------------------------
------------------------------------------------------------
------------------------------------------------------------
-----------------------------------------PGP---KG--MKGENSGT00900000140804_1_Linker_6_7 (1015, 1023)
DSRTIT
TKPF01391 // ENSGT00900000140804_1_Domain_7 (1023, 1082)
GERGQPGVPGVPGLKGTDGIPGPPGLDGFHGL-----PGP-PGDGIKGPRGDAGQPGA
PGTKGLPGERGPPGLGLPGPKGERGFPGDAGLPGPPGFPGPPGLPGAPGQTDCD---SGVKRPIGADGQETIQ--PGTV--TTGPL--ETVYR-------------VCAHGERPGVPP-GA----------------RNRAKGLRGVPGFSGADGVPGLKGLPGDPGREGFPGPP-GFMGPRGSKGAVGPPGLDGLPGASGLPGPAGPPGDRGLPGEVLGAQPGPRGDSGLPGRPGLKGPPGERGPPGFRGS-----QGMPGMPGQKGQPGSPGFSGQPGLPGAPGQHGF-PGAPGREGPLGPPGAP-GF--------------------EGLPGDRGEPGDTGVPGPVGRKGGSGDRGDPGQQGERGHPGLPGFKGVSGMPGTPGLK-------GARGSPGMDGFEGMLGLKGRPGLPGIKGEAGFFGI----PGLKGLAGEPGVKGSRGDPGPPGPPPIILPG-MKDIKGEKGDEGPMGLKGYLGLK-------------GLPGMPGIPGLSGIP-GLPGRPGHIKGV-KGDTGVPGVPGSPGFPGVPGSPGVMGFQGFTGSRGDKGAPGR------------------------------AGLFGEVGETGDF---GDIGDTIDLPGSPGLKGERGTTGIP-GQKGFLGERGTEGDVGFPGITGLAGVQGPPGFQGQKGFPGLTGLQGPQGDPGRAGAPG--IKGDSGWPGNPGLP---------------------------------GLPGLRGISGLHGLPGNKGFPGSPGADVHG----------D---PGFPGPAGDKGDPGEANTLPGPTGAPGQKGERGAP-----------------------------------GERGPIGSPGLQGFPGI-TPPSNISGSPGDIGA----PGIFGLEGYRGPPGPPGPAALPGSKGDEGSPGTPGNPGIKGWVGDPGPQGRPGVFGLPGEKG---------PKGEPGFMGNIGPT-----GSPGDRGPRGPKGDRGLPG------AP-GAVGAPGITGIPQRIAVERGPVGPQGRRGPPGAQGEMGPQGPPGEP----------------------------------------------------GFRGAQGKAGPQGRGGVSAVPGFRGDQGPVGLQGPVGFE----GEPGRPGSPGLPGMPGRSISIGYLLVKHSQTDKEPMCPVGMNKLWSGYSLLYFEGQEKAHNQDLGLAGSCLARFSTMPFLYCNPGDICYYASRNDKSYWLSTTAPLPMMPVAEED--IRPYISRCSVCEAPAVAIAVHSQ--DVSIPHCPAGWRSLWIGYSFLMHTAAGDEGGGQS----LVSPGSCLEDFRAT---PFIECNG-ARGTCHYYANKYS--FWLTTIPEQSFQGAPSA-----DTLKAGLIRTHISRCQVCMKNL----------------
PF01391 // ENSGT00900000140804_1_Domain_9 (1070, 1127)
GPRGDAGQPGAPGTKGLPGERGPPGLGLPGPKGERGFPGDAGLPGPPGFPGPPGLPGAPGQTDCD---SGV
KRPIGADGQETIQ--PGTV--TTGPL--ETVYR-------------VCAHGERPGVPP-G
A----------------RNRAKGLRGVPGFSGADGVPGLKGLPGDPGREGFPGPP-GFMG
PRGSKGAVGPPGLDGLPGASGLPGPAGPPGDRGLPGEVLGAQPGPRGDSGLPGRPGLKGP
PGERGPPGFRGS-----QGMPGMPGQKGQPGSPGFSGQPGLPGAPGQHGF-PGAPGREGP
LGPPGAP-GF--------------------EGLPGDRGEPGDTGVPGPVGRKGGSGDRGD
PGQQGERGHPGLPGFKGVSGMPGTPGLK-------
PF01391 // ENSGT00900000140804_1_Domain_13 (1476, 1538)
GARGSPGMDGFEGMLGLKGRPGLPG
IKGEAGFFGI----PGLKGLAGEPGVKGSRGDPGPPGPPPIILPG-MKDIKGEKGDEGPM
GLKGYLGLK-------------GLPGMPGIPGLSGIP-GLPGRPGHIKGV-K
PF01391 // ENSGT00900000140804_1_Domain_15 (1613, 1700)
GDTGVPGV
PGSPGFPGVPGSPGVMGFQGFTGSRGDKGAPGR---------------------------
---AGLFGEVGETGDF---GDIGDTIDLPGSPGLKGERGTTGIP-GQKGFLGERGTEGDV
GFPGITGLAGVQGPPGFQGQKGFPGLTGLQGPQGDPGRAGAPG--IKGDSGWPGNPGLP-
--------------------------------GLPGLRGISGLHGLPGNKGFPGSPGADV
HG----------D---PGFPGPAGDKGDPGEANTLPGPTGAPGQKGERGAP---------
--------------------------GERGPIGSPGLQGFPGI-TPPSNIS
PF01391 // ENSGT00900000140804_1_Domain_19 (1972, 2034)
GSPGDIGA-
---PGIFGLEGYRGPPGPPGPAALPGSKGDEGSPGTPGNPGIKGWVGDPGPQGRPGVFGL
PGEKG---------PKGEPGFMGNIGPT-----GSPGDRGPRGPKGDRGLPG------AP
-GAVGAPGITGIPQRIAVERGPVGPQGRRGPPGAQGEMGPQGPPGEP-------------
---------------------------------------GFRGAQGKAGPQGRGGVSAVP
GFRGDQGPVGLQGPVGFE----GEPGRPGSPGLPGMPGRSISIGY
PF01413 // ENSGT00900000140804_1_Domain_23 (2266, 2371)
LLVKHSQTDKEPMCP
VGMNKLWSGYSLLYFEGQEKAHNQDLGLAGSCLARFSTMPFLYCNPGDICYYASRNDKSY
WLSTTAPLPMMPVAEED--IRPYISRCSVCENSGT00900000140804_1_Linker_23_25 (2371, 2376)
EAPAVPF01413 // ENSGT00900000140804_1_Domain_25 (2376, 2504)
AIAVHSQ--DVSIPHCPAGWRSLWI
GYSFLMHTAAGDEGGGQS----LVSPGSCLEDFRAT---PFIECNG-ARGTCHYYANKYS
--FWLTTIPEQSFQGAPSA-----DTLKAGLIRTHISRCQVCMKNL--------------
--------------
MGP_PahariEiJ_P0046588 (view gene)
--------------MDRVR-FTASGPPLRGWLLLATVTVGLLAQGVLGGVKKLDVPCGGR
DCS-GGCQCYPEKGARGQPGAVGPQGYNGPPGLQGFPGLQGRKGDKGERGIPGPTGPKGD
VGARGVSGFP
PF01391 // ENSGT00900000140804_1_Domain_0 (131, 189)
GADGIP---GHPGQGGPRGRPGYDGCNGTRGDAGPQGPSGSGGFPGLPGP
QGPKGQKGENSGT00900000140804_1_Linker_0_1 (189, 203)
EPYALSKEDRDKYRPF01391 // ENSGT00900000140804_1_Domain_1 (203, 251)
GEPGEPGLVGYQGPPGRPGPIGQMGPMGAPGRPGPPGP
PGPKGQPGNRENSGT00900000140804_1_Linker_1_2 (251, 321)
GLGF--YGEKGEKGDIGQPGPNGIPSDI---TLIGPTTSTIHPDLYKGEK
GDRG---EQGTPGV----ISPF01391 // ENSGT00900000140804_1_Domain_2 (321, 448)
KGEEGIMGFPGVRGVPGL---DGEKGVLGE----------
---------------------------------------------------------KGS
RGLDGFQGPS-GPRGPKGERGEQGPPGPSVYSPHP-------------------------
-----------------------------------------SLAKGARGDPGFQGAPGEP
GSRGEPGEPGTTGP---P--------------------------GPSVGDED-SMRGLPG
EMGPKGFSGEPGPPARYPGPPGADGRPGPQGVPGPAGSPGPD------------------
-GFLFGLKGSEGRVGYPGPSGFPGARGQKGWKG---------------------------
-----------------------EAGDC-QCG---QVIA
PF01391 // ENSGT00900000140804_1_Domain_6 (760, 1015)
GLPGLPGPKGFPGVNGELGKK
GDQGDPGLHGIPGFPGFKGAPGIAGA----------------------------------
------------------------------------------------------------
------------------------------------------------------------
-----------------------------------------PGP---KG--IKGDSRTIT
TKGERGQPGVPGVHGMKGDDGVPGRDGLDGFPGL-----PGP-PGDG
PF01391 // ENSGT00900000140804_1_Domain_9 (1068, 1127)
IKGPPGDAGLPGI
PGTKGFPGEVGPPGHGLPGPKGERGFPGDAGLPGPPGFPGPPGPPGENSGT00900000140804_1_Linker_9_10 (1127, 1241)
TPGQRDCD---TGV
KRPIGDGQQVIV--QPGCI--GGPT-----------------------GSPGQPGPP---
G----------------PTGAKGVRGMPGFPGASGEQGLKPF01391 // ENSGT00900000140804_1_Domain_10 (1241, 1297)
GFPGDPGREGFPGPP-GFMG
PRGSKGTTGLPGPDGPVGPIGLPGPAGPPGDRGIPGEVLGAQPGTRGDAGLPGQPGLKGL
PGEIGAPGFRGS-----QGMPGMPGLKGQPGFPGPSGQPGQSGPPGQHGF-PGAPGREGP
LGQPGSP-GL--------------------GGLPGDRGEPGDPGVPGPVGMKGLSGDRGD
AGMSGERGHPGSPGFKGMAGMPGIPGQK-------GDRGSPGMDGFQGMLGLKGRPGFPG
TKGEAGFFGV----PGLKGLPGEPGVKGNRGDRGPPGPPPLILPG-MKDIKGEKGDEGPM
GLKGYLGLK-------------GIQGMPGVPGVSGFP-GLPGRPGFIKGV-K
PF01391 // ENSGT00900000140804_1_Domain_15 (1613, 1700)
GDIGVPGT
PGLPGFPGVSGPPGITGFPGFTGSRGEKGTPGV---------------------------
---AGLFGETGPTGDF---GDIGDTVDLPGSPGLKGERGVTGIP-GLKGFFGEKGAEGDI
GFPGITGMAGAQGSPGLKGQTGFPGLTGLQGPQGEPGRIGIPG--DKGDFGWPGVPGLP-
--------------------------------GFPGIRGISGLHGLPGTKGFPGSPGVDA
HG----------D---PGFPGPTGDRGDRGEANTLPGPVGAPGQKGERGTP---------
--------------------------GERGPAGSPGLQGFPGI-SPPSNISGSPGDVGA-
---PGIFGLQGYQGPPGPPGPNALPGVKGEEGSSGAAGFPGEKGWVGDPGPQ
PF01391 // ENSGT00900000140804_1_Domain_20 (2033, 2111)
GQPGVLGL
PGEKG---------PRGEQGFMGNTGPS-----GAVGDRGPKGPKGDQGFPG-------A
PGSMGSPGIPGVPQKIAVQPGTLGPQGRRGLPGALGEIGPQGPPGDP-------------
---------------------------------------GFRGAPGKAGPQGRGGVSAVP
GFRGDQGPMGQQGPVGQE----GEPGRPGSPGLPGMPGRSVSIGY
PF01413 // ENSGT00900000140804_1_Domain_23 (2266, 2371)
LLVKHSQTDQEPMCP
VGMNKLWSGYSLLYFEGQEKAHNQDLGLAGSCLARFSTMPFLYCNPGDVCYYASRNDKSY
WLSTTAPLPMMPVAEEE--IKPYISRCSVCENSGT00900000140804_1_Linker_23_25 (2371, 2376)
EAPAVPF01413 // ENSGT00900000140804_1_Domain_25 (2376, 2504)
AIAVHSQ--DTSIPHCPAGWRSLWI
GYSFLMHTAAGDEGGGQS----LVSPGSCLEDFRAT---PFIECNG-GRGTCHYFANKYS
--FWLTTIPEQNFQSTPSA-----DTLKAGLIRTHISRCQVCMKNL--------------
--------------
ENSRNOP00000054270 (view gene)
------------------------------------------------------------
----------------
PF01391 // ENSGT00900000140804_1_Domain_0 (77, 134)
GQPGEVGPQGYNGPPGLQGFPGLQGRKGDKGERGAPGPTGPKGD
VGARGVSGFPGADGIP---GHPGQGGPRGRPGYDGCNGTRGDAGPQGPSGTGGFPGLPGP
QGPKGQKGEPYALSKEDRDKYRGLPGEPARSGYQGPPGRPGPIGQMGPMGAPGRPGPPGP
PGPKGQPGNRGLGF--YGEKGEKGDVGQPGPNGIPSDI---TLIGPTPSTYHPDMYKGEK
GSQG---EPGIPGI----TLK
PF01391 // ENSGT00900000140804_1_Domain_2 (322, 449)
GEEGIMGFPGTRGFPGL---DGEKGVSGQ----------
---------------------------------------------------------KGS
RGLDGFQGPS-GPRGPKGERGELGPPGPPAYSPHP-------------------------
-----------------------------------------SLAKGARGDPGFQGAHGEP
GSRGEPGDPGPVGP---P--------------------------GLSIGDED-SKRGLPG
EMGPKGFSGEPGPSAYYPGPPGADGKPGPQGLPGPAGPPGPD------------------
-GFLFGLKGSEGRVGYPGPSGFPGGRGQKGWKG---------------------------
-----------------------EAGDC-QCG---QVI
PF01391 // ENSGT00900000140804_1_Domain_6 (759, 1015)
PGLPGLPGPKGFPGVNGEFGKK
GDQGDPGLHGIPGFPGFKGAPGIAGA----------------------------------
------------------------------------------------------------
------------------------------------------------------------
-----------------------------------------PGP---KG--VKGDSRTIT
TKGERGQPGIPGVHGMKGDDGVPGRDGLDGFPGL-----PGP-PGDGI
PF01391 // ENSGT00900000140804_1_Domain_9 (1069, 1127)
KGPPGDAGLPGT
PGTKGFPGEVGPPGQGLPGPKGERGFPGDAGLPGPPGFPGPPGLPGTPGQADCD---TGV
KRPIGGGQQVVI--QPGCV--EGPA-----------------------GSPGQPGPP---
G----------------PTGAKGIRGIPGFPGASGEQGLKGFPGDPGREGFPGPP-GFMG
PRGSKGAPGLPGPDGPPGPIGLPGPAGPPGDRGIPGEVLGAQPGARGDAGLPGQPGLKGF
PGEIGAPGFRGD-----QGMPGMPGLKGQPGFPGPSGQPGLSGPPGQHGF-PGAPGREGP
LGLPGSP-GL--------------------GGLPGDRGEPGEPG-PGPVGMKGVSGDRGD
AGVSGERGHPGSPGF-GMAGMPGIPGQK-------GDR
PF01391 // ENSGT00900000140804_1_Domain_13 (1479, 1539)
GSPGMDGFQGMLGLKGRPGFPG
IKGEAGFFGV----PGLKGLPGEPGVKGNRGDRGPPGPPPLILPG-MKDIKGEKGDEGPM
GLKGYLGLK-------------GIQGMPGVPGLSGIP-GLPGRPGFIKGA-KGDIGVPGT
PGLPGFPGVSGPPGITGFPGFTGSRGEKGTPGV---------------------------
---AGVFGETGPTGDF---GDIGDTVDLP
PF01391 // ENSGT00900000140804_1_Domain_17 (1710, 1765)
GSPGLKGERGVTGIP-GLKGLFGEKGAEGDV
GFPGITGMAGAQGSPGLKGQTGFPGLTGLQGPQGEPGRIGIPG--DKGDFGWPGVPGRP-
--------------------------------GIPGIRGISGLHGLPGTKGFPGSPGVDA
HG----------D---PGFPGPTGDRGDRGEANTLPGPVGAPGQKGEQGIP---------
--------------------------GERGPVGSPGLQGFPGI-SPPSNISGLPGDVGA-
---PGIFGLQGYQGPPGPPGPNALPGIKGDEGSSGAAGFPGEKGWVGDPGPQ
PF01391 // ENSGT00900000140804_1_Domain_20 (2033, 2111)
GQPGVHGL
PGEKG---------PKGEQGFMGNTGPS-----GAVGDRGPKGPKGDQGFPG-------A
PGSMGSPGIPGIPQKIAVQPGTMGPQGRRGLPGALGEMGPQGPPGDP-------------
---------------------------------------GFRGAPGKAGPQGRGGVSAVP
GFRGDQGPMGHQGPIGQE----GEPGRPGSPGLPGMPGRSVSIGY
PF01413 // ENSGT00900000140804_1_Domain_23 (2266, 2371)
LLVKHSQTDQEPMCP
VGMNKLWSGYSLLYFEGQEKAHNQDLGLAGSCLARFSTMPFLYCNPGDVCYYASRNDKSY
WLSTTAPLPMMPVAEEE--IKPYISRCSVCENSGT00900000140804_1_Linker_23_25 (2371, 2376)
EAPAVPF01413 // ENSGT00900000140804_1_Domain_25 (2376, 2504)
AIAVHSQ--DVSIPHCPAGWRSLWI
GYSFLMHTAAGDEGGGQS----LVSPGSCLEDFRAT---PFIECNG-GRGTCHYFANKYS
--FWLTTIPEQNFQSTPSA-----DTLKAGLIRTHISRCQVCMKNL--------------
--------------
ENSPANP00000001006 (view gene)
--------------MGRDQ-RAGAGPALRRWLLLGTVTVGFLAQGVLAGVKKFDVPCGGR
DCS-GGCQCYPEKGGRGQPGPVGPQGYNGPPGLQGFPGLQGRKGDKGERGAPGITGPKGD
VGARGVSGFP
PF01391 // ENSGT00900000140804_1_Domain_0 (131, 189)
GADGIP---GHPGQGGPRGRPGYDGCNGTQGDSGPQGPPGSEGFTGPPGP
QGPKGQKGEPYALPREERDRYRGEPGEPGLVGFQGPPGRPGPVGQMGPVGAPGRPGPPGP
PGPKGQQGNRGLGF--YGVKGEKGDVGQPGPNGIPSDT-LHPIIAPTGVTFHPDQYKGEK
GS---EGEPGIRGI----SLK
PF01391 // ENSGT00900000140804_1_Domain_2 (322, 447)
GEEGIMGFPGPRGSPGL---SGEKGSPGQ----------
---------------------------------------------------------KGS
RGLDGYQGPD-GPRGPKGEAGDLGPPGLPAYSPHP-------------------------
-----------------------------------------SLAKGARGDPGFPGAQGEP
GSQGEPGDPGPRGP---P--------------------------GLSIGDAD-QRRGLPG
EMGPKGFIGDPGIPALYP
PF01391 // ENSGT00900000140804_1_Domain_5 (619, 747)
GPPGPDGKPGPPGPPGLPGPPGPD------------------
-GFLFGRKGAKGTPGFPGLSGSPGARGPKGWKG---------------------------
-----------------------DAGENSGT00900000140804_1_Linker_5_6 (747, 760)
DC-RCAEGDEAVRPF01391 // ENSGT00900000140804_1_Domain_6 (760, 1015)
GLPGLPGPKGFAGINGEPGRK
GDKGDPGQHGLPGFPGLKGVPGNVGA----------------------------------
------------------------------------------------------------
------------------------------------------------------------
-----------------------------------------PGP---KG--AKGDSRTIT
TKGERGQPGVPGVPGMKGDDGSPGRDGLDGFPGL-----PGP-PGDGIKGPPGDAGYPGI
PGTKGTPGEMGPPGLGLPGFKGQRGFPGDAGLPGPPGFPGPPGPAGTPGQIDCD---TDV
KRPIGGDRQEAV--QPGCI--GGPK-----------------------GLPGLPGPP---
G----------------PSGAKGLRGIPGFSGADGGPGPKGLPGDPGREGFPGPP-GFIG
PRGSKGIVGLPGPDGPPGPIGLPGPDGPPGDRGLPGEVLGAQPGPRGDAGVPGQPGFKGL
PGDRGTPGFRGS-----QGMPGMPGLKGQPGLPGPSGQPGLSGPPGQHGF-PGAPGQEGP
LGLPGIP-GL--------------------EGLPGDRGDPGDIGAPGPVGMKGVSGDRGD
AGLAGERGHPGSPGFKGIDGMPGTPGLK-------
PF01391 // ENSGT00900000140804_1_Domain_13 (1476, 1538)
GERGSPGMDGFQGMPGLKGRAGFPG
SKGEAGFFGI----PGLKGLAGEPGFKGSRGDPGPPGPPPIILPG-MKDIKGEKGDEGPM
GLKGYLGAK-------------GIQGMPGIPGLSGIP-GLPGRPGHIKGV-KGDIGAPGI
PGLPGFPGVAGPPGITGFPGFIGSRGDKGAPGR---------------------------
---AGLYGEIGPTGDF---GDIGDTINLPGRPGLKGEQGTAGIP-GLKGFFGEKGTEGDI
GFPGITGVTGVQGPPGLKGQTGFP
PF01391 // ENSGT00900000140804_1_Domain_17 (1765, 1857)
GLTGPPGSQGEPGRIGLPG--GKGDDGWPGIPGLP-
--------------------------------GFPGLRGIRGLHGLPGTKGFPGSPGADI
HG----------D---PGFPGPPGDRGNPGDANTLPGPVGVPGQKGEQGAP---------
--------------------------GERGPPGSPGLQGFPGI-TPPSNISGAPGDKGA-
---PGIFGLKGYRGPPGPPGSAALPGSKGDAGNPGAPGTPGTKGWAGDS
PF01391 // ENSGT00900000140804_1_Domain_20 (2030, 2108)
GPQGRPGVFGL
PGEKG---------PRGEQGFMGNTGPA-----GAVGDRGPKGPKGDPGFPG-------A
PGIVGAPGIAGIPQKIAIPPGTVGPQGRRGPPGAPGEMGPQGPPGEP-------------
---------------------------------------GFRGAPGKAGPQGRGGVSAVP
GFRGDEGPVGHQGPIGQE----GAPGRPGSPGLPGMPGRSVSIGY
PF01413 // ENSGT00900000140804_1_Domain_23 (2266, 2371)
LLVKHSQTDQEPMCP
VGMNKLWSGYSLLYFEGQEKAHNQDLGLAGSCLARFSTMPFLYCNPGDVCYYASRNDKSY
WLSTTAPLPMMPVAEDE--IKPYISRCSVCENSGT00900000140804_1_Linker_23_25 (2371, 2376)
EAPAVPF01413 // ENSGT00900000140804_1_Domain_25 (2376, 2504)
AIAVHSQ--DVSIPHCPAGWRSLWI
GYSFLMHTAAGDEGGGQS----LVSPGSCLEDFRAT---PFIECNG-GRGTCHYYANKYS
--FWLTTIPEQSFQGSPSA-----DTLKAGLIRTHISRCQVCMKNL--------------
--------------
ENSCGRP00000009304 (view gene)
ENSDORP00000009566 (view gene)
ENSMFAP00000004048 (view gene)
-MDL-PCNSRGRTSRARTL-TCLFLPFSCRWLLLGTVTVGFLAQGVLAGVKKFDVPCGGR
DCS-GGCQCYPEKGGR
PF01391 // ENSGT00900000140804_1_Domain_0 (77, 132)
GQPGPVGPQGYNGPPGLQGFPGLQGRKGDKGERGAPGITGPKGD
VGARGVSGFPGADGIP---GHPGQGGPRGRPGYDGCNGTQGDSGPQGPPGSEGFTGPPGP
QGPKGQKGEPYALPREERDRYR
PF01391 // ENSGT00900000140804_1_Domain_1 (203, 251)
GEPGEPGLVGFQGPPGRPGPVGQMGPVGAPGRPGPPGP
PGPKGQQGNRENSGT00900000140804_1_Linker_1_2 (251, 322)
GLGF--YGVKGEKGDVGQPGPNGIPSDT-LHPIVAPTGVTFHPDQYKGEK
GS---EGEPGIRGI----SLKPF01391 // ENSGT00900000140804_1_Domain_2 (322, 447)
GEEGIMGFPGPRGPPGL---SGEKGSPGQ----------
---------------------------------------------------------KGS
RGLDGYQGPD-GPRGPKGEAGDLGPPGLPAYSPHP-------------------------
-----------------------------------------SLAKGARGDPGFPGAQGEP
GSQGEPGDPGPRGP---P--------------------------GLSIGDED-QRRGLPG
EMGPKGFIGDPGIPALYP
PF01391 // ENSGT00900000140804_1_Domain_5 (619, 747)
GPPGPDGKPGPPGPPGLPGPPGPD------------------
-GFLFGRKGAKGTPGFPGLSGSPGARGPKGWKG---------------------------
-----------------------DAGENSGT00900000140804_1_Linker_5_6 (747, 760)
DC-RCAEGDEAVRPF01391 // ENSGT00900000140804_1_Domain_6 (760, 1015)
GLPGLPGPKGFAGINGEPGRK
GDKGDPGQHGLPGFPGLKGVPGNVGA----------------------------------
------------------------------------------------------------
------------------------------------------------------------
-----------------------------------------PGP---KG--AKGDSRTIT
TKGERGQPGVPGVPGMKGDDGSPGRNGLDGFPGL-----PGP-PGDGIKGPPGDAGYPGI
PGTKGTPGEMGPPGLGLPGFKGQRGFPGDAGLPGPPGFPGPPGPAGTPGQIDCD---TDV
KRPIGGDRQEAV--QPGCI--GGPK-----------------------GLPGLPGPP---
G----------------PSGAKGLRGIPGFSGADGGPGPKGLPGDPGREGFPGPP-GFIG
PRGSKGAVGLPGPDGPPGPIGLPGPDGPPGDRGLPGEVLGAQPGPRGDAGVPGQPGFKGL
PGDRGPPGFRGS-----QGMPGMPGLK
PF01391 // ENSGT00900000140804_1_Domain_12 (1348, 1428)
GQPGLPGASGQPGLSGPPGQHGF-PGAPGQEGP
LGLPGIP-GL--------------------EGLPGDRGDPGDIGAPGPVGMKGVSGDRGD
AGLAGERGHPGSPGFKGIDGMPGTPGLK-------GERGSPGMDGFQGMPGLKGRPGFPG
SKGEAGFFGI----PGLKGLAGEPGFKGSRGDPGPPGPPPIILPG-MKDIKGEKGDEGPM
GLKGYLGAK-------------GIQGMPGIPGLSGIP-GLPGRPGHI
PF01391 // ENSGT00900000140804_1_Domain_15 (1608, 1694)
KGV-KGDIGAPGI
PGLPGFPGVAGPPGITGFPGFIGSRGDKGAPGR---------------------------
---AGLYGEIGPTGDF---GDIGDTINLPGRPGLKGEQGTAGIP-GLKGFFGEKGTEGDI
GFPGITGVTGVQGPPGLKGQTGFPGLTGPPGSQGEPGRIGLPG--GKGDDGWPGIAGLP-
--------------------------------GFPGLRGIRGLHGLPGTKGFPGSPGADI
HG----------D---PGFPGPPGDRGDPGDANTLPGPVGVPGQKGEQGAP---------
--------------------------GERGPPGSPGLQGFPGI-TPPSNISGAPGDKGA-
---PGIFGLKGYRGPPGPPGSAALPGSKGDAGNPGAPGTPGTKGWAGDS
PF01391 // ENSGT00900000140804_1_Domain_20 (2030, 2108)
GPQGRPGVFGL
PGEKG---------PRGEQGFMGNTGPA-----GAVGDRGPKGPKGDPGFPG-------A
PGIVGAPGIAGIPQKIAIPPGTVGPQGRRGPPGAPGEMGPQGPPGEP-------------
---------------------------------------GFRGAPGKAGPQGRGGVSAVP
GFRGDEGPVGHQGPIGQE----GAPGRPGSPGLPGMPGRSVSIGY
PF01413 // ENSGT00900000140804_1_Domain_23 (2266, 2371)
LLVKHSQTEQEPMCP
VGMNKLWSGYSLLYFEGQEKAHNQDLGLAGSCLARFSTMPFLYCNPGDVCYYASRNDKSY
WLSTTAPLPMMPVAEDE--IKPYISRCSVCENSGT00900000140804_1_Linker_23_25 (2371, 2376)
EAPAVPF01413 // ENSGT00900000140804_1_Domain_25 (2376, 2504)
AIAVHSQ--DVSIPHCPAGWRSLWI
GYSFLMHTAAGDEGGGQS----LVSPGSCLEDFRAT---PFIECNG-GRGTCHYYANKYS
--FWLTTIPEQSFQGSPSA-----DTLKAGLIRTHISRCQVCMKNL--------------
--------------
ENSTBEP00000004421 (view gene)
------------------------------------------------GVKKFDVPCGGR
DCS-GGCQCYPEKGGR
PF01391 // ENSGT00900000140804_1_Domain_0 (77, 136)
GQPGPVGPQGYSGPPGLQGFPGLQGRKGDKGERGIPGTTGPKGD
VGPRGVSGFPGADGIP---XXXXXXXXXXXXXXXGCNGTRGDAGPQGTSGP-GFPGSPXX
XXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXGPPGP
PGPKGQPGNRGLGF--YGEKGEKGDIGQPGPNGVPSDN-LHPIVGPTKTTVHPDHYKGEK
GS---EGEPGAQGI----SLK
PF01391 // ENSGT00900000140804_1_Domain_2 (322, 422)
GEEGIMGFPGPRGTSGF---DGEKGSPGQ----------
---------------------------------------------------------KGS
RGLXXXXXXX-XXXXXXGEAGAQGPPGLPAYSPHPSLA----------------------
------------------------------------KXXXXXXXXXXXXXXXXXXXXXX-
-----------XXXXXXX--------------------------XXXXXXXXXERRGLPG
EMGPKGFRGDKGFPALYPGPPGTDGEPGLRGPPGPPGLPGPD------------------
-XXXXXXXXXXGSVGYPGPSGFPGA-GQKGWKG---------------------------
-----------------------DSGDC-KCA-GDQFIGGLPGLPGPKGFPGVSGEPGKK
GDQGDPGQHGIPGFPGFK
PF01391 // ENSGT00900000140804_1_Domain_6 (799, 1064)
GAPGLVGL----------------------------------
------------------------------------------------------------
------------------------------------------------------------
-----------------------------------------PGP---KG--IKGDSRTIT
TKGERGQRGVPGVPGMKGDDGLSGQDGLDGFPGL-----PGP-ENSGT00900000140804_1_Linker_6_9 (1064, 1070)
PGDGIQPF01391 // ENSGT00900000140804_1_Domain_9 (1070, 1128)
GPPGDTGHSGA
PGTKGLPGEMGPPGLGLPGPKGLRGTPGDAGLPGPPGFPGPPGPPGTPGQIDCD---SGV
KRPFGGDGQRPV--QPGCV--GGPK-----------------------GSPGRPGPQG--
-----------------PTGAKGLRGLPGLSGADGAPGLKGLPGDPGREGFPGPP-GFVG
PRGSKGAVGLPGPD
PF01391 // ENSGT00900000140804_1_Domain_11 (1275, 1333)
GRPGPSGLPGPVGPPGDRGLPGEVLGAQPGPRGDAGLPGHPGLKGL
PGDRGSPGFRGXXXXXXXXXXXXXXXXXXXXXXXXX---------XXXXX----------
XXXXXXX-XXXXXXXXXXXXXXXXXXXXXXXXXXXXX-X--XX----XXXXXXX---XXX
XXXXXXXXXXXXXXXXXXXXXXXXXXXXXX----XXGRGSPGMDGFQGLPGLQGRPGIPG
SKGEAGFFGV----PGLKGLAGEPGVKGXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXX
XXXXXXXXX-------------XXXXXXXXXXXXXXX-XX--XXXXXXXXXXXXXXXXXX
XXXXXXXXXXXXXXXXXXXXXXXXXGEKGAPGR---------------------------
---PGLYGEIGPTGDLXXXXXXXXXXXXXXXXXXXXXXXXXXXX----XXXXXXXXXXXX
XXXXXXXXXXXXXXXXXXXXXXFP
PF01391 // ENSGT00900000140804_1_Domain_17 (1765, 1859)
GLTGLRGPQGEPGRIGVPG--DKGDYGWPGSPGLP-
--------------------------------GFPGLRGIGGLHGLPGTKGFPGSPGADI
YG----------D---PGSPGPSGDRGEPGEPNTLPGPTGLPGQKGEPGAPXXXXXXXXX
XXXXXXXXXXXXXXXXXXXXXX-------------------------------XXXXXX-
---XXXXXXX-X-----------X--XXXXXXXXXXXXXXXXXXXXXXX--XXXXXXXXX
XXXXX---------XXXXXXXXXXXXXX-----XXXXXX--X-XXXXXXXXX-------X
-XXXXXXXXXXXXXXXXXXXXXX----------------------XX---XXXXXXXXXX
XXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXX-X-----X-XXXXXX
XXXXXXXXXXXXXXXXXX---------XXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXX
XXXXXXX
PF01413 // ENSGT00900000140804_1_Domain_23 (2288, 2373)
SGYSLLYFEGQEKAHNQDLGLAGSCLARFSTMPFLYCNPGDVCYYASRNDKSY
WLSTTAPLPMMPVAEDD--IKPYISRCSVCEAENSGT00900000140804_1_Linker_23_24 (2373, 2374)
PPF01413 // ENSGT00900000140804_1_Domain_24 (2374, 2406)
AVAIAVHSQ--DVSIPHCPAGWRSLWI
GYSFLM------------------------------------------------------
------------------------------------------------------------
--------------
ENSCHOP00000001082 (view gene)
--------------MGRDQ-CAASCPTLRRWLLLGAVTAGFLVQSVLAGVKKFDVPCGGR
DCS-RGCQCYPEKGGRGQPGPV
PF01391 // ENSGT00900000140804_1_Domain_0 (83, 145)
GPQGYTGPPGLQGFPGLQGRKGDKGERGVPGITGPKGD
VGARGVSGFPGADGIP---GHPGQGGSRGRPGNDGCNGTXXXXXXXXXXXXXXXXXXXGP
QGPKGQKGEPYALSKEDRDKYRGEP
PF01391 // ENSGT00900000140804_1_Domain_1 (206, 266)
GDPGLAGFQGPQGHPGPIGQMGPVGFPGRPGPPGP
PGPKGQQGNRGLGF--YGEKGDKGDENSGT00900000140804_1_Linker_1_2 (266, 298)
IGQPGPNGIPSDS-DRAGTGPTKETIHLDQYKPF01391 // ENSGT00900000140804_1_Domain_2 (298, 434)
GEK
GS---EGEPGRKGI----SLKGEEGIMGYPGTRGAPGF---NGEQGSPGQ----------
---------------------------------------------------------KGS
TGPDGYAGSD-GPKGPKXXXXXXXXXXXXXXXXXX-------------------------
-----------------------------------------XXXXXXXXXXXXXXXXXXX
XXXXXXXXXXXXXX---X--------------------------XXXXXXXX-XXXXXXX
XXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXX------------------
-XFLFGLKGAEGRVGYPGPSGFPGA-GQKGWKG---------------------------
-----------------------DAGDC-KCAEGDQF
PF01391 // ENSGT00900000140804_1_Domain_6 (758, 1016)
ITGLPGLQGPKGFPGDNGEPGKK
GDQGDPGQHGVPGFPGFKGTPGDLGP----------------------------------
------------------------------------------------------------
------------------------------------------------------------
-----------------------------------------PGR---KG--EKGDENSGT00900000140804_1_Linker_6_7 (1016, 1022)
SRTIT
TPF01391 // ENSGT00900000140804_1_Domain_7 (1022, 1058)
KGDRGQPGVPGVPGVKGDDGVPGRDGLDGFPGP---GPRXX-XXXXXXXXXXXXXXXXX
XXXXXXXXXXX---XXXX--XXXXXXXXXXXXXXXXXXX-X-XXXXXXXXXXXX---XXX
XXXXXXXXXXXXXXXXXXX--XXXX-----------------------XXXXXXXXX---
X----------------XXXXXTKLQGTGFSGADGPPGLKGP--GN
PF01391 // ENSGT00900000140804_1_Domain_10 (1247, 1297)
GREGYPGPP-GFVG
PRGSKGAVGLPGPDGHPGPTGLPGPVGPPGDRGLPGEVLGAQAGPRGDAGLPGHSGLKGL
PGERGTAGFRGS-----QGMPGMPGLKGQPGFPGPSGQPGQPGPPGQHGF-PGAAGPEGP
LGTPGVP-GF--------------------EGLPGDRGAPGDIGVPGPVGMKGVSGDRGD
PGSLGEKGHPGNIGFKGMAGMPGIPGPK-------GDRGSPGMDGFQGMVGLKGRPGFPG
NKGEAGVFGF----PGLKGLAGEPGVKGNRGDPGPPGPPPVILPG-MKDI
PF01391 // ENSGT00900000140804_1_Domain_14 (1551, 1607)
KGEKGDQGPM
GLKGYLGLK-------------GLQGMQGVPGLAGIP-GSPGKPGHIKGL-KGDIGFPGL
PGLPGFPGVTGPPGISGFPGFIGTRGEKGAPGR---------------------------
---AGLYGATGPTGDF---GDIGDTID-------------------LKGFFGEKGTEGNR
GFPGITGITGAQGPPGLKGQTGFPGLTGLQGPQGEPGRIGVPG--EKGDYGWPGAAGPP-
--------------------------------GFPGIRGIDGLHGLPGTKGFPGSPGADI
SG----------D---PGFPGTTGDRGDPGEPNTFPGPTGVPGQKGERGIP---------
--------------------------GERGPMGSPGLQGFPGV-TPPSNISGLPGEKGA-
---PGLFGLKGYQGPQGPPGPAALPGSQGDGGNPGASGNPGTKGWV
PF01391 // ENSGT00900000140804_1_Domain_20 (2027, 2107)
GDPGPQGRPGVFGI
PGEKG---------TRGEQGLMGNTGAT-----GSMGDRGPKGPKGDRGLPG------PP
-GSLGSENSGT00900000140804_1_Linker_20_21 (2107, 2136)
PGIVGIPQRIVPEPGTLGPQGRRGFPGAPPF01391 // ENSGT00900000140804_1_Domain_21 (2136, 2251)
GEMGPQGPPGEP-------------
---------------------------------------GFRGEQGRTGPQGRGGVSAVP
GFRGDQGPVGHQGPIGQE----GEPGRPGSENSGT00900000140804_1_Linker_21_23 (2251, 2264)
PGLPGMPGRSVSIPF01413 // ENSGT00900000140804_1_Domain_23 (2264, 2373)
GYLLVKHSQTDQEPMCP
VGMNTLWSGYSLLYFEGQEKAHNQDLGLAGSCLARFSTMPFLYCNPGDVCYYASRNDKSY
WLSTTAPLPMMPVAEED--IKPYISRCSVCEAENSGT00900000140804_1_Linker_23_25 (2373, 2374)
PPF01413 // ENSGT00900000140804_1_Domain_25 (2374, 2505)
AVAIAVHSQ--DVSIPHCPAGWRSLWI
GYSFLMHTAAGDEGGGQS----LVSPGSCLEDFRAT---PFIECNG-GRGTCHYYANKYS
--FWLTTIPEQSFQGSPSA-----DTLKAGLIRTHISRCQVCMKNL--------------
--------------
ENSOGAP00000009095 (view gene)
ENSSHAP00000011227 (view gene)
--------------MGSRM-YPTSRLARQRCLLLF-VLITVLTERAHAGVKKYDVPCGGR
DCS-GGCQCYPEK
PF01391 // ENSGT00900000140804_1_Domain_0 (74, 132)
GARGQPGQVGPQGYDGPPGLQGIPGLQGRKGDKGERGAPGITGPKGD
VGQRGVSGFPGADGIP---GHPGQGGPRGRPGSDGCNGTTGDAGTQGPPGSGGLPGLPGP
QGPKGQKGEPYAFSPAERDKYR
PF01391 // ENSGT00900000140804_1_Domain_1 (203, 251)
GDSGEPGFIGLPGPVGYPGSPGKIGPVGAPGRPGPPGP
PGPKGQPGNRENSGT00900000140804_1_Linker_1_2 (251, 322)
GLGF--YGEKGEKGDVGRPGPTGLAPDSDLA-TSFLENVVDHLDQYKGEK
GN---WGEPGWKGI----SEKPF01391 // ENSGT00900000140804_1_Domain_2 (322, 448)
GEEGIVGFPGQRGLPGN---NGREGDQGE----------
---------------------------------------------------------KGI
RGDVGFPGID-GFKGLPGDIGDRGPPGPPSYSPHP-------------------------
-----------------------------------------SVIKGIRGDPGDTGAYGPP
GNPGAQGDPGQPGP---P--------------------------GTTIGDGD-ER
PF01391 // ENSGT00900000140804_1_Domain_5 (596, 662)
RGLPG
EMGAKGFIGDPGIPALYPGPPGIDGKPGLPGKPGSPGPPGPD------------------
-ENSGT00900000140804_1_Linker_5_6 (662, 759)
GFLFGLKGAEGNVGYPGPSGIPGTRGQKGWKG---------------------------
-----------------------DPGDC-KC----DIPPF01391 // ENSGT00900000140804_1_Domain_6 (759, 1015)
SGLPGPPGPIGRPGANGEPGRK
GEIGDPGAHGIPGFPGFKGNPGIPGL----------------------------------
------------------------------------------------------------
------------------------------------------------------------
-----------------------------------------PGF---KG--QKGENSGT00900000140804_1_Linker_6_7 (1015, 1022)
DSKTIT
TPF01391 // ENSGT00900000140804_1_Domain_7 (1022, 1079)
KGSRGDQGNPGMPGLQGEDGFPGRDGLDGFPGL-----PGL-PGDGIKGLPGDSGFPGVPGLKGIPGESGPPGLGLPGPKGERGFPGDPGLSGSPGFPGPPGPPGTPGQTDCD---GGVKEPIGQE---II--RPGCV--IPPK-----------------------GSQGLPGLP---G----------------SPGAKGQRGLPGLPGVDGFIGLKGFQGDAGREGFPGPP-GFVGPRGSKGATGLPGLDGSPGSSGLPGSVGPAGQKGLPGEVLGAQTGLPGDPGLPGIPGLKGEQGNRGPQGFRGS-----PGMPGMPGLKGQPGFPGPSGQPGLPGPPGTHGF-PGPQGQAGLPGLPGPS-GF--------------------AGLPGDRGDRGDIGVPGPVGMKGLDGDRGDTGFIGEQGRRGEPGFQGIPGLPGIPGTK-------GFKGSPGMEGFKGMLGIKGRSGIIGYKGEIGDFGI----PGLKGLTGEPGLKGSRGEPGPPGAPPKFLPG-MKDFKGEKGDEGPVGGKGYWGIK-------------GGQGMPGIPGQAGIP-GLPGRPGHIKGA-KGEIGVPGVPGLQGFPGVIGSPGIRGFPGFVGVRGDKGMPGQ------------------------------EGLYGETGSIGDS---GDEGDTINLPGRPGLKGEVGTSGLA-GEKGFTGEKGTLGDHGFTGLMGIKGSQGLPGLKGQTGFPGLTGLPGPQGDPGRAGPPG--EKGDQGWPGSPGLS---------------------------------GFPGIRGISGLHGLPGTKGFPGSPGTDGYG----------N---PGFPGLFGDKGERGEPNTIPGPLGAPGQKGERGTI-----------------------------------GEQGPSGSPGPQGLPGF-TSPSNISGLPGDKGS----PGLFGPQGYRGPPGAPGPTSFPGIKGEVGSPGFTGERGPKGWPGDPGPQGRPGVYGLPGEKG---------PKGQQGLMGYHGIP-----GSIGDRGPKGPKGDRGFPG-------IPGAMGSPGIPGVPLRIATEPGVAGPHGKRGPPGMQGEMGPQGPPGDP----------------------------------------------------GLRGLPGKPGPQGRGGMSALPGFRGDEGPMGHQGPVGQE----GEPGRPGSPGLPGMPGRSVSIGYLLVKHSQTDQEPMCPVGMNKLWSGYSLLYFEGQEKAHNQDLGLAGSCLARFSTMPFLYCNPGDICHYASRNDKSYWLSTTAPLPMMPVAEDE--IKPYISRCSVCEAPAVAIAVHSQ--DVSIPHCPAGWRSLWIGYSFLMHTAAGDEGGGQS----LVSPGSCLEDFRAT---PFIECNG-GRGTCHYFANKYS--FWLTTIPEQNFQASPSA-----DTLKAGLIRTHISRCQNLQFLPKKQTNCHEVLYQQAHLMKMSDY
PF01391 // ENSGT00900000140804_1_Domain_9 (1073, 1130)
GDSGFPGV
PGLKGIPGESGPPGLGLPGPKGERGFPGDPGLSGSPGFPGPPGPPGTPGENSGT00900000140804_1_Linker_9_10 (1130, 1188)
QTDCD---GGV
KEPIGQE---II--RPGCV--IPPK----------------------PF01391 // ENSGT00900000140804_1_Domain_10 (1188, 1257)
-GSQGLPGLP---
G----------------SPGAKGQRGLPGLPGVDGFIGLKGFQGDAGREGFPGPP-GFVG
PRGSKGATGLPGLDGSPGSSGLPGSVGPAGQKGLPGEVLGAQTGLPGDPGLPGIPGLKGE
QGNRGPQGFRGS-----PGMPGMPGLKGQPGFPGPSGQPGLPGPPGTHGF-PGPQGQAGL
PGLPGPS-GF--------------------AGLPGDRGDRGDIGVPGPVGMKGLDGDRGD
TGFIGEQGRRGEPGFQGIPGLPGIPGTK-------GFKGSPGMEGFKGMLGIKGRSGIIG
YKGEIGDFGI----PGLKGLTGEPGLKGSRGEPGPPGAPPKFLPG-MKDFKGEKGDEGPV
GGKGYWGIK-------------GGQGMPGIPGQAGIP-GLPGRPGHIKGA-KGEIGVPGV
PGLQGFPGVIGSPGIRGFPGFVGVRGDKGMPGQ---------------------------
---EGLYGETGSIGDS---GDEGDTINL
PF01391 // ENSGT00900000140804_1_Domain_17 (1709, 1765)
PGRPGLKGEVGTSGLA-GEKGFTGEKGTLGDH
GFTGLMGIKGSQGLPGLKGQTGFPGLTGLPGPQGDPGRAGPPG--EKGDQGWPGSPGLS-
--------------------------------GFPGIRGISGLHGLPGTKGFPGSPGTDG
YG----------N---PGFPGLFGDKGERGEPNTIPGPLGAPGQKGERGTI---------
--------------------------GEQGPSGSPGPQGLPGF-TSPSNISGLPGDKGS-
---PGLFGPQ
PF01391 // ENSGT00900000140804_1_Domain_19 (1991, 2058)
GYRGPPGAPGPTSFPGIKGEVGSPGFTGERGPKGWPGDPGPQGRPGVYGL
PGEKG---------PKGENSGT00900000140804_1_Linker_19_22 (2058, 2203)
QQGLMGYHGIP-----GSIGDRGPKGPKGDRGFPG-------I
PGAMGSPGIPGVPLRIATEPGVAGPHGKRGPPGMQGEMGPQGPPGDP-------------
---------------------------------------GLRPF01391 // ENSGT00900000140804_1_Domain_22 (2203, 2259)
GLPGKPGPQGRGGMSALP
GFRGDEGPMGHQGPVGQE----GEPGRPGSPGLPGMPGENSGT00900000140804_1_Linker_22_23 (2259, 2265)
RSVSIGPF01413 // ENSGT00900000140804_1_Domain_23 (2265, 2372)
YLLVKHSQTDQEPMCP
VGMNKLWSGYSLLYFEGQEKAHNQDLGLAGSCLARFSTMPFLYCNPGDICHYASRNDKSY
WLSTTAPLPMMPVAEDE--IKPYISRCSVCEENSGT00900000140804_1_Linker_23_25 (2372, 2375)
APAPF01413 // ENSGT00900000140804_1_Domain_25 (2375, 2500)
VAIAVHSQ--DVSIPHCPAGWRSLWI
GYSFLMHTAAGDEGGGQS----LVSPGSCLEDFRAT---PFIECNG-GRGTCHYFANKYS
--FWLTTIPEQNFQASPSA-----DTLKAGLIRTHISRCQNLQFLPKKQTNCHEVLYQQA
HLMKMSDYMKKCMR
ENSMOCP00000024736 (view gene)
--------------MDRVK-LAASGHALRGWLLLATVTVLLLAPSVLAGVKKLDVPCGGR
DCS-GGCQCYPE
PF01391 // ENSGT00900000140804_1_Domain_0 (73, 143)
KGARGQPGPVGTQGYNGPPGLQGFPGLQGRKGDKGERGAPGPTGPKGD
V------------------GHPGQGGPRGRPGYDGCNGTKGDAGPQGPPGPGGFPGLPGP
QGPKGQKGEPYALSDKDRDKYR
PF01391 // ENSGT00900000140804_1_Domain_1 (203, 251)
GESGEPGVPGYQGPPGRPGPIGQMGPIGAPGRPGPPGP
PGPKGQPGNRENSGT00900000140804_1_Linker_1_2 (251, 322)
GLGF--YGEKGEKGDIGQPGPNGIPSDV---TLIGPTPSTYDRDLYKGEK
GSRG---DQGIPGP----HSKPF01391 // ENSGT00900000140804_1_Domain_2 (322, 448)
GEEGIMGFPGTRGLPGL---DGEKGVFGQ----------
---------------------------------------------------------KGS
RGLDGFQGPS-GPRGPKGERGEQGPPGPPVYSPHP-------------------------
-----------------------------------------SLAKGARGDPGFPGAHGEP
GSRGEPGDPGPMGP---Q--------------------------GPSVGDKD-SMRGLPG
EMGPKGFSGEPGRPAQYPGPPGADGKPGLQGAPGPAGAPGPD------------------
-GFLFGLKGSEGREGYPGPSGFPGARGQKGWKG---------------------------
-----------------------EAGDC-QCD---QFIG
PF01391 // ENSGT00900000140804_1_Domain_6 (760, 1014)
GLPGLPGPKGFPGVNGELGKK
GDQGEPGVHGIPGFPGFKGAPGLTGL----------------------------------
------------------------------------------------------------
------------------------------------------------------------
-----------------------------------------PGP---KG--IKGDSRTVT
TKGERGQPGIPGVHGMKGDNGVPGRDGLDGFPGL-----PGP-PGDGIKGPPGDAGLPGL
PGTKGFPGEMGPPGQGLPGPKGERGFPGDAGLPGLPGFDGPPGPPGTPGQLDCD---TGV
KRPIGDGQQVIV--QPGCI--GGPP-----------------------GPPGQPGPP---
G----------------PTGIKGSRGMPGVPGEFGGPGLKGFPGDPGREGFPGPP-GFMG
PRGSKGATGLPGPDGPPGPIGLPGPAGPPGDRGIPGEVLGAQPGPRGDAGLPGHPGLKGP
PGERGPPGFRGS-----QGMPGMPGSK
PF01391 // ENSGT00900000140804_1_Domain_12 (1348, 1428)
GQPGFPGPSGQPGLSGPPGQHGF-PGAPGREGP
LGLPGSP-GL--------------------GGLPGDRGEPGEPGVPGPVGMKGLSGDRGD
AGVSGERGHPGSPGFKGMAGMPGVPGQK-------GVRGSPGMDGFQGLLGLKGRPGFPG
VKGEAGFFGV----PGLKGLPGEPGVKGNRGDRGPPGPPPLILPG-MKDIKGEKGDEGPM
GLKGYLGLK-------------GNLGMPGIPGQAGIP-GLPGRPGYIK
PF01391 // ENSGT00900000140804_1_Domain_15 (1609, 1694)
GV-KGDIGVPGI
PGTPGQPGVAGPPGITGFPGFVGSRGEKGAPGV---------------------------
---AGLFGETGPTGDF---GDIGDTVNLAGSPGLKGERGVTGIP-GIKGLFGEKGAEGDI
GFPGITGMAGAQGSPGLKGQTGFPGLTGLQGPQGDPGRSGLPG--DKGDYGWPGVAGLP-
--------------------------------GFPGIRGISGLHGLPGTKGFPGSPGADV
HG----------D---PGFPGPAGDRGDRGEPNTFPGPVGPAGQKGERGTP---------
--------------------------GERGPAGSPGLQGFPGI-SPPSNITGSPGDVGA-
---PGIFGLEGYR
PF01391 // ENSGT00900000140804_1_Domain_19 (1994, 2060)
GPPGPPGSDALPGIKGDEGSAGAAGFPGQKGWVGDPGPQGQPGVVGL
PGEKG---------PKGEQGFMGNTGPS-----GTVGDRGPKGPKGDQGFPG-------A
PGSMGAPGIQGIPQKIATQPGIVGPQGRRGLPGAQGEIGPQGPPGDP-------------
---------------------------------------GFRGAPGKAGPQGRGGVSAVP
GFRGDQGPMGQQGPIGQE----GEPGRPGSPGLPGMPGRSVSIGY
PF01413 // ENSGT00900000140804_1_Domain_23 (2266, 2371)
LLVKHSQTDQEPMCP
VGMNKLWSGYSLLYFEGQEKAHNQDLGLAGSCLARFSTMPFLYCNPGDVCYYASRNDKSY
WLSTTAPLPMMPVAEEE--IKPYISRCSVCENSGT00900000140804_1_Linker_23_25 (2371, 2376)
EAPAVPF01413 // ENSGT00900000140804_1_Domain_25 (2376, 2504)
AIAVHSQ--DVSIPHCPAGWRSLWI
GYSFLMHTAAGDEGGGQS----LVSPGSCLEDFRAT---PFIECNG-GRGTCHYYANKYS
--FWLTTIPEQNFQGTPSA-----DTLKAGLIRTHISRCQVCMKNL--------------
--------------
ENSMMUP00000029330 (view gene)
--------------MGRDQ-RAVAGPALRRWLLLGTVTVGFLAQGVLAGVKKFDVPCGGR
DCS-GGCQCYPEKGGRGQPGPVGPQGYNGPPGLQGFPGLQGRKGDKGERGAPGITGPKGD
VGARGVSGFP
PF01391 // ENSGT00900000140804_1_Domain_0 (131, 189)
GADGIP---GHPGQGGPRGRPGYDGCNGTQGDSGPQGPPGSEGFTGPPGP
QGPKGQKGENSGT00900000140804_1_Linker_0_1 (189, 203)
EPYALPREERDRYRPF01391 // ENSGT00900000140804_1_Domain_1 (203, 251)
GEPGEPGLVGFQGPPGRPGPVGQMGPVGAPGRPGPPGP
PGPKGQQGNRENSGT00900000140804_1_Linker_1_2 (251, 322)
GLGF--YGVKGEKGDVGQPGPNGIPSDT-LHPIVAPTGVTFHPDQYKGEK
GS---EGEPGIRGI----SLKPF01391 // ENSGT00900000140804_1_Domain_2 (322, 447)
GEEGIMGFPGPRGPPGL---SGEKGSPGQ----------
---------------------------------------------------------KGS
RGLDGYQGPD-GPRGPKGEAGDLGPPGLPAYSPHP-------------------------
-----------------------------------------SLAKGARGDPGFPGAQGEP
GSQGEPGDPGPRGP---P--------------------------GLSIGDED-QRRGLPG
EMGPKGFIGDPGIPALYP
PF01391 // ENSGT00900000140804_1_Domain_5 (619, 747)
GPPGPDGKPGPPGPPGLPGPPGPD------------------
-GFLFGRKGAKGTPGFPGLSGSPGARGPKGWKG---------------------------
-----------------------DAGENSGT00900000140804_1_Linker_5_6 (747, 760)
DC-RCAEGDEAVRPF01391 // ENSGT00900000140804_1_Domain_6 (760, 1015)
GLPGLPGPKGFAGINGEPGRK
GDKGDPGQHGLPGFPGLKGVPGNVGA----------------------------------
------------------------------------------------------------
------------------------------------------------------------
-----------------------------------------PGP---KG--AKGDSRTIT
TKGERGQPGVPGVPGMKGDDGSPGRNGLDGFPGL-----PGP-PGDGIKGPPGDAGYPGI
PGTKGTPGEMGPPGLGLPGFKGQRGFPGDAGLPGPPGFPGPPGPAGTPGQIDCD---TDV
KRPIGGDRQEAV--QPGCI--GGPK-----------------------GLPGLPGPP---
G----------------PSGAKGLRGIPGFSGADGGPGPKGLPGDPGREGFPGPP-GFIG
PRGSKGAVGLPGPDGPPGPIGLPGPDGPPGDRGLPGEVLGAQPGPRGDAGVPGQPGFKGL
PGDRGPPGFRGS-----QGMPGMPGLK
PF01391 // ENSGT00900000140804_1_Domain_12 (1348, 1428)
GQPGLPGASGQPGLSGPPGQHGF-PGAPGQEGP
LGLPGIP-GL--------------------EGLPGDRGDPGDIGAPGPVGMKGVSGDRGD
AGLAGERGHPGSPGFKGIDGMPGTPGLK-------GERGSPGMDGFQGMPGLKGRPGFPG
SKGEAGFFGI----PGLKGLAGEPGFKGSRGDPGPPGPPPIILPG-MKDIKGEKGDEGPM
GLKGYLGAK-------------GIQGMPGIPGLSGIP-GLPGRPGHIKGV-KGDIGAPGI
PGLPGFPGVAGPPGITGFPGFIGSRGDKGAPGR---------------------------
---AGLYGEIGPTGDF---GDIGDTINLPGRPGLKGEQGTAGIP-GLKGFFGEKGTEGDI
GFPGITGVTGVQGPPGLKGQTGFP
PF01391 // ENSGT00900000140804_1_Domain_17 (1765, 1857)
GLTGPPGSQGEPGRIGLPG--GKGDDGWPGIPGLP-
--------------------------------GFPGLRGIRGLHGLPGTKGFPGSPGADI
HG----------D---PGFPGPPGDRGDPGDANTLPGPVGVPGQKGEQGAP---------
--------------------------GERGPPGSPGLQGFPGI-TPPSNISGAPGDKGA-
---PGIFGLKGYRGPPGPPGSAALPGSKGDAGNPGAPGTPGTKGWAGDS
PF01391 // ENSGT00900000140804_1_Domain_20 (2030, 2108)
GPQGRPGVFGL
PGEKG---------PRGEQGFMGNTGPA-----GAVGDRGPKGPKGDPGFPG-------A
PGIVGAPGIAGIPQKIAIPPGTVGPQGRRGPPGAPGEMGPQGPPGEP-------------
---------------------------------------GFRGAPGKAGPQGRGGVSAVP
GFRGDEGPVGHQGPIGQE----GAPGRPGSPGLPGMPGRSVSIGY
PF01413 // ENSGT00900000140804_1_Domain_23 (2266, 2371)
LLVKHSQTEQEPMCP
VGMNKLWSGYSLLYFEGQEKAHNQDLGLAGSCLARFSTMPFLYCNPGDVCYYASRNDKSY
WLSTTAPLPMMPVAEDE--IKPYISRCSVCENSGT00900000140804_1_Linker_23_25 (2371, 2376)
EAPAVPF01413 // ENSGT00900000140804_1_Domain_25 (2376, 2504)
AIAVHSQ--DVSIPHCPAGWRSLWI
GYSFLMHTAAGDEGGGQS----LVSPGSCLEDFRAT---PFIECNG-GRGTCHYYANKYS
--FWLTTIPEQSFQGSPSA-----DTLKAGLIRTHISRCQVCMKNL--------------
--------------
ENSCATP00000017946 (view gene)
-MDL-PCNSRGRTSRARTL-TCLFLPFSCRWLLLGTVTVGFLAQGVLAGVKKFDVPCGGR
DCS-GGCQCYPEKGGRGQPGPVGPQGYNGPPGLQGFPGLQGRKGDKGERGAPGITGPKGD
VGARGVSGFP
PF01391 // ENSGT00900000140804_1_Domain_0 (131, 189)
GADGIP---GHPGQGGPRGRPGYDGCNGTQGDSGPQGPPGSEGFTGPPGP
QGPKGQKGEPYALPREERDRYRGEPGEPGLVGFQGPPGRPGPVGQMGPVGAPGRPGPPGP
PGPKGQQGNRGLGF--YGVKGEKGDVGQPGPNGIPSDT-LHPIIAPTGVTFHPDQYKGEK
GS---EGEPGIRGI----SLK
PF01391 // ENSGT00900000140804_1_Domain_2 (322, 447)
GEEGIMGFPGPRGSPGL---SGEKGSPGQ----------
---------------------------------------------------------KGS
RGLDGYQGPD-GPRGPKGEAGDLGPPGLPAYSPHP-------------------------
-----------------------------------------SLAKGARGDPGFPGAQGEP
GSQGEPGDPGPRGP---P--------------------------GLSIGDED-QRRGLPG
EMGPKGFIGDPGIPALYP
PF01391 // ENSGT00900000140804_1_Domain_5 (619, 747)
GPPGPDGKPGPPGPPGLPGPPGPD------------------
-GFLFGLKGPEGRPGFPGLSGSPGARGPKGWKG---------------------------
-----------------------DAGENSGT00900000140804_1_Linker_5_6 (747, 760)
DC-RCAEGDEAVRPF01391 // ENSGT00900000140804_1_Domain_6 (760, 1015)
GLPGLPGPKGFAGINGEPGRK
GDKGDPGQHGLPGFPGLKGVPGNVGA----------------------------------
------------------------------------------------------------
------------------------------------------------------------
-----------------------------------------PGP---KG--AKGDSRTIT
TKGERGQPGVPGVPGMKGDDGSPGRDGLDGFPGL-----PGP-PGDGIKGPPGDAGYPGI
PGTKGTPGEMGPPGLGLPGFKGQRGFPGDAGLPGPPGFPGPPGPAGTPGQIDCD---TDV
KRPIGGDRQEAI--QPGCI--GGPK-----------------------GLPGLPGPP---
G----------------PSGAKGLRGIPGFSGADGGPGPKGLPGDPGREGFPGPP-GFIG
PRGSKGAVGLPGPDGPPGPIGLPGPDGPPGDRGLPGEVLGAQPGPRGDAGVPGQPGFKGL
PGDRGTPGFRGS-----QGMPGMPGLKGQPGLPGPSGQPGLSGPPGQHGF-PGAPGQEGP
LGLPGIP-GL--------------------EGLPGDRGDPGDIGAPGPVGMKGVSGDRGD
AGLAGERGHPGSPGFKGIDGMPGTPGLK-------
PF01391 // ENSGT00900000140804_1_Domain_13 (1476, 1538)
GERGSPGMDGFQGMPGLKGRPGFPG
SKGEAGFFGI----PGLKGLAGEPGFKGSRGDPGPPGPPPIILPG-MKDIKGEKGDEGPM
GLKGYLGAK-------------GIQGMPGIPGLSGIP-GLPGRPGHIKGV-KGDIGAPGI
PGLPGFPGVAGPPGITGFPGFIGSRGDKGAPGR---------------------------
---AGLYGEIGPTGDF---GDIGDTINLPGRPGLKGEQGTAGIP-GLKGFFGEKGTEGDI
GFPGITGVTGVQGPPGLKGQTGFP
PF01391 // ENSGT00900000140804_1_Domain_17 (1765, 1848)
GLTGPPGSQGEPGRIGLPG--GKGDDGWPGIPGLP-
--------------------------------GFPGLRGIRGLHGLPGTKGFPGSPGADI
HG----------D---PGFPGPPGDRGNPGDANTLPGPVGVPGQKGEQGAP---------
--------------------------GERGPPGSPGLQGFPGI-TPPSNIS
PF01391 // ENSGT00900000140804_1_Domain_19 (1972, 2034)
GAPGDKGA-
---PGIFGLKGYRGPPGPPGSAALPGSKGDAGNPGAPGTPGTKGWAGDSGPQGRPGVFGL
PGEKG---------PRGEQGFMGNTGPA-----GAVGDRGPKGPKGDPGFPG-------A
PGIVGAPGIAGIPQKIAIPPGTVGPQGRRGPPGAPGEMGPQGPPGEP-------------
---------------------------------------GFRGAPGKAGPQGRGGVSAVP
GFRGDEGPVGHQGPIGQE----GAPGRPGSPGLPGMPGRSVSIGY
PF01413 // ENSGT00900000140804_1_Domain_23 (2266, 2371)
LLVKHSQTDQEPMCP
VGMNKLWSGYSLLYFEGQEKAHNQDLGLAGSCLARFSTMPFLYCNPGDVCYYASRNDKSY
WLSTTAPLPMMPVAEDE--IKPYISRCSVCENSGT00900000140804_1_Linker_23_25 (2371, 2376)
EAPAVPF01413 // ENSGT00900000140804_1_Domain_25 (2376, 2504)
AIAVHSQ--DVSIPHCPAGWRSLWI
GYSFLMHTAAGDEGGGQS----LVSPGSCLEDFRAT---PFIECNG-GRGTCHYYANKYS
--FWLTTIPEQSFQGSPSA-----DTLKAGLIRTHISRCQVCMKNL--------------
--------------
ENSPSIP00000014136 (view gene)
ENSXETP00000005650 (view gene)
ENSPTRP00000058373 (view gene)
--------------MGRDQ-RAVAGPALRRWL-LGTVTVGFLAQSVLAGVKKFDVPCGGR
DCS-GGCQCYPEKGGRGQPGPVGPQGYNGPPGLQGFPGLQGRKGDKGERGAPGITGPKGD
VGARGVSGFP
PF01391 // ENSGT00900000140804_1_Domain_0 (131, 190)
GADGIP---GHPGQGGPRGRPGYDGCNGTQGDSGPQGPPGSEGFTGPPGP
QGPKGQKGEENSGT00900000140804_1_Linker_0_1 (190, 203)
PYALPKEERDRYRPF01391 // ENSGT00900000140804_1_Domain_1 (203, 251)
GEPGEPGLVGFQGPPGRPGHVGQMGPVGAPGRPGPPGP
PGPKGQQGNRENSGT00900000140804_1_Linker_1_2 (251, 322)
GLGF--YGVKGEKGDVGQPGPNGIPSDL-LHPIIAPTGVTFHPDQYKGEK
GS---EGEPGIRGI----SLKPF01391 // ENSGT00900000140804_1_Domain_2 (322, 447)
GEEGIMGFPGLRGYPGL---SGEKGSPGQ----------
---------------------------------------------------------KGS
RGLDGYQGPD-GPRGPKGEAGDPGPPGLPAYSPHP-------------------------
-----------------------------------------SLAKGARGDPGFPGAQGEP
GSQGEPGDPGLPGA---P--------------------------GLSIGDGD-QRRGLPG
EMGPKGFIGDPGIPALYG
PF01391 // ENSGT00900000140804_1_Domain_5 (619, 747)
GPPGPDGKRGPPGPPGLPGPPGPD------------------
-GFLFGLKGAKGRAGFPGLPGSPGARGPKGWKG---------------------------
-----------------------DAGENSGT00900000140804_1_Linker_5_6 (747, 760)
EC-RCTEGDEAIKPF01391 // ENSGT00900000140804_1_Domain_6 (760, 1015)
GLPGLPGPKGFAGINGEPGRK
GDKGDPGQHGLPGFPGLKGVPGNVGA----------------------------------
------------------------------------------------------------
------------------------------------------------------------
-----------------------------------------PGP---KG--AKGDSRTIT
TKGERGQPGVPGVPGMKGDDGSPGRDGLDGFPGL-----PGP-PGDGIKGPPGDPGYPGI
PGTKGTPGEMGPPGLGLPGLKGQRGFPGDAGLPGPPGFLGPPGPAGTPGQIDCD---TDV
KRAIGGDRQEAI--QPGCV--GGPK-----------------------GLPGLPGPP---
G----------------PTGAKGLRGIPGFSGADGGPGPKGLPGDAGREGFPGPP-GFIG
PRGSKGAVGLPGPDGSPGPIGLPGPDGPPGERGLPGEVLGAQPGPRGDAGVPGQPGLKGL
PGDRGPPGFRGS-----QGMPGMPGLKGQPGLPGPSGQPGLYGPPGLHGF-PGAPGQEGP
LGLPGIP-GR--------------------EGLPGDRGDPGDTGAPGPVGMKGLSGDRGD
AGFTGERGHPGSPGFKGIDGMPGTPGLK-------
PF01391 // ENSGT00900000140804_1_Domain_13 (1476, 1538)
GDRGSPGMDGFQGMPGLKGRPGFPG
SKGEAGFFGI----PGLKGLAGEPGFKGSRGDPGPPGPPPVILPG-MKDIKGEKGDEGPM
GLKGYLGAK-------------GIQGMPGIPGLSGIP-GLPGRPGHIKGV-KGDIGAPGI
PGLPGFPGVAGPPGITGFPGFIGSRGDKGAPGR---------------------------
---AGLYGEIGATGDF---GDIGDTINLPGRPGLKGERGTTGIP-GLKGFFGEKGTEGDI
GFPGITGVTGVQGPPGLKGQTGFP
PF01391 // ENSGT00900000140804_1_Domain_17 (1765, 1856)
GLTGPPGSQGEPGRIGLPG--GKGDDGWPGAPGLP-
--------------------------------GFPGLRGIRGLHGLPGTKGFPGSPGSDI
HG----------D---PGFPGPPGERGDPGEANTLPGPVGVPGQKGDQGAP---------
--------------------------GERGPPGSPGLQGFPGI-TPPSNIS
PF01391 // ENSGT00900000140804_1_Domain_19 (1972, 2034)
GAPGDKGA-
---PGIFGLKGYRGPPGPPGSAALPGSKGDTGNPGAPGTPGTKGWAGDSGPQGRPGVFGL
PGEKG---------PRGEQGFMGNTGPT-----GAVGDRGPKGPKGDPGFPG-------A
PGTVGAPGIAGIPQKIAVQPGTVGPQGRRGPPGAPGEMGPQGPPGEP-------------
---------------------------------------GFRGAPGKAGPQGRGGVSAVP
GFRGDEGPIGHQGPIGQE----GAPGRPGSPGLPGMPGRSVSIGY
PF01413 // ENSGT00900000140804_1_Domain_23 (2266, 2371)
LLVKHSQTDQEPMCP
VGMNKLWSGYSLLYFEGQEKAHNQDLGLAGSCLARFSTMPFLYCNPGDVCYYASRNDKSY
WLSTTAPLPMMPVAEDE--IKPYISRCSVCENSGT00900000140804_1_Linker_23_25 (2371, 2376)
EAPAVPF01413 // ENSGT00900000140804_1_Domain_25 (2376, 2504)
AIAVHSQ--DVSIPHCPAGWRSLWI
GYSFLMHTAAGDEGGGQS----LVSPGSCLEDFRAT---PFIECNG-GRGTCHYYANKYS
--FWLTTIPEQSFQGSPSA-----DTLKAGLIRTHISRCQVCMKNL--------------
--------------
ENSCLAP00000003695 (view gene)
------------------------------------------------GVKKFDVPCGGR
DCS-GGCQCYPEKGGR
PF01391 // ENSGT00900000140804_1_Domain_0 (77, 132)
GQPGPVGPQGFSGPPGLQGFPGLQGRKGDKGERGAPGTPGPKGD
VGPRGVSGFPGADGIP---GHPGQGGSRGRPGYDGCNGTRGDSGPQGPSGSEGFPGSPGP
QGPKGQKGEPYALPPEERDRYR
PF01391 // ENSGT00900000140804_1_Domain_1 (203, 251)
GETGEPGLVGFQGPPGRPGPIGPMGPVGAPGRPGPPGP
PGPKGQPGNRENSGT00900000140804_1_Linker_1_2 (251, 321)
GLGF--YGEKGEKGDQGLPGPNGIPSDT-THPAVAPTSSTI--LPYKGEK
GS---QGEPGTKGI----SYPF01391 // ENSGT00900000140804_1_Domain_2 (321, 448)
KGEEGIVGFPGTRGPPGL---DGEKGFPGQ----------
---------------------------------------------------------KGN
RGLDGYQGPD-GAQGLKGESGDPGPPGPPAYSPHP-------------------------
-----------------------------------------ALAKGVRGDPGFQGIYGEP
GSRGVPGDPGPPGL---P--------------------------GLPAGDHG-DRRGLLG
EMGPKGFAGEPGIPARTP
PF01391 // ENSGT00900000140804_1_Domain_5 (619, 747)
GPKGADGRPGLAGPPGPAGPPGPD------------------
-GFLFGLKGTQGSRGYPGHSGFPGMRGQKGWKG---------------------------
-----------------------DTGENSGT00900000140804_1_Linker_5_6 (747, 763)
DC-RCGGDQF--IPLPPF01391 // ENSGT00900000140804_1_Domain_6 (763, 1014)
GLPGLKGFPGANGEPGRK
GEPGDPGQHGIPGFPGFKGTPGSVGA----------------------------------
------------------------------------------------------------
------------------------------------------------------------
-----------------------------------------PGP---KG--AKGDSRMVT
TKGEKGQPGVPGLHGMKGDDGTPGRDGLDGFPGL-----PGP-PGDGIKGTPGDAGYPGV
PGAKGVPGEMGPPGLGLPGPKGEQGFPGDAGFPGPPGLPGPPGPQGSPGQTDCD---TGV
KRPIGGDRQESV--QPGCV--GGPK-----------------------GSPGLPGPP---
G----------------APGDKGPRGAPGLSGEDGPRGLKGFPGDPGREGFPGPP-GFMG
PRGSKGAVGLPGPDGFQGPNGLPGPAGAPGDRGIPGEVLGAQPGPRGDAGLPGHPGLKGV
PGEQGAPGFRGS-----QGMPGMPGLKGQLGFPGPSGQQGLPGPPGLHGF-PGAPGPEGP
LGLPGVP-GR--------------------EGLPGDR
PF01391 // ENSGT00900000140804_1_Domain_12 (1418, 1482)
GDPGDTGAPGPVGMKGLAGDRGD
AGAAGDRGQPGTPGFKGMVGMPGSPGRK-------GDRGSPGMDGFQGMIGLKGRPGLPG
TKGEAGLSGL----PGVRGPPGQRGGKGTRGDPGPPGPPPVILPG-MKYI
PF01391 // ENSGT00900000140804_1_Domain_14 (1551, 1619)
KGEKGDEGPM
GLKGHLGLK-------------GVQGMPGVPGQAGVP-GLPGRPGYIKGI-KGDIGLPGS
PGLPGFPGVVGPPGIIGFPGFTGSRGDKGTQGR---------------------------
---TGLYGEVGQPGDF---GDIGDTINLPGSPGLKGERGVTGTP-GLKGFFGERGTQGEV
GFPGITGLAGPQGPPGIKGQTGFPGLTGLQGPQGDPGQIGFPG--QKGDDGWPGMPGLP-
--------------------------------GFPGLRGISGLHGLPGPKGSPGSPGADM
PG----------D---PGFPGPIGDRGDPGEPNSSPGPTGVPGQKGEQGAP---------
--------------------------GERGPAGRPGIQGFPGI-SPASNISGLP
PF01391 // ENSGT00900000140804_1_Domain_19 (1975, 2036)
GDTGS-
---PGIFGLKGYPGPQGPPGPAALPGSKGDTGSAGAPGVPGTKGWVGDPGPQGRPGVFGV
PGEKG---------PRGEPGLMGNTGPT-----GAVGDRGPKGPKGDQGFPG-------A
PGSVGSPGIAGIPQRISAPAGTMGPEGRRGSPGAQGEMGPQGPPGEP-------------
---------------------------------------GFRGVPGKPGPQGRGGVSALP
GFRGDPGPVGQQGPIGQE----GEPGRPGSPGLPGMPGRSVSIGY
PF01413 // ENSGT00900000140804_1_Domain_23 (2266, 2371)
LLVKHSQTDQEPMCP
VGMNRLWSGYSLLYFEGQEKAHNQDLGLAGSCLARFSTMPFLYCNPGDVCYYASRNDKSY
WLSTTAPLPMMPVAEDD--IKPYISRCSVCENSGT00900000140804_1_Linker_23_25 (2371, 2376)
EAPAVPF01413 // ENSGT00900000140804_1_Domain_25 (2376, 2504)
AIAVHSQ--DVSIPHCPAGWRSLWI
GYSFLMHTAAGDEGGGQS----LVSPGSCLEDFRAT---PFIECNG-GRGTCHYFANKYS
--FWLTTIPEASFQGSPSA-----DTLKAGLIRTHISRCQVCMKNL--------------
--------------
ENSETEP00000000138 (view gene)
ENSPPYP00000006266 (view gene)
ENSMLUP00000016067 (view gene)
ENSMUSP00000033899 (view gene)
--------------MDRVR-FKASGPPLRGWLLLATVTVGLLAQSVLGGVKKLDVPCGGR
DCS-GGCQCYPEKGARGQPGAVGPQGYNGPPGLQGFPGLQGRKGDKGERGVPGPTGPKGD
VGARGVSGFP
PF01391 // ENSGT00900000140804_1_Domain_0 (131, 189)
GADGIP---GHPGQGGPRGRPGYDGCNGTRGDAGPQGPSGSGGFPGLPGP
QGPKGQKGEPYALSKEDRDKYRGEPGEPGLVGYQGPPGRPGPIGQMGPMGAPGRPGPPGP
PGPKGQPGNRGLGF--YGQKGEKGDIGQPGPNGIPSDI---TLVGPTTSTIHPDLYKGEK
GDEG---EQGIPGV----IS
PF01391 // ENSGT00900000140804_1_Domain_2 (321, 448)
KGEEGIMGFPGIRGFPGL---DGEKGVVGQ----------
---------------------------------------------------------KGS
RGLDGFQGPS-GPRGPKGERGEQGPPGPSVYSPHP-------------------------
-----------------------------------------SLAKGARGDPGFQGAHGEP
GSRGEPGEPGTAGP---P--------------------------GPSVGDED-SMRGLPG
EMGPKGFSGEPGSPARYLGPPGADGRPGPQGVPGPAGPPGPD------------------
-GFLFGLKGSEGRVGYPGPSGFPGTRGQKGWKG---------------------------
-----------------------EAGDC-QCG---QVIG
PF01391 // ENSGT00900000140804_1_Domain_6 (760, 1014)
GLPGLPGPKGFPGVNGELGKK
GDQGDPGLHGIPGFPGFKGAPGVAGA----------------------------------
------------------------------------------------------------
------------------------------------------------------------
-----------------------------------------PGP---KG--IKGDSRTIT
TKGERGQPGIPGVHGMKGDDGVPGRDGLDGFPGL-----PGP-PGDG
PF01391 // ENSGT00900000140804_1_Domain_9 (1068, 1127)
IKGPPGDAGLPGV
PGTKGFPGDIGPPGQGLPGPKGERGFPGDAGLPGPPGFPGPPGPPGTPGQRDCD---TGV
KRPIGGGQQVVV--QPGCI--EGPT-----------------------GSPGQPGPP---
G----------------PTGAKGVRGMPGFPGASGEQGLKGFPGDPGREGFPGPP-GFMG
PRGSKGTTGLPGPDGPPGPIGLPGPAGPPGDRGIPGEVLGAQPGTRGDAGLPGQPGLKGL
PGETGAPGFRGS-----QGMPGMPGLKGQPGFPGPSGQPGQSGPPGQHGF-PGTPGREGP
LGQPGSP-GL--------------------GGLPGDR
PF01391 // ENSGT00900000140804_1_Domain_12 (1418, 1482)
GEPGDPGVPGPVGMKGLSGDRGD
AGMSGERGHPGSPGFKGMAGMPGIPGQK-------GDRGSPGMDGFQGMLGLKGRQGFPG
TKGEAGFFGV----PGLKGLPGEPGVKGNRGDRGPPGPPPLILPG-MKD
PF01391 // ENSGT00900000140804_1_Domain_14 (1550, 1619)
IKGEKGDEGPM
GLKGYLGLK-------------GIQGMPGVPGVSGFP-GLPGRPGFIKGV-KGDIGVPGT
PGLPGFPGVSGPPGITGFPGFTGSRGEKGTPGV---------------------------
---AGVFGETGPTGDF---GDIGDTVDLPGSPGLKGERGITGIP-GLKGFFGEKGAAGDI
GFPGITGMAGAQGSPGLKGQTGFPGLTGLQGPQGEPGRIGIPG--DKGDFGWPGVPGLP-
--------------------------------GFPGIRGISGLHGLPGTKGFPGSPGVDA
HG----------D---PGFPGPTGDRGDRGEANTLPGPVGVPGQKGERGTP---------
--------------------------GERGPAGSPGLQGFPGI-SPPSNISGSPGDV
PF01391 // ENSGT00900000140804_1_Domain_19 (1978, 2035)
GA-
---PGIFGLQGYQGPPGPPGPNALPGIKGDEGSSGAAGFPGQKGWVGDPGPQGQPGVLGL
PGEKG---------PKGEQGFMGNTGPS-----GAVGDRGPKGPKGDQGFPG-------A
PGSMGSPGIPGIPQKIAVQPGTLGPQGRRGLPGALGEIGPQGPPGDP-------------
---------------------------------------GFRGAPGKAGPQGRGGVSAVP
GFRGDQGPMGHQGPVGQE----GEPGRPGSPGLPGMPGRSVSIGY
PF01413 // ENSGT00900000140804_1_Domain_23 (2266, 2371)
LLVKHSQTDQEPMCP
VGMNKLWSGYSLLYFEGQEKAHNQDLGLAGSCLARFSTMPFLYCNPGDVCYYASRNDKSY
WLSTTAPLPMMPVAEEE--IKPYISRCSVCENSGT00900000140804_1_Linker_23_25 (2371, 2376)
EAPAVPF01413 // ENSGT00900000140804_1_Domain_25 (2376, 2504)
AIAVHSQ--DTSIPHCPAGWRSLWI
GYSFLMHTAAGDEGGGQS----LVSPGSCLEDFRAT---PFIECNG-GRGTCHYFANKYS
--FWLTTIPEQNFQSTPSA-----DTLKAGLIRTHISRCQVCMKNL--------------
--------------
ENSTTRP00000005937 (view gene)
ENSTGUP00000009048 (view gene)
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
-PGAPGKPI----------PGK------PGAPL---SPFS-P---FCPGNPGVPG-----
-EPGTPGRPFSPGVPSKGKRKMAGPGGPGNPLGPGRPTPGPGGPGNPGGPGKPSLPGIPG
EGN----PGEPF-SPFSPLSPGTPGLLGSQGFTGPPGLMGIP------------------
-----GVQGPKGHKG---ERGYPGITGPKGEVG---------------------------
-------------------------------------HPGQPGPRGK---PGHDGCNGTA
GDPGDSGTPGLSGFPGRSPAFFAGPPGIPGKDGSP-------------------------
------------------------------------------------------------
---------------------G--------------------------------------
-------------------------------------IRGPPGPIGAPGKKGINL
PF01391 // ENSGT00900000140804_1_Domain_7 (1016, 1095)
QGSTG
PRGPPGFPGTPGQPGIKGEPGD-----------------------------QGPHGLPGY
PGAKGFPGEAGLPGEGFPGPKGFRGPP------------GLPGLPGEPG-----------
------------------------------------------------------------
-----------------QLGAKGARGFPGDPGPMGFPGLNGTRGDPGREGFPG-------
-PPDRGPNGLPGLQGHPGLTGKSGAPGAPGPKGLP--------GSRGDVGLPGFPGLKGA
PGDQGVPGTR-----------------GLPGPKGDPGPQGLPGTPGTHGF-PGPPGNRGP
DG------------------------------PPGARGEDGEQGFPGPVGLKGLSGDKGD
MGHTGLPGSKGMPGLPGKTGTPGSPGHPSYVAGVKGDIGAKGLT
PF01391 // ENSGT00900000140804_1_Domain_13 (1485, 1540)
GVKGYPGPAGSPGIRG
FPGTTGVRVSQKGAPGWPGLPGQAGIYGVPGYPGPQGQPGIPAPP------GSKGESGEAGRTGEIGPK-------------GS---------------------------RGDPGLSGRPGLPGFPGPKGF------------QGPPGSPG---------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------
PF01391 // ENSGT00900000140804_1_Domain_14 (1537, 1652)
GQPGIPAPP------GSKGESGEA
GRTGEIGPK-------------GS---------------------------RGDPGLSGR
PGLPGFPGPKGF------------QGPPGSPG----------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
--------------
ENSCANP00000013008 (view gene)
--------------MGRDQ-RAVAGPALRRWLLLATVTVGFLAQGVLAGVKKFDVPCGGR
DCS-GGCQCYPEKGGRGQPGPVGPQGYNGPPGLQGFPGLQGRKGDKGERGAPGITGPKGD
VGARGVSGFP
PF01391 // ENSGT00900000140804_1_Domain_0 (131, 189)
GADGIP---GHPGQGGPRGRPGYDGCNGTQGDSGPQGPPGSEGFTGPPGP
QGPKGQKGENSGT00900000140804_1_Linker_0_1 (189, 203)
EPYALPREERDRYRPF01391 // ENSGT00900000140804_1_Domain_1 (203, 251)
GEPGEPGLVGFQGPPGRPGPVGQMGPVGAPGRPGPPGP
PGPKGQQGNRENSGT00900000140804_1_Linker_1_2 (251, 322)
GLGF--YGVKGEKGDVGQPGPNGIPSDT-LHPIVAPTGVTFHPDQYKGEK
GS---EGEPGIRGI----SLKPF01391 // ENSGT00900000140804_1_Domain_2 (322, 447)
GEEGIMGFPGLRGYPGL---SGEKGSPGQ----------
---------------------------------------------------------KGS
RGLDGYQGPD-GPRGPKGEAGDLGPPGLPAYSPHP-------------------------
-----------------------------------------SLAKGARGDPGFPGAQGEP
GSQGEPGDPGPRGP---P--------------------------GLFIGDED-QRRGLPG
EMGPKGFIGDPGIPALYP
PF01391 // ENSGT00900000140804_1_Domain_5 (619, 747)
GPPGPDGRPGPPGPPGPPGPPGPD------------------
-GFLFGLKGAEGRPGFPGLSGSPGARGPKGWKG---------------------------
-----------------------DAGENSGT00900000140804_1_Linker_5_6 (747, 760)
DC-RCAEGDEPVRPF01391 // ENSGT00900000140804_1_Domain_6 (760, 1014)
GPPGLPGPKGFAGINGEPGRK
GDKGDPGQHGLPGFPGLKGVPGNVGA----------------------------------
------------------------------------------------------------
------------------------------------------------------------
-----------------------------------------PGP---KG--AKGDSRTIT
TKGERGQPGVPGVPGMKGDDGSPGRDGLDGFPGL-----PGP-PGDGIRGPPGDAGYPGT
PGTKGTPGEVGPPGLGLPGFKGQRGFPGDAGLPGPPGFPGPPGPAGTPGQRDCD---TDE
KRPIGGDRQEAI--QPGCV--GGPK-----------------------GLPGLPGPP---
G----------------PSGAKGLRGIPGFSGADGGPGPKGLPGDPGREGFPGPP-GFIG
PRGSKGAVGLPGPDGPPGPIGLPGPDGPPGDRGLPGEVLGAQPGPRGDAGVPGQPGFKGL
PGDRGAPGFRGS-----QGMPGMPGLK
PF01391 // ENSGT00900000140804_1_Domain_12 (1348, 1427)
GQPGLPGPSGQPGLSGPPGQHGF-PGAPGQEGP
LGLPGIP-GL--------------------EGLPGDRGDPGDIGAPGPVGMKGLSGDRGD
AGLAGERGHPGSPGFKGIDGMPGTPGLK-------GERGSPGMDGFQGMPGLKGRPGFPG
SKGEAGFFGI----PGLKGLAGEPGFKGSRGDPGPPGPPPVILPG-MKDIKGEKGDEGPM
GLKGYLGAK-------------GIPGMPGIPGLSGIP-GLPGRPGHIKGV-KGDIGAPGI
PGLPGFPGVAGPPGITGFPGFIGSRGDKGAPGR---------------------------
---AGLYGEIGPTGDF---GDIGDTINLP
PF01391 // ENSGT00900000140804_1_Domain_17 (1710, 1767)
GRPGLKGEQGTAGIP-GLKGFFGEKGTEGDI
GFPGITGVTGVQGPPGLKGQTGFPGLTGPPGSQGEPGRIGLPG--GKGDDGWPGTPGLP-
--------------------------------GFPGLRGIRGLHGLPGTKGFPGSPGADI
HG----------D---PGFPGPPGDRGGPGDANTLPGPVGVPGQKGEPGAP---------
--------------------------GERGPPGSPGLQGFPGI-TPPSNIS
PF01391 // ENSGT00900000140804_1_Domain_19 (1972, 2034)
GVPGDKGA-
---PGIFGLKGYRGPPGPPGSAALPGSKGDTGNPGAPGTPGTKGWAGDSGPQGRPGVFGL
PGEKG---------PRGEQGFMGNTGPA-----GAVGNRGPKGPKGDPGFPG-------A
PGIVGAPGIAGIPQKIAIPPGTVGPQGRRGPPGAQGEMGPQGPPGEP-------------
---------------------------------------GFRGAPGKAGPQGRGGVSAVP
GFRGDEGPIGHQGPTGQE----GAPGRPGSPGLPGMPGRSVSIGY
PF01413 // ENSGT00900000140804_1_Domain_23 (2266, 2371)
LLVKHSQTDQEPMCP
VGMNKLWSGYSLLYFEGQEKAHNQDLGLAGSCLARFSTMPFLYCNPGDVCYYASRNDKSY
WLSTTAPLPMMPVAEDE--IKPYISRCSVCENSGT00900000140804_1_Linker_23_25 (2371, 2376)
EAPAVPF01413 // ENSGT00900000140804_1_Domain_25 (2376, 2504)
AIAVHSQ--DVSIPHCPAGWRSLWI
GYSFLMHTAAGDEGGGQS----LVSPGSCLEDFRAT---PFIECNG-GRGTCHYYANKYS
--FWLTTIPEQSFQGSPSA-----DTLKAGLIRTHISRCQVCMKNL--------------
--------------
ENSECAP00000018971 (view gene)
ENSOCUP00000011443 (view gene)
ENSFALP00000011009 (view gene)
ENSCAPP00000010546 (view gene)
ENSOPRP00000012047 (view gene)
--------------MARER-LAAPGPALQWWLLLGTVTMGLLWQSALA-NKKLDVPCGGR
DCS-GGCQCYPEKGGRGQPGPV
PF01391 // ENSGT00900000140804_1_Domain_0 (83, 145)
GP-GYTGPPGLQGFAGLQGRKGDKGERGAPGITGPKGD
VGARGVSGFPGADGIP---XXXXXXGPRGRPGYDGCNGTKGR-GLQGPPG-DGFPGSPGP
QGPKGQKGEPYTLPEEERGRY
PF01391 // ENSGT00900000140804_1_Domain_1 (202, 237)
RGETGEPGLVGIQGPPGRPGPMGPMGPVGAAGQPGPLSS
LENXXXXXXXXXXXXXXXX------XXX--XXXXXXXX-XXXXXXXXXXXXXXXXXXXXX
XXX---XXXXXXXX-XXXXXXXXXXXXXXXXXXXXXXX---XXXXXXX-X----------
---------------------------------------------------------X-X
XXXXXXXXXX------XXXXXXXXXXXXXXXXXXXXXX----------------------
------------------------------------XXXXXXXXXX
PF01391 // ENSGT00900000140804_1_Domain_3 (527, 587)
ARGDPGFPGAHGEP
GSRGEPGDPGPPGP---P--------------------------GRPARGED-EMRGLPG
EMGPKGFIGEPGIPASFSDPPGADGRPGLPGAPGSSGPPWTDXXXXXXXXXXXXXXX---
-----------------XXXXXXXXXXXXXXXX---------------------------
-----------------------EAGDC-KCLEGDQYVG
PF01391 // ENSGT00900000140804_1_Domain_6 (760, 1019)
GLPGLPGPKGLPGFNGEPGRK
GDQGDPGQHGIPGFPGLKGTPGLPGV----------------------------------
------------------------------------------------------------
------------------------------------------------------------
-----------------------------------------PGP---KG--AKGDSRTENSGT00900000140804_1_Linker_6_7 (1019, 1023)
VS
TKPF01391 // ENSGT00900000140804_1_Domain_7 (1023, 1058)
GERGQPGVPGVPGMKGDDGVPGHDGLDGFPGLPGPPXXXXXXXXXXXXXXXXXXXX--
---X-XXXXXX---XXXXX-XXXXXXXXXXXXXXX-XXXXXXXXXXXXXXXXXXXXXXXX
XXXXXXXXXXXXXXXXXCV--AGPK-----------------------GLPGPPGPP---
G----------------PAGAKGLQGTPGFSGADGPPGLKGLPGDPGREGYPGPPXXXXX
XXXXXXXXXXXXXXXXXXXXXXX-------XXXXXXXXXXXXX-XXXXXXXXXXXXXXXX
XXXXXXXXXXXX-----XXXXXX--XXXX-----------X-XXXXXXXX-XXXXXXXXX
XXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXX
PF01391 // ENSGT00900000140804_1_Domain_12 (1413, 1453)
LPGERGDPGDTGVPGPVGMKGLAGDRGD
AGYSGERGQPGSPGFKGMVGMPGIPGPK-------GERGSPGMDGFQGMLGLKGRPGLPG
IKGETGFFGR----TGEKGPAGEPGDKXXXXXXXXXXXXXXX-XXXXXXXXXXXXXX
PF01391 // ENSGT00900000140804_1_Domain_14 (1558, 1605)
GPM
GLKGYLGLK-------------GIQGMPGVPGLAGIP-GLPGRPGHIKGV-KGDIGVPGV
PGLPGFPGVAGPPGIIGFPGFTGSRGDKGAPGR---------------------------
---AGLYGEVGQTGDL---GDVGDTIDLPGSPGLKGEQGTAGVP-GVKGFFGEKGAEGEV
GFPGITGLTGAQGPPGLKGQTGFPGLTGLQGPQGDPGQIGRPG--DKGDQGWPGRPGLP-
--------------------------------GFPGLRGIGGLHGLPGTKGLQGSPGADI
HG----------D---PGFPGPTGDRGDPGEPNTLPGPVGVPGHK
PF01391 // ENSGT00900000140804_1_Domain_18 (1906, 2005)
GEQGVP---------
--------------------------GERGPAGSPGLQGFPGI-TPPSNISGLPGDKGA-
---AGIFGPQGYPGLPGPPGPAALPGSKGEDGSPGTAGIPGSKGWVGXXXXXXXXXXXX-
--------XXXXXXPRGEQGFMGNTGAT-----GAAGDRGPKGPKGDQGFPG-------A
PGPVGSPGIVGLPQRVAVQHGPVGPQGRRGPPGAQGEM
PF01391 // ENSGT00900000140804_1_Domain_21 (2139, 2254)
GPQGPPGEP-------------
---------------------------------------GLRGAPGKAGPQGRGGVSAVP
GIRGDQGPMGHPGPVGQE----GEPGRPGSPGLENSGT00900000140804_1_Linker_21_23 (2254, 2264)
PGMPGRSVSIPF01413 // ENSGT00900000140804_1_Domain_23 (2264, 2373)
GYLLVKHSQTDQEPMCP
TGMNKLWSGYSLLYFEGQEKAHNQDLGLAGSCLARFSTMPFLYCNPGDICYYASRNDKSY
WLSTTAPLPMMPVAEEE--IRPYISRCSVCEAENSGT00900000140804_1_Linker_23_24 (2373, 2374)
PPF01413 // ENSGT00900000140804_1_Domain_24 (2374, 2406)
AVAIAVHSQ--DVSIPHCPAGWRSLWI
GYSFLM------------------------------------------------------
------------------------------------------------------------
--------------
ENSMICP00000040801 (view gene)
--------------MGRDR-RAASGPALQRWLLLGTVTVGVLAQSALAGVKKLDVPCGGR
DCS-GGCQCYPEKGGRGQPGPVGPQGYNGPPGLQGFPGLQGRKGDKGERGAPGITGPKGD
VGARGVSGFP
PF01391 // ENSGT00900000140804_1_Domain_0 (131, 190)
GADGIP---GHPGQGGPRGRPGYDGCNGTRGDAGPQGPYGSDGFPGPTGP
PGPKGQKGEENSGT00900000140804_1_Linker_0_1 (190, 203)
PYALSREDRDRYRPF01391 // ENSGT00900000140804_1_Domain_1 (203, 251)
GEPGEPGLVGYQGPPGRPGPIGPMGPVGAAGRPGPPGP
PGPKGEPGNRENSGT00900000140804_1_Linker_1_2 (251, 321)
GLGF--HGQKGEKGDVGLPGPNGIPSDH-YPPSVGPTIS-YISDMYKGEK
GS---EGDPGIKGI----SVPF01391 // ENSGT00900000140804_1_Domain_2 (321, 447)
KGEEGIVGFSGMRGSPGF---DGEKGSPGQ----------
---------------------------------------------------------KGS
RGLDGFQGPD-GPRGPKGEAGDRGPPGLPAYSPHP-------------------------
-----------------------------------------SLAKGARGDPGFPGAHGEP
GSQGEPGDPGPPGL---P--------------------------GTSIIDED-GKRGLPG
EMGPKGFIGDPGIPARYPGPPGGDGKPGLRGPPGPAGEPGPD------------------
-GFLFGLKGTEGRVGFPGHSGSPGTRGQKGWKG---------------------------
-----------------------DAGDCRCAAGDDQFVT
PF01391 // ENSGT00900000140804_1_Domain_6 (760, 1015)
GVPGLPGPKGFPGLNGDPGRK
GEPGDPGQHGIPGFPGFKGAPGNVGV----------------------------------
------------------------------------------------------------
------------------------------------------------------------
-----------------------------------------PGP---KG--AKGDSRTIT
TKGERGQPGVPGVPGIKGDDGLPGREGLDGFPGL-----PGP-PGDGIPGPPGDQGYPGA
PGTKGVPGERGSPGQGLPGPKGERGFPGDTGLPGPSGFPGPPGPPGTPGQIDCD---TGV
KRPIGGDGQEAV--QPGCI--GGPK-----------------------GLPGLPGSP---
G----------------LTGAKGLRGIPGFPGADGEPGLKGFAGDPGREGFPGPP-GFVG
PRGSKGAAGLPGPDGPPGPTGLPGPAGPPGDRGIPGEVLGAQPGPRGDAGLPGHPGLKGL
PGDRGPPGFRGS-----QGMPGMPGLKGHLGFPGPSGQPGLPGPPGLHGF-PGNPGQEGP
LGQPGAP-GL--------------------GGLPGDR
PF01391 // ENSGT00900000140804_1_Domain_12 (1418, 1482)
GEPGDTGVPGPVGMKGFSGDRGD
AGQSGDRGHPGSPGFKGMLGMPGPPGLK-------GDRGSPGMDGFQGMPGLKGRPGLPG
SKGESGFFGI----PGLKGLAGEPGVKGSRGDPGPPGPPPIILPG-MKDMKGEKGDEGPM
GLKGYLGLK-------------GTQGMPGIPGLSGTP-GLPGRPGYIKGV-KGDIGVPGV
PGLPGLPGVAGPPGITGFPGFTGSRGDKGAPGS---------------------------
---AGLYGKTGPTGDF---GDTGDTVDLPGRPGLKGEQGSAGMP-GLKGFFGEKGTEGSI
GFPGIPGAPGVQGPPGLKGQTGFP
PF01391 // ENSGT00900000140804_1_Domain_17 (1765, 1857)
GLTGLQGPQGEPGQVGPPG--SKGDDGWPGVPGLP-
--------------------------------GFPGSRGISGLHGLPGTKGLLGSPGADI
YG----------D---PGFQGPPGDRGDPGEANTFPGPVGAPGQKGEQGAP---------
--------------------------GQRGPAGSPGLQGFPGI-TPPSNISGSPGDKGA-
---PGIFGLEGYRGPPGPPGPAALPGGKGDAGNPGAPGVPGTKGWVGDP
PF01391 // ENSGT00900000140804_1_Domain_20 (2030, 2108)
GPQGRPGVFGL
PGEKG---------PRGEQGLMGSTGAS-----GTVGDRGPKGPKGDQGFPG-------A
PGPAGSPGIAGVPPRIAVQPGILGPQGRRGLPGAQGEMGPQGPPGDP-------------
---------------------------------------GLRGDPGKAGPQGRGGVSAVP
GFRGDQGPVGHQGPTGQE----GEPGRPGSPGLPGMPGRSVSIGY
PF01413 // ENSGT00900000140804_1_Domain_23 (2266, 2371)
LLVKHSQTDQEPMCP
VGMNRLWSGYSLLYFEGQEKAHNQDLGLAGSCLARFSTMPFLYCNPGDVCYYASRNDKSY
WLSTTAPLPMMPVAEDD--IKPYISRCSVCENSGT00900000140804_1_Linker_23_25 (2371, 2376)
EAPAVPF01413 // ENSGT00900000140804_1_Domain_25 (2376, 2504)
AIAVHSQ--DVSIPHCPPGWRSLWI
GYSFLMHTAAGDEGGGQS----LVSPGSCLEDFRAT---PFIECNG-GRGTCHYYANKYS
--FWLTTIPEQSFQGSPSA-----DTLKAGLIRTHISRCQVCMKNL--------------
--------------
ENSP00000353654 (view gene)
ENSCGRP00001022430 (view gene)
--------------MDRVK-FSVSGPALRGWLLLATVTVGLLAQSVLAGVKKLDVPCGGR
DCS-GGCQCYPEKGAR
PF01391 // ENSGT00900000140804_1_Domain_0 (77, 132)
GQPGPVGTQGYNGPPGLQGFPGLQGRKGDKGERGAPGPTGPKGD
VGARGVSGFPGADGIP---GHPGQGGPRGRPGYDGCNGTRGDAGPQGPSGSGGFYGSPGP
QGPKGQKGEPYVLRKEDQDKYR
PF01391 // ENSGT00900000140804_1_Domain_1 (203, 251)
GEPGEPGQIGYQGPPGRLGLIGQMGPMGAPGRPGPPGP
PGPKGQPGNRENSGT00900000140804_1_Linker_1_2 (251, 322)
GLGF--YGKKGEKGDVGQPGPNGIPSDI---TLIGPTTTTYRPDQYKGEK
GSRG---DQGIPGP----SLKPF01391 // ENSGT00900000140804_1_Domain_2 (322, 448)
GEEGIMGFPGTRGHPGI---DGEKGIFGQ----------
---------------------------------------------------------KGS
RGLDGFQGPS-GPRGPKGERGQQGPPGPPVYSPHP-------------------------
-----------------------------------------SLAKGARGDPGFPGAHGEP
GSRGEPGDPGPMGP---P--------------------------GPSIGDED-TKRGLPG
EMGPKGFSGEPGHPAPYPGPPGADGKPGLQGVPGPAGPPGPD------------------
-GFLFGLKGSEGRVGYPGPSGFPGARGQKGWKG---------------------------
-----------------------EAGDC-QCG---QVIG
PF01391 // ENSGT00900000140804_1_Domain_6 (760, 1015)
GLPGLPGPKGSPGVNGELGKK
GDQGDPGLHGIPGFPGFKGAPGPTGI----------------------------------
------------------------------------------------------------
------------------------------------------------------------
-----------------------------------------PGP---KG--IKGDSRTVT
TKGERGQPGVPGVHGMKGDDGVPGRDGLDGFPGL-----PGP-PGDGIKGPPGDAGSPGL
PGTKGFPGEVGPPGQGLPGPKGERGFPGDAGFPGSPGYPGPPGPPGAPGQLDCD---TGV
KRPIGGGQQVII--QPGCG--EGPT-----------------------GSPGNPGPP---
G----------------PTGVKGIRGMPGFPGDTGAPGLKGFPGDPGREGFPGPP-GFMG
PRGSKGATGLPGPDGLPGPTGLPGPAGPPGDRGIPGEVLGAQPGPRGDAGLPGHPGLKGP
PGERGPPGVTGS-----QGMPGIPGLKGQPGFPGPSGQPGLPGPPGQDGF-PGAPGREGP
LGLPGSL-GL--------------------RGLPGDRGEPGEPGVPGPVGMKGLSGDRGD
VGISGERGHPGSPGIKGVAGMPGNPGQK-------
PF01391 // ENSGT00900000140804_1_Domain_13 (1476, 1538)
GDRGSPGMDGFQGMLGLKGRPGFPG
VKGEAGFFGV----PGLKGMPGEPGVKGNRGDRGPPGPPPVILPG-MKDIKGEKGDEGPM
GLKGYLGLK-------------GIQGMPGVPGLAGIP-GLPGRPGYIKGV-KGDIGVPGT
PGLPGQPGVVGTPGITGFPGFIGSRGEKGAPGR---------------------------
---AGLFGEIGLTGDF---GDIGDTVDLPGSPGLKGERGVTGIP-GLKGILGEKGTEGEV
GFPGITGMAGPQGSPGLKGQTGF
PF01391 // ENSGT00900000140804_1_Domain_17 (1764, 1855)
PGLTGLQGPQGDPGRSGVPG--DKGDYGWPGVSGLP-
--------------------------------GFPGIRGISGLHGLPGTKGFPGSPGADV
HG----------D---PGFPGPAGDRGDRGEANTLPGPVGAAGQKGERGTP---------
--------------------------GERGPAGSPGLQGFPGI-SPLSNISGSPGDVGA-
---PGIFGSEGYRGPPGPPGPNALPGIKGDEGSVGAAGFPGQKGWV
PF01391 // ENSGT00900000140804_1_Domain_20 (2027, 2106)
GDPGLQGQPGVVGL
PGEKG---------PKGEQGFMGNTGPS-----GAVGDRGPKGPKGDQGFPG-------A
PGSVGSPGIPGIPQKIAIQPGTMGPQGRRGLPGAQGEMGPQGPPGDP-------------
---------------------------------------GFRGAPGKTGPQGRGGVSAVP
GFRGDQGPMGQQGPIGQE----GEPGRPGSPGLPGMPGRSVSIGY
PF01413 // ENSGT00900000140804_1_Domain_23 (2266, 2371)
LLVKHSQTDQEPMCP
VGMNKLWSGYSLLYFEGQEKAHNQDLGLAGSCLARFSTMPFLYCNPGDVCYYASRNDKSY
WLSTTAPLPMMPVAEEE--IKPYISRCSVCENSGT00900000140804_1_Linker_23_25 (2371, 2376)
EAPAVPF01413 // ENSGT00900000140804_1_Domain_25 (2376, 2504)
AIAVHSQ--DVSIPHCPAGWRSLWI
GYSFLMHTAAGDEGGGQS----LVSPGSCLEDFRAT---PFIECNG-GRGTCHYFANKYS
--FWLTTIPEQSFQGTPSA-----DTLKAGLIRTHISRCQVCMKNL--------------
--------------
ENSCSAP00000013500 (view gene)
--------------MGRDQ-RAVAGPALRRWLLLGTVTVGFLAQGVLAGVKKFDVPCGGR
DCS-GGCQCYPEKGGRGQPGPVGPQGYNGPPGLQGFPGLQGRKGDKGERGAPGITGPKGD
VGARGVSGFP
PF01391 // ENSGT00900000140804_1_Domain_0 (131, 189)
GADGIP---GHPGQGGPRGRPGYDGCNGTQGDSGPQGPPGSEGFTGPPGP
QGPKGQKGEPYALPREERDRYRGEPGEPGLVGFQGPPGRPGPVGQMGPVGAPGRPGPPGP
PGPKGQQGNRGLGF--YGVKGEKGDVGQPGPNGIPSDL-LHPIIAPTGVTFHPDQYKGEK
GS---EGEPGIRGI----SLK
PF01391 // ENSGT00900000140804_1_Domain_2 (322, 447)
GEEGIMGFPGPRGSPGL---SGEKGSPGQ----------
---------------------------------------------------------KGS
RGLDGYQGPD-GPRGPKGEAGDLGPPENSGT00900000140804_1_Linker_2_3 (447, 525)
GLPAYSPHP-------------------------
-----------------------------------------SLAPF01391 // ENSGT00900000140804_1_Domain_3 (525, 612)
KGARGDPGFPGAQGEP
GSQGEPGDPGPRGP---P--------------------------GLSIGDAD-QRRGLPG
EMGPKGFIGDPGIPALYPGPPGLDGKPGPPGPPGLPGPPGPD-------------------GFLFGLKGPEGGPGVPGLSGSPGARGPKGWKG--------------------------------------------------DAGDC-RCAEGDEAVRGLPGLPGPKGFAGINGEPGRKGDKGDPGQHGLPGFPGLKGVPGNVGA---------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------PGP---KG--AKGDSRTITTKGERGQPGVPGVPGMKGDDGGPGRDGLDGFPGL-----PGP-PGDGIKGPPGDAGYPGIPGTKGTPGEMGPPGLGLPGFKGQRGFPGDAGLPGPPGFPGPPGPAGTPGQIDCD---TDVKRPIGGDRQEAV--QPGCV--GGPK-----------------------GLPGLPGPP---G----------------PSGAKGLRGIPGFSGADGGPGPKGLPGDPGREGFPGPP-GFIGPRGSKGAVGRPGPDGPPGPIGLPGPDGPPGDRGLPGEVLGAQPGPRGDAGVPGQPGFKGLPGDRGPPGFRGS-----QGMPGMPGLKGQPGLPGPSGQPGLSGPPGQHGF-PGAPGQEGPLGLPGIP-GL--------------------EGLPGDRGDPGDIGAPGPVGMKGLSGDRGDAGLAGERGHPGSPGFKGIDGMPGTPGLK-------GERGSPGMDGFQGMPGLKGRPGFPGSKGEAGFFGI----PGLKGLAGEPGFKGSRGDPGPPGPPPIILPG-MKDIKGEKGDEGPMGLKGYLGAK-------------GIQGMPGIPGLSGIP-GLPGRPGHIKGV-KGDIGAPGIPGLPGFPGVAGPPGITGFPGFIGSRGDKGAPGR------------------------------AGLYGEIGPTGDF---GDIGDTINLPGRPGLKGEQGTAGIP-GLKGFFGEKGTEGDIGFPGITGVTGVQGPPGLKGQTGFPGLTGPPGSQGQPGRIGLPG--SKGDDGWPGTPGLP---------------------------------GFPGLRGIRGLHGLPGTKGFPGSPGADIHG----------D---PGFPGPPGDRGDPGDANTLPGPVGVPGQKGEQGAP-----------------------------------GERGPPGSPGLQGFPGI-TPPSNISGAPGDKGA----PGIFGLKGYRGPPGPPGSAALPGSKGDTGNPGAPGTPGTKGWAGDSGPQGRPGVFGLPGEKG---------PRGEQGFMGNTGPA-----GTVGDRGPKGPKGDPGFPG-------APGIVGAPGIAGIPQKIAIPPGTVGPQGRRGPPGAPGEMGPQGPPGEP----------------------------------------------------GFRGAPGKAGPQGRGGVSAVPGFRGDEGPIGHQGPIGQE----GAPGRPGSPGLPGMPGRSVSIGYLLVKHSQTDQEPMCPVGMNKLWSGYSLLYFEGQEKAHNQDLGLAGSCLARFSTMPFLYCNPGDVCYYASRNDKSYWLSTTAPLPMMPVAEDE--IKPYISRCSVCEAPAVAIAVHSQ--DVSIPHCPAGWRSLWIGYSFLMEKKIQILGKEGR--------DKQYWRFNST---RRRE----TKAAASHWCHRAA--VWRTSAPRHSSNAMEAA-----APATTTPISTASG---------------------
PF01391 // ENSGT00900000140804_1_Domain_5 (596, 662)
RGLPGEMGPKGFIGDPGIPALYPGPPGLDGKPGPPGPPGLPGPPGPD------------------
-ENSGT00900000140804_1_Linker_5_6 (662, 760)
GFLFGLKGPEGGPGVPGLSGSPGARGPKGWKG---------------------------
-----------------------DAGDC-RCAEGDEAVRPF01391 // ENSGT00900000140804_1_Domain_6 (760, 1015)
GLPGLPGPKGFAGINGEPGRK
GDKGDPGQHGLPGFPGLKGVPGNVGA----------------------------------
------------------------------------------------------------
------------------------------------------------------------
-----------------------------------------PGP---KG--AKGDSRTIT
TKGERGQPGVPGVPGMKGDDGGPGRDGLDGFPGL-----PGP-PGDGIKGPPGDAGYPGI
PGTKGTPGEMGPPGLGLPGFKGQRGFPGDAGLPGPPGFPGPPGPAGTPGQIDCD---TDV
KRPIGGDRQEAV--QPGCV--GGPK-----------------------GLPGLPGPP---
G----------------PSGAKGLRGIPGFSGADGGPGPKGLPGDPGREGFPGPP-GFIG
PRGSKGAVGRPGPDGPPGPIGLPGPDGPPGDRGLPGEVLGAQPGPRGDAGVPGQPGFKGL
PGDRGPPGFRGS-----QGMPGMPGLKGQPGLPGPSGQPGLSGPPGQHGF-PGAPGQEGP
LGLPGIP-GL--------------------EGLPGDRGDPGDIGAPGPVGMKGLSGDRGD
AGLAGERGHPGSPGFKGIDGMPGTPGLK-------
PF01391 // ENSGT00900000140804_1_Domain_13 (1476, 1538)
GERGSPGMDGFQGMPGLKGRPGFPG
SKGEAGFFGI----PGLKGLAGEPGFKGSRGDPGPPGPPPIILPG-MKDIKGEKGDEGPM
GLKGYLGAK-------------GIQGMPGIPGLSGIP-GLPGRPGHI
PF01391 // ENSGT00900000140804_1_Domain_15 (1608, 1694)
KGV-KGDIGAPGI
PGLPGFPGVAGPPGITGFPGFIGSRGDKGAPGR---------------------------
---AGLYGEIGPTGDF---GDIGDTINLPGRPGLKGEQGTAGIP-GLKGFFGEKGTEGDI
GFPGITGVTGVQGPPGLKGQTGFPGLTGPPGSQGQPGRIGLPG--SKGDDGWPGTPGLP-
--------------------------------GFPGLRGIRGLHGLPGTKGFPGSPGADI
HG----------D---PGFPGPPGDRGDPGDANTLPGPVGVPGQKGEQGAP---------
--------------------------GERGPPGSPGLQGFPGI-TPPSNISGAPGDKGA-
---PGIFGLKGYRGPPGPPGSAALPGSKGDTGNPGAPGTPGTKGWAGDS
PF01391 // ENSGT00900000140804_1_Domain_20 (2030, 2108)
GPQGRPGVFGL
PGEKG---------PRGEQGFMGNTGPA-----GTVGDRGPKGPKGDPGFPG-------A
PGIVGAPENSGT00900000140804_1_Linker_20_22 (2108, 2203)
GIAGIPQKIAIPPGTVGPQGRRGPPGAPGEMGPQGPPGEP-------------
---------------------------------------GFRPF01391 // ENSGT00900000140804_1_Domain_22 (2203, 2259)
GAPGKAGPQGRGGVSAVP
GFRGDEGPIGHQGPIGQE----GAPGRPGSPGLPGMPGENSGT00900000140804_1_Linker_22_23 (2259, 2265)
RSVSIGPF01413 // ENSGT00900000140804_1_Domain_23 (2265, 2372)
YLLVKHSQTDQEPMCP
VGMNKLWSGYSLLYFEGQEKAHNQDLGLAGSCLARFSTMPFLYCNPGDVCYYASRNDKSY
WLSTTAPLPMMPVAEDE--IKPYISRCSVCEENSGT00900000140804_1_Linker_23_24 (2372, 2375)
APAPF01413 // ENSGT00900000140804_1_Domain_24 (2375, 2409)
VAIAVHSQ--DVSIPHCPAGWRSLWI
GYSFLMEKKIQILGKEGR--------DKQYWRFNST---RRRE----TKAAASHWCHRAA
--VWRTSAPRHSSNAMEAA-----APATTTPISTASG-----------------------
--------------
ENSMNEP00000004318 (view gene)
--------------MERDQ-RAVAGPALRRWLLLGTVTVGFLAQGVLAGVKKFDVPCGGR
DCS-GGCQCYPEKGGRGQPGPVGPQGYNGPPGLQGFPGLQGRKGDKGERGAPGITGPKGD
VGARGVSGFP
PF01391 // ENSGT00900000140804_1_Domain_0 (131, 189)
GADGIP---GHPGQGGPRGRPGYDGCNGTQGDSGPQGPPGSEGFTGPPGP
QGPKGQKGENSGT00900000140804_1_Linker_0_1 (189, 203)
EPYALPREERDRYRPF01391 // ENSGT00900000140804_1_Domain_1 (203, 251)
GEPGEPGLVGFQGPPGRPGPVGQMGPVGAPGRPGPPGP
PGPKGQQGNRENSGT00900000140804_1_Linker_1_2 (251, 322)
GLGF--YGVKGEKGDVGQPGPNGIPSDT-LHPIVAPTGVTFHPDQYKGEK
GS---EGEPGIRGI----SLKPF01391 // ENSGT00900000140804_1_Domain_2 (322, 447)
GEEGIMGFPGPRGPPGL---SGEKGSPGQ----------
---------------------------------------------------------KGS
RGLDGYQGPD-GPRGPKGEAGDLGPPGLPAYSPHP-------------------------
-----------------------------------------SLAKGARGDPGFPGAQGEP
GSQGEPGDPGPRGP---P--------------------------GLSIGDED-QRRGLPG
EMGPKGFIGDPGIPALYP
PF01391 // ENSGT00900000140804_1_Domain_5 (619, 747)
GPPGPDGKPGPPGPPGLPGPPGPD------------------
-GFLFGRKGAKGTPGFPGLSGSPGARGPKGWKG---------------------------
-----------------------DAGENSGT00900000140804_1_Linker_5_6 (747, 760)
DC-RCAEGDEAVRPF01391 // ENSGT00900000140804_1_Domain_6 (760, 1015)
GLPGLPGPKGFAGINGEPGRK
GDKGDPGQHGLPGFPGLKGVPGNVGA----------------------------------
------------------------------------------------------------
------------------------------------------------------------
-----------------------------------------PGP---KG--AKGDSRTIT
TKGERGQPGVPGVPGMKGDDGSPGRNGLDGFPGL-----PGP-PGDGIKGPPGDAGYPGI
PGTKGTPGEMGPPGLGLPGFKGQRGFPGDAGLPGPPGFPGPPGPAGTPGQIDCD---TDV
KRPIGGDRQEAV--QPGCI--GGPK-----------------------GLPGLPGPP---
G----------------PSGAKGLRGIPGFSGADGGPGPKGLPGDPGREGFPGPP-GFIG
PRGSKGAVGLPGPDGPPGPIGLPGPDGPPGDRGLPGEVLGAQPGPRGDAGVPGQPGFKGL
PGDRGPPGFRGS-----QGMPGMPGLKGQPGLPGPSGQPGLSGPPGQHGF-PGAPGQEGP
LGLPGIP-GL--------------------EGLPGDRGDPGDIGAPGPVGMKGVSGDRGD
AGLAGERGHPGSPGFKGIDGMPGTPGLK-------
PF01391 // ENSGT00900000140804_1_Domain_13 (1476, 1538)
GERGSPGMDGFQGMPGLKGRPGFPG
SKGEAGFFGI----PGLKGLAGEPGFKGSRGDPGPPGPPPIILPG-MKDIKGEKGDEGPM
GLKGYLGAK-------------GIQGMPGIPGLSGIP-GLPGRPGHIKGV-KGDIGAPGI
PGLPGFPGVAGPPGITGFPGFIGSRGDKGAPGR---------------------------
---AGLYGEIGPTGDF---GDIGDTINLP
PF01391 // ENSGT00900000140804_1_Domain_17 (1710, 1765)
GRPGLKGEQGTAGIP-GLKGFFGEKGTEGDI
GFPGITGVTGVQGPPGLKGQTGFPGLTGPPGSQGEPGRIGLPG--GKGDDGWPGIPGLP-
--------------------------------GFPGLRGIRGLHGLPGTKGFPGSPGADI
HG----------D---PGFPGPPGDRGDPGDANTLPGPVGVPGQKGEQGAP---------
--------------------------GERGPPGSPGLQGFPGI-TPPSNISGAPGDKGA-
---PGIFGLKGYRGPPGPPGSAALPGSKGDAGNPGAPGTPGTKGWAGDS
PF01391 // ENSGT00900000140804_1_Domain_20 (2030, 2108)
GPQGRPGVFGL
PGEKG---------PRGEQGFMGNTGPA-----GAVGDRGPKGPKGDPGFPG-------A
PGIVGAPGIAGIPQKIAIPPGTVGPQGRRGPPGAPGEMGPQGPPGEP-------------
---------------------------------------GFRGAPGKAGPQGRGGVSAVP
GFRGDEGPVGHQGPIGQE----GAPGRPGSPGLPGMPGRSVSIGY
PF01413 // ENSGT00900000140804_1_Domain_23 (2266, 2371)
LLVKHSQTDQEPMCP
VGMNKLWSGYSLLYFEGQEKAHNQDLGLAGSCLARFSTMPFLYCNPGDVCYYASRNDKSY
WLSTTAPLPMMPVAEDE--IKPYISRCSVCENSGT00900000140804_1_Linker_23_24 (2371, 2376)
EAPAVPF01413 // ENSGT00900000140804_1_Domain_24 (2376, 2409)
AIAVHSQ--DVSIPHCPAGWRSLWI
GYSFLMVRGIYPVPL--------PSHTLAGDTARTP---AFVRVIGQGSSGCALHE---G
--TWPRERER------------------AGI---QLTLC---------------------
--------------
ENSPCAP00000014473 (view gene)
--------------MGRQWRAAAPYCALWRWLLLGTVTVGFFAQSALAGVKKFGVACGGR
DCS-GGCQCYPEKGARXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXX
XXXXXXXXXXX
PF01391 // ENSGT00900000140804_1_Domain_0 (132, 193)
XXXXX---GHPGQGGPRGRPGYDGCNGTAGDAGAQGAPGSAGIPGYPGP
QGPKGQKGEPYAENSGT00900000140804_1_Linker_0_1 (193, 202)
LSQEDRDKYPF01391 // ENSGT00900000140804_1_Domain_1 (202, 251)
RGEPGEPGLVGLQGPSGHPGYVGPMGPIGAPGRPGPPAP
PGPKGQPGNRENSGT00900000140804_1_Linker_1_2 (251, 334)
GLGF--YGEKGEKGDIGRPGPNGIPSDN-DRAIVGPQKEVIHPEQYKXXX
XXXXXXXXXXX-------XXXXXXXXXXXXXXXPF01391 // ENSGT00900000140804_1_Domain_2 (334, 449)
GFPGV---DGEPGTPGQ----------
---------------------------------------------------------KGS
KGLDGYQGPV-GPKGVKGEVGDQGPPGLPAYTPHPSLE----------------------
------------------------------------KXXXXXXXXXXXXXXXXXXXXXX-
------------XXXXXX--------------------------XXXXXXXXXXERGLP-
EMGPKGFAGD-GIPAVFP
PF01391 // ENSGT00900000140804_1_Domain_5 (619, 663)
GPPGADGTPGLPGPPGPPGLPGPD------------------
-GFLFGLKGVAGAVGYPGPSGFPGPSGQKGHKX---------------------------
-----------------------XXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXX
XXXXXXXXXXX--XX-XX-XXXXXXX----------------------------------
------------------------------------------------------------
------------------------------------------------------------
-----------------------------------------XXX---XX--XXXXXXXXX
XXXXXXXX---XXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXGDGIKGLP
PF01391 // ENSGT00900000140804_1_Domain_9 (1073, 1131)
GDTGYPGV
PGTKGMPGEVGPPGLGLPGPKGERGFPGDVGIPGSPGFPGPPGPPGTPGQIDCD---NGV
KRPIAGAGQDAV--LPGCV--GGPK-----------------------GLPGLPGPQ---
G----------------PPGAKGLRGLPGFSGPDGEPGLKGFPGDPGREGFPGPQXXXXX
XXXXXXXXXXXXXXXXXXXXXXX-XXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXX
PGQRGPPGFRGS-----QGMPGMPGPKGQPGFPGPSGQTGLPGPPGQHGF-PGAPGQPGP
LGLPGAP-GR--------------------GGLPGDRGDPGDTGFPGPVGMKGLPGDGGD
AGIVGDRGQPGNPGFKGMAGMPGIPGRK-------
PF01391 // ENSGT00900000140804_1_Domain_13 (1476, 1539)
GDKGSPGMQGFQGMVGLKGRAGYPG
NKGEPGFFGF----PGPKGLAGEPGVKGNRGDPGLPGPENSGT00900000140804_1_Linker_13_14 (1539, 1555)
PPIILPG-MKDIKGEKPF01391 // ENSGT00900000140804_1_Domain_14 (1555, 1605)
GDQGPM
GVKGSLGLK-------------GLQGMPGVPGLSGIP-GLPGRPGHIKGT-KGDTGVPGI
PGSPGFPGVTGPPGISGFPGFIGSRGDKGAPGR---------------------------
---AGPSGQTGETGDY---GDIGDTIDLPGNPGVKGERGITGTP-GQKG---SEKGAGGS
GFPGIVGMKGAQGP-RLKGQTGFPGLTGLQGPQGEPGRIGLHG--EKGDYGWQGPPGLP-
--------------------------------GLPGIRGISGLHGLPGTKGLPGSPGADI
PG----------D---PGFPGIAGDRGDPGEPNTRPGPV
PF01391 // ENSGT00900000140804_1_Domain_18 (1900, 1999)
GVPGQKGQQGTP---------
--------------------------GGRGPVGSPGLPGFPGI-TPPSNISGSPGDQGA-
---QGIFGPQGYQGPPGPPGAAALPGSKGDAGNPGFSGNPGPKGWVGDPGPEGIPGVFGL
PGEKG---------PRGEPGFMGITGVP-----GSVGDRGPKGPKGDRGLPG-------P
PGSLGSPGIAGIPQK-ADEPGILGPYGRRGLFGAPGEMGPQGPPD-P-------------
--------------------------------------
PF01391 // ENSGT00900000140804_1_Domain_22 (2199, 2254)
-GLRGAPGKAGPQGRGGVSAVP
GFRGDQGPIGQQGPAGQE----GEPGRQGSPGLENSGT00900000140804_1_Linker_22_23 (2254, 2264)
PGMPGRSVSIPF01413 // ENSGT00900000140804_1_Domain_23 (2264, 2306)
GYLLVKHSQTAQEPMCP
VGMNKLWSGYSLLYFEGQEKAHNQDLXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXX
XXXXXXXXXXXXXXXXX----XXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXX
XXXX
PF01413 // ENSGT00900000140804_1_Domain_25 (2405, 2505)
XXHTAAGDEGGQS------LFPGNCLEDFRAT---PFIECNG-GGGTCHYYANKYS
--FWLTTIPEQDLQGSPSA-----DTVKAGLIRTHISRCQVCMKNLRPGG----------
--------------
ENSODEP00000016445 (view gene)
--------------------AFAMGPAPTGWLLLGAVTVGLVTHSVLAGVKKLEVPCGGR
DCS-GGCQCYPEKGGRGQPGPVGPQGFSGPPGLQGFPGLQGRKGDKGERGSPGTPGPKGD
VGPRGVSGFP
PF01391 // ENSGT00900000140804_1_Domain_0 (131, 190)
GADGIP---GHPGQGGPRGRPGYDGCNGTRGDTGPQGPSGSEGFPGTPGP
QGPKGQKGEENSGT00900000140804_1_Linker_0_1 (190, 203)
PYALPPEERDRYRPF01391 // ENSGT00900000140804_1_Domain_1 (203, 251)
GETGEPGLVGFQGPPGRPGPVGPMGPVGAPGRPGPPGP
PGPKGQPGNRENSGT00900000140804_1_Linker_1_2 (251, 322)
GLGF--YGEKGEKGEQGLPGPNGIPSDT-THPIVAPTSSTI--LPYKGEK
GS---QGEPGMTGI----SYKPF01391 // ENSGT00900000140804_1_Domain_2 (322, 448)
GEEGIVGFPGTRGPPGL---DGEKGLPGQ----------
---------------------------------------------------------KGS
RGHDGYQGPD-GAQGQKGEIGEPGLPGPPAYSPHP-------------------------
-----------------------------------------ALAKGVRGDPGFQGTYGEP
GSRGEPGEPGLPGP---P--------------------------GLPAGDRG-DRRGLTG
EMGPKGFTGDPGIIAPVP
PF01391 // ENSGT00900000140804_1_Domain_5 (619, 747)
GPKGADGRPGLPGRPGPPGPPGPD------------------
-GFLFGLKGAPGSRGYPGHSGFPGLRGQKGWKG---------------------------
-----------------------DAGENSGT00900000140804_1_Linker_5_6 (747, 762)
DC-RCAGGDQ---FIPF01391 // ENSGT00900000140804_1_Domain_6 (762, 1014)
PGLPGLKGFPGANGEPGRK
GEPGDPGQHGIPGFPGFKGAPGNVGA----------------------------------
------------------------------------------------------------
------------------------------------------------------------
-----------------------------------------PGP---KG--AKGDVRAVT
TKGEKGQPGIPGVHGMKGDDGTPGRDGLDGFPGL-----PGP-PGDGIKGPPGDAGYPGA
PGTKGIPGETGLPGLGLPGPKGEQGFPGDAGFPGPPGLPGPPGPPGSPGQTDCD---TGV
KRPIGDDRQESV--QPGCI--GGPK-----------------------GSPGRPGPP---
G----------------VPGDKGPRGIPGFAGEDGPQGLKGFPGDPGRDGLPGPP-GIIG
PRGSKGAVGRPGPDGLQGPDGLPGTVGAPGDRGIPGEVLGAEPGSRGDAGLPGQPGLKGV
PGERGPPGFRGS-----QGMPGMPGLKGQLGLPGPSGLQGRPGPPGRHGF-PGAPGPEGP
LGLPGVP-GR--------------------EGLRGDRGEPGDTGAPGPVGMKGLAGDRGD
AGAGGDRGHPGTPGFKGIVGMPGSPGHK-------
PF01391 // ENSGT00900000140804_1_Domain_13 (1476, 1538)
GERGSPGMDGFQGMIGLKGRPGVPG
TKGEAGLSGQ----PGVRGPPGQRGGKGTRGDPGPPGPPPAILPV-IKYLKGEKGDEGPM
GLKGHLGLK-------------GIQGMPGVPGQAGVP-GLPGRPGLIKGV-KGDIGVPGS
PGLPGFPGVVGPPGIVGFPGFTGSRGDKGAPGR---------------------------
---AGLYGEAGQPGDF---GDTGDTINLPGSPGLKGERGLTGTP-GLKGFFGERGTQGEV
GFPGISGLAGPQGPPGIKGQTGFP
PF01391 // ENSGT00900000140804_1_Domain_17 (1765, 1858)
GLTGLQGPQGDPGQIGFPG--DKGDDGWPGVPGLP-
--------------------------------GFPGLRGIGGLHGLPGPKGSPGSPGADM
PG----------D---PGFPGSIGDRGDRGEANTSPGPTGVPGQKGEQGAP---------
--------------------------GERGPAGRPGIQGFPGI-SPASNISGLPGDLGS-
---PGIFGLKGYPGPQGPPGPPALPGSKGDSGSAGAPGFPGTKGWGGDP
PF01391 // ENSGT00900000140804_1_Domain_20 (2030, 2108)
GPQGQPGVFGV
PGEKG---------PRGDPGLMGNTGPT-----GTVGDRGPKGPKGDLGFPG-------A
PGSVGSPGIAGIPQRILAPAGTIGPEGRRGPPGARGEMGPQGPPGEP-------------
---------------------------------------GFRGVPGKPGPQGRGGVSALP
GFRGDPGPVGQQGPLGQE----GEPGRPGSPGLPGMPGRSVSIGY
PF01413 // ENSGT00900000140804_1_Domain_23 (2266, 2371)
LLVKHSQTDQEPMCP
VGMNRLWSGYSLLYFEGQEKAHNQDLGLAGSCLARFSTMPFLYCNPGDVCYYASRNDKSY
WLSTTAPLPMMPVAEDD--IKPYISRCSVCENSGT00900000140804_1_Linker_23_25 (2371, 2376)
EAPAVPF01413 // ENSGT00900000140804_1_Domain_25 (2376, 2504)
AIAVHSQ--DVSIPHCPAGWRSLWI
GYSFLMHTAAGDEGGGQS----LVSPGSCLEDFRAT---PFIECNG-ARGTCHYFANKYS
--FWLTTIPEASFQGTPSA-----DTLKAGLIRTHISRCQVCMKNL--------------
--------------
ENSANAP00000018465 (view gene)
--------------MGRNQ-RAAAGPALPRWLLLGTVTVGFLSQSVLAGVKKSDVPCGGR
DCS-GGCQCFPEKGGRGQPGPVGPQGYTGPPGLQGFPGLQGRKGDKGERGAPGITGPKGD
VGARGVSGFP
PF01391 // ENSGT00900000140804_1_Domain_0 (131, 189)
GADGIP---GHPGQGGPRGRPGYDGCNGTQGDAGPQGPPGSEGFTGPPGP
QGPKGQKGEPYALPEEERARYRGEPGEPGLVGFQGPPGRPGPVGQMGPVGAPGRPVS---
---TWTQGNRGLGF--YGVKGEKGDVGQPGPNGIPSET--HPVIAPTGVTYHPEQDKVKN
SQ---PLVRIRTGI----SLKGEEGIMGFPGPRGYPGL---SGEKGSPGQ----------
---------------------------------------------------------KVS
WARELQCALG-PRSPTEGETGDPGPPGLPAYSPHP-------------------------
-----------------------------------------SLAKGARGDPGFPGARGEP
GSQGEPGDPGPLGP---P--------------------------GISIGAED-LGRGLPG
EMGPKGFIGDPGIPALFP
PF01391 // ENSGT00900000140804_1_Domain_5 (619, 747)
GPPGLDGKPGLPGPPGLPGPPGPD------------------
-GFLFGLKGAEGRAGFPGLSGSPGARGQKGWKG---------------------------
-----------------------DAGENSGT00900000140804_1_Linker_5_6 (747, 760)
DC-RCAEGDEPVRPF01391 // ENSGT00900000140804_1_Domain_6 (760, 1015)
GLPGLPGPKGFPGINGEPGRK
GDKGDPGQHGLPGFPGLKGVPGNVGA----------------------------------
------------------------------------------------------------
------------------------------------------------------------
-----------------------------------------PGP---KG--TKGDSRTIT
TKGERGQPGVPGVPGMKGDDGIPGREGLDGFPGL-----PGP-PGDGIK
PF01391 // ENSGT00900000140804_1_Domain_9 (1070, 1129)
GPPGDPGYPGI
PGTKGSPGEMGPPGLGLPGLKGQRGFPGDAGLPGPPGFPGPSGPPGTPGRTDCD---PDV
RRPIAGDRQEAV--QPCPFILVGPD-----------------------V---LCGLF---
G----------------CLGAKGLRGTPGFSGADGGPGPKGLSGDPGREGFPGPP-GFIG
PRGSKGAAGLPGPDGLPGPVGLPGPDGPPGDKGLPGEVLGAQPGPRGDAGVPGHPGLKGL
PGDRGFPGFRGL-----SG------------------------PPGQRGF-PGAPGPEGP
FGLPGTL-GLEGKAPALPQTGGVLTGAARRTCLPGDRGEPGDVGAPGPVGMKGVSGDRGD
TGLAGERGHPGSPGFKGIDGMPGAPGLK-------
PF01391 // ENSGT00900000140804_1_Domain_13 (1476, 1537)
GERGSPGMDGFQGMPGLKGRPGFPG
SKGEAGFFGI----PGLKGLAGEPGFKGSRGDPGPPGPPPVILPG-MKDIKGDKGDEGPM
GLKGYLGAK-------------GIQGMPGIPGLSGIP-GLPGRPGHIK
PF01391 // ENSGT00900000140804_1_Domain_15 (1609, 1694)
GV-KGDIGAPGI
PGLPGFPGVAGPPGITGFPGFTGSRGDKGSPGR---------------------------
---AGLYGEIGRTGDF---GDIGDTINLPGRPGLKGERGTTGIP-GLKGFFGEKGTEGDI
GFPGITGVTGVQGPPGLKGQAGFPGLTGPPGPQGEPGRIGLPG--GKGDDGWPGAPGLP-
--------------------------------GFPGPRGISGLHGLPGTKGFPGSPGADI
HG----------D---PGFSGPPGERGDPGEANTLPGPVGVPGQKGEQGAP---------
--------------------------GQRGPPGSPGLQGFPGI-TPPSNISGVPGDKGA-
---PGIFGLKGYRGPPGPPGSAALPGSKGDTGNPGAPGTPGTKGWVGDP
PF01391 // ENSGT00900000140804_1_Domain_20 (2030, 2108)
GPQGRPGVLGL
PGEKG---------PRGEQGFMGNPGLP-----GSVGDRGPKGPKGDPGFPG-------A
PGIVGAPGIAGIPQKIATQPGTVGPQGRRGPPGAPGEMGPQGPPGEP-------------
---------------------------------------GFRGAPGKAGPQGRGGVSAVP
GFRGDEGPVGHQGPTGQE----GVPGRPGSPGLPGMPGRSVSIGY
PF01413 // ENSGT00900000140804_1_Domain_23 (2266, 2371)
LLVKHSQTDQEPMCP
VGMNKLWSGYSLLYFEGQEKAHNQDLGLAGSCLARFSTMPFLYCNPGDVCYYASRNDKSY
WLSTTAPLPMMPVAEDE--IKPYISRCSVCENSGT00900000140804_1_Linker_23_25 (2371, 2376)
EAPAVPF01413 // ENSGT00900000140804_1_Domain_25 (2376, 2504)
AIAVHSQ--DVSIPHCPAGWRSLWI
GYSFLMDTKGEHEGGGQQSNRLFTGQSTCLEDFAPYFNTSSIHSSAVTRHPATNYANKYK
ELLVLTTIPEQSFQELCSLSLAPITPLSARLLIRGNCRCQVCMKTL--------------
--------------
ENSCAFP00000009112 (view gene)
---------------------------------------------LLQGVKKFDVPCGGR
DCS-GGCQCYPEKGGRGQPGPV
PF01391 // ENSGT00900000140804_1_Domain_0 (83, 143)
GPQGYTGPPGLQGFPGLQGRKGDKGERGSPGITGPKGD
VGARGVSGFPGADGIP---GHPGQGGPRGPPGYDGCNGTRGDVGRQGPEGPAGFLGPPGP
QGPKGQKGEPYALSSVDRDKYR
PF01391 // ENSGT00900000140804_1_Domain_1 (203, 251)
GEPGEPGLVGFQGPPGRPGPIGQMGPVGAPGRPGPPGP
PGPKGQPGNRENSGT00900000140804_1_Linker_1_2 (251, 298)
GLGF--YGEKGEKGDMGLPGPNGIPSDT-HHAIIGPMKEMFHPDQYKPF01391 // ENSGT00900000140804_1_Domain_2 (298, 436)
GEK
GS---EGEPGRKGI----SLKGEEGIMGFSGTRGAPGY---DGEKGSPGQ----------
---------------------------------------------------------KGS
RGLDGYQGPD-GPPGENSGT00900000140804_1_Linker_2_3 (436, 526)
PKKCALQVGTGSSPVLTMGP-------------------------
-----------------------------------------ISVAPF01391 // ENSGT00900000140804_1_Domain_3 (526, 613)
GARGDPGFPGTHGDP
GSRGEPGDPGPPGL---P--------------------------GPSIRDED-GKRGLPG
EMGPKGFIGDPGIPAPYPGPPGTDGKPGLRGPPGAAGPPGPD-------------------GFLFGLKGTEGSAGFPGPSGFPGARGQKGWKG--------------------------------------------------DAGDC-KCAEGDQFVGGLPGLPGPKGFPGINGEPGRKGSQGDPGQHGIPGFPGFKGAPGTVGP---------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------PGP---KG--LKGDSRAITTKGERGQPGVPGVPGLKGDDGVPGRDGLDGFPGL-----PGP-PGDGIKGPPGDSGYPGAPGTKGTPGERGPPGLGLPGPKGERGFPGDAGLPGPPGFPGPPGPPGTPGQIDCD---TGVKRPIGGDGQEAV--QPDCV--RGPM-----------------------GSPGLPGPP---G----------------PTGAKGLQGIPGLSGADGTPGLRGLPGDPGREGFPGPP-GFVGPRGSKGAVGLPGPDGHPGPTGLPGPVGPPGDRGLPGEVLGAQPGPRGDAGLPGHPGLKGPPGERGPPGFRGS-----QGMLRMPGLKGQPGFPGLSGQPGLPGPSGQHGF-QS-SGREGPLGLPGSLRSR--------------------RGLPGDRGDPGDTGVPGPVGMKGLSGDRGDPGLLGERGPPGSPGFKGMSGMPGIPGQK-------GGRGSPGMDGFQGMVGLKGRPGLPGNKGEAGFFGI----PGLKGLAGEPGVKGSRGDPGPPGPPPIIQPG-MKDIKGEKGDEGPMGLKGYLGLK-------------GLPGMPGIPGLSGVP-GLPGKPGHIKGI-KGDIGIPGVPGLPGFPGVPGPPGIIGFPGFTGSRGDKGAPGR------------------------------AGLYGEVGPTGDF---GDSGDIINLPGSPGLKGERGIAGAP-GLKGFFGEKGTEGEVGFPGITGVPGVQGPPGPKGQTGFPGLTGLQGPQGDPGRAGVPG--NKGDYGWPGIPGSP---------------------------------GFPGIRGISGLHGLPGTKGLPGTPGADIHG----------D---PGFPGPAGDRGDPGEANIRPGPIGAPGQKGERGDP-----------------------------------GGRGPIGSPGLQGFPGI-TPPSNMSGLPGDKGA----PGIFGLEGYRGPPGPPGPAALPGSKGDEGNPGAPGTPGTKGWGGDPGPQGRPGVFGLPGEKG---------PRGEPGFMGNTGAT-----GSVGDRGPKGPKGDRGLPG-------APGSVGSPGIAGIPQKITIQPGPVGPQGRRGPPGMQGEMGPQGPPGDP----------------------------------------------------GFRGAPGKAGPQGRGGVSAVPGFRGDQGPMGHQGPVGQE----GEPGRPGTPGLPGMPGRSVSIGYLLVKHSQTDQEPMCPVGMNKLWSGYSLLYFEGQEKAHNQDLGLAGSCLARFSTMPFLYCNPGDVCYYASRNDKSYWLSTTAPLPMMPVAEDD--IKPYISRCSVCEAPAVAIAVHSQ--DVSIPHCPAGWRSLWIGYSFLMHTAAGDEGGGQS----LVSPGSCLEDFRAT---PFIECNG-GRGTCHYYANKYS--FWLTTIPEQSFQGSPSA-----DTLKAGLIRTHISRCQVCMKNL--------
PF01391 // ENSGT00900000140804_1_Domain_5 (593, 662)
-GKRGLPGEMGPKGFIGDPGIPAPYPGPPGTDGKPGLRGPPGAAGPPGPD------------------
-ENSGT00900000140804_1_Linker_5_6 (662, 760)
GFLFGLKGTEGSAGFPGPSGFPGARGQKGWKG---------------------------
-----------------------DAGDC-KCAEGDQFVGPF01391 // ENSGT00900000140804_1_Domain_6 (760, 1015)
GLPGLPGPKGFPGINGEPGRK
GSQGDPGQHGIPGFPGFKGAPGTVGP----------------------------------
------------------------------------------------------------
------------------------------------------------------------
-----------------------------------------PGP---KG--LKGENSGT00900000140804_1_Linker_6_7 (1015, 1026)
DSRAIT
TKGERPF01391 // ENSGT00900000140804_1_Domain_7 (1026, 1092)
GQPGVPGVPGLKGDDGVPGRDGLDGFPGL-----PGP-PGDGIKGPPGDSGYPGA
PGTKGTPGERGPPGLGLPGPKGERGFPGDAGLPGPPGFPGPPGPPGTPGQIDCD---TGVKRPIGGDGQEAV--QPDCV--RGPM-----------------------GSPGLPGPP---G----------------PTGAKGLQGIPGLSGADGTPGLRGLPGDPGREGFPGPP-GFVGPRGSKGAVGLPGPDGHPGPTGLPGPVGPPGDRGLPGEVLGAQPGPRGDAGLPGHPGLKGPPGERGPPGFRGS-----QGMLRMPGLKGQPGFPGLSGQPGLPGPSGQHGF-QS-SGREGPLGLPGSLRSR--------------------RGLPGDRGDPGDTGVPGPVGMKGLSGDRGDPGLLGERGPPGSPGFKGMSGMPGIPGQK-------GGRGSPGMDGFQGMVGLKGRPGLPGNKGEAGFFGI----PGLKGLAGEPGVKGSRGDPGPPGPPPIIQPG-MKDIKGEKGDEGPMGLKGYLGLK-------------GLPGMPGIPGLSGVP-GLPGKPGHIKGI-KGDIGIPGVPGLPGFPGVPGPPGIIGFPGFTGSRGDKGAPGR------------------------------AGLYGEVGPTGDF---GDSGDIINLPGSPGLKGERGIAGAP-GLKGFFGEKGTEGEVGFPGITGVPGVQGPPGPKGQTGFPGLTGLQGPQGDPGRAGVPG--NKGDYGWPGIPGSP---------------------------------GFPGIRGISGLHGLPGTKGLPGTPGADIHG----------D---PGFPGPAGDRGDPGEANIRPGPIGAPGQKGERGDP-----------------------------------GGRGPIGSPGLQGFPGI-TPPSNMSGLPGDKGA----PGIFGLEGYRGPPGPPGPAALPGSKGDEGNPGAPGTPGTKGWGGDPGPQGRPGVFGLPGEKG---------PRGEPGFMGNTGAT-----GSVGDRGPKGPKGDRGLPG-------APGSVGSPGIAGIPQKITIQPGPVGPQGRRGPPGMQGEMGPQGPPGDP----------------------------------------------------GFRGAPGKAGPQGRGGVSAVPGFRGDQGPMGHQGPVGQE----GEPGRPGTPGLPGMPGRSVSIGYLLVKHSQTDQEPMCPVGMNKLWSGYSLLYFEGQEKAHNQDLGLAGSCLARFSTMPFLYCNPGDVCYYASRNDKSYWLSTTAPLPMMPVAEDD--IKPYISRCSVCEAPAVAIAVHSQ--DVSIPHCPAGWRSLWIGYSFLMHTAAGDEGGGQS----LVSPGSCLEDFRAT---PFIECNG-GRGTCHYYANKYS--FWLTTIPEQSFQGSPSA-----DTLKAGLIRTHISRCQVCMKNL------
PF01391 // ENSGT00900000140804_1_Domain_9 (1070, 1127)
GPPGDSGYPGAPGTKGTPGERGPPGLGLPGPKGERGFPGDAGLPGPPGFPGPPGPPGTPGQIDCD---TGV
KRPIGGDGQEAV--QPDCV--RGPM-----------------------GSPGLPGPP---
G----------------PTGAKGLQGIPGLSGADGTPGLRGLPGDPGREGFPGPP-GFVG
PRGSKGAVGLPGPDGHPGPTGLPGPVGPPGDRGLPGEVLGAQPGPRGDAGLPGHPGLKGP
PGERGPPGFRGS-----QGMLRMPGLKGQPGFPGLSGQPGLPGPSGQHGF-QS-SGREGP
LGLPGSLRSR--------------------RGLPGDRGDPGDTGVPGPVGMKGLSGDRGD
PGLLGERGPPGSPGFKGMSGMPGIPG
PF01391 // ENSGT00900000140804_1_Domain_13 (1467, 1535)
QK-------GGRGSPGMDGFQGMVGLKGRPGLPG
NKGEAGFFGI----PGLKGLAGEPGVKGSRGDPGPPGPPPIIQPG-MKDIKGEKGDEGPM
GLKGYLGLK-------------GLPGMPGIPGLSGVP-GLPGKPGHIKGI-K
PF01391 // ENSGT00900000140804_1_Domain_15 (1613, 1703)
GDIGIPGV
PGLPGFPGVPGPPGIIGFPGFTGSRGDKGAPGR---------------------------
---AGLYGEVGPTGDF---GDSGDIINLPGSPGLKGERGIAGAP-GLKGFFGEKGTEGEV
GFPGITGVPGVQGPPGPKGQTGFPGLTGLQGPQGDPGRAGVPG--NKGDYGWPGIPGSP-
--------------------------------GFPGIRGISGLHGLPGTKGLPGTPGADI
HG----------D---PGFPGPAGDRGDPGEANIRPGPIGAPGQKGERGDP---------
--------------------------GGRGPIGSPGLQGFPGI-TPPSNMSGLPGDKGA-
---PGIFGLEGYRGPPGPPGPAALPGSKGDEGNPGAPGTPGTKGWGGDPGPQ
PF01391 // ENSGT00900000140804_1_Domain_20 (2033, 2111)
GRPGVFGL
PGEKG---------PRGEPGFMGNTGAT-----GSVGDRGPKGPKGDRGLPG-------A
PGSVGSPGIAENSGT00900000140804_1_Linker_20_21 (2111, 2120)
GIPQKITIQPF01391 // ENSGT00900000140804_1_Domain_21 (2120, 2221)
PGPVGPQGRRGPPGMQGEMGPQGPPGDP-------------
---------------------------------------GFRGAPGKAGPQGRGGVSAVPENSGT00900000140804_1_Linker_21_23 (2221, 2265)
GFRGDQGPMGHQGPVGQE----GEPGRPGTPGLPGMPGRSVSIGPF01413 // ENSGT00900000140804_1_Domain_23 (2265, 2372)
YLLVKHSQTDQEPMCP
VGMNKLWSGYSLLYFEGQEKAHNQDLGLAGSCLARFSTMPFLYCNPGDVCYYASRNDKSY
WLSTTAPLPMMPVAEDD--IKPYISRCSVCEENSGT00900000140804_1_Linker_23_25 (2372, 2375)
APAPF01413 // ENSGT00900000140804_1_Domain_25 (2375, 2505)
VAIAVHSQ--DVSIPHCPAGWRSLWI
GYSFLMHTAAGDEGGGQS----LVSPGSCLEDFRAT---PFIECNG-GRGTCHYYANKYS
--FWLTTIPEQSFQGSPSA-----DTLKAGLIRTHISRCQVCMKNL--------------
--------------
ENSMLEP00000037549 (view gene)
--------------MGRDQ-RAVAGPALRRWLLLGTVTVGFLAQGVLAGVKKFDVPCGGR
DCS-GGCQCYPEKGGRGQPGPVGPQGYNGPPGLQGFPGLQGRKGDKGERGAPGITGPKGD
VGARGVSGFP
PF01391 // ENSGT00900000140804_1_Domain_0 (131, 189)
GADGIP---GHPGQGGPRGRPGYDGCNGTQGDSGPQGPPGSEGFTGPPGP
QGPKGQKGENSGT00900000140804_1_Linker_0_1 (189, 203)
EPYALPREERDRYRPF01391 // ENSGT00900000140804_1_Domain_1 (203, 251)
GEPGEPGLVGFQGPPGRPGPVGQMGPVGAPGRPGPPGP
PGPKGQQGNRENSGT00900000140804_1_Linker_1_2 (251, 321)
GLGF--YGVKGEKGDVGQPGPNGIPSDT-LHPIIAPTGVTFHPDQYKGEK
GS---EGEPGIRGI----SLPF01391 // ENSGT00900000140804_1_Domain_2 (321, 447)
KGEEGIMGFPGPRGSPGL---SGEKGSPGQ----------
---------------------------------------------------------KGS
RGLDGYQGPD-GPRGPKGEAGDIGPPGLPAYSPHP-------------------------
-----------------------------------------SLAKGARGDPGFPGAQGAP
GSQGEPGDPGPQGP---P--------------------------GLSIGDED-QRRGLPG
EMGPKGFIGDPGIPALFP
PF01391 // ENSGT00900000140804_1_Domain_5 (619, 747)
GPPGPDGKPGPPGPPGLPGPPGPD------------------
-GFLFGLKGPEGRPGFPGLSGSPGARGPKGWKG---------------------------
-----------------------DAGENSGT00900000140804_1_Linker_5_6 (747, 760)
DC-RCAEGDEAVTPF01391 // ENSGT00900000140804_1_Domain_6 (760, 1015)
GLPGLPGPKGFAGINGEPGRK
GDKGDPGQHGLPGFPGLKGVPGNVGA----------------------------------
------------------------------------------------------------
------------------------------------------------------------
-----------------------------------------PGP---KG--AKGENSGT00900000140804_1_Linker_6_7 (1015, 1023)
DSRTIT
TKPF01391 // ENSGT00900000140804_1_Domain_7 (1023, 1082)
GERGQPGVPGVPGIKGDDGSPGRDGLDGFPGL-----PGP-PGDGIKGPPGDAGYPGI
PGTKGTPGEMGPPGLGLPGFKGQRGFPGDAGLPGPPGFPGPPGPSGTPGQIDCD---TDVKRPIGGDRQEAV--QPGCI--GGPK-----------------------GLPGLPGPP---G----------------PSGAKGLRGIPGFSGADGGPGPKGLPGDPGHEGFPGPP-GFIGPRGSKGAVGLPGPDGPPGPIGLPGPDGPPGDRGLPGEVLGAQPGPRGDAGVPGQPGFKGLPGDRGTPGFRGS-----QGMPGMPGLKGQPGLPGPSGQPGLSGPPGQHGF-PGAPGQEGPLGLPGIP-GL--------------------EGLPGDRGDPGDIGAPGPVGMKGVSGDRGDAGLAGERGHPGSPGFKGIDGMPGTPGLK-------GERGSPGMDGFQGMPGLKGRPGFPGSKGEAGFFGI----PGLKGLAGEPGFKGSRGDPGPPGPPPIILPG-MKDIKGEKGDEGPMGLKGYLGAK-------------GIQGMPGIPGLSGIP-GLPGRPGHIKGV-KGDIGAPGIPGLPGFPGVAGPPGITGFPGFIGSRGDKGAPGR------------------------------AGLYGEIGPTGDF---GDIGDTINLPGRPGLKGEQGTAGIP-GLKGFFGEKGTEGDIGFPGITGVTGVQGPPGLKGQTGFPGLTGPPGSQGEPGRIGLPG--GKGDDGWPGTSGLP---------------------------------GFPGLRGIRGLHGLPGTKGFPGSPGADIHG----------D---PGFPGPPGDRGDPGDANTLPGPVGVPGQKGQQGAP-------------------------------------------------------------------------------GYRGPPGPPGSAALPGSKGDAGNPGAPGTPGTKGWAGDSGPQGRPGVFGLPGEKG---------PRGEQGFMGNTGPA-----GAVGDRGPKGPKGDPGFPG-------APGIVGAPGIAGIPQKIAIPPGTVGPQGRRGPPGAPGEMGPQGPPGEP----------------------------------------------------GFRGAPGKAGPQGRGGVSAVPGFRGDEGPVGHQGPIGQE----GAPGRPGSPGLPGMPGRSVSIGYLLVKHSQTDQEPMCPVGMNKLWSGYSLLYFEGQEKAHNQDLGLAGSCLARFSTMPFLYCNPGDVCYYASRNDKSYWLSTTAPLPMMPVAEDE--IKPYISRCSVCEAPAVAIAVHSQ--DVSIPHCPAGWRSLWIGYSFLMHTAAGDEGGGQS----LVSPGSCLEDFRAT---PFIECNG-GRGTCHYYANKYS--FWLTTIPEQSFQGSPSA-----DTLKAGLIRTHISRCQVCMKNL----------------
PF01391 // ENSGT00900000140804_1_Domain_9 (1070, 1129)
GPPGDAGYPGIPGTKGTPGEMGPPGLGLPGFKGQRGFPGDAGLPGPPGFPGPPGPSGTPGQIDCD---TDV
KRPIGGDRQEAV--QPGCI--GGPK-----------------------GLPGLPGPP---
G----------------PSGAKGLRGIPGFSGADGGPGPKGLPGDPGHEGFPGPP-GFIG
PRGSKGAVGLPGPDGPPGPIGLPGPDGPPGDRGLPGEVLGAQPGPRGDAGVPGQPGFKGL
PGDRGTPGFRGS-----QGMPGMPGLK
PF01391 // ENSGT00900000140804_1_Domain_12 (1348, 1428)
GQPGLPGPSGQPGLSGPPGQHGF-PGAPGQEGP
LGLPGIP-GL--------------------EGLPGDRGDPGDIGAPGPVGMKGVSGDRGD
AGLAGERGHPGSPGFKGIDGMPGTPGLK-------GERGSPGMDGFQGMPGLKGRPGFPG
SKGEAGFFGI----PGLKGLAGEPGFKGSRGDPGPPGPPPIILPG-MKDIKGEKGDEGPM
GLKGYLGAK-------------GIQGMPGIPGLSGIP-GLPGRPGHI
PF01391 // ENSGT00900000140804_1_Domain_15 (1608, 1694)
KGV-KGDIGAPGI
PGLPGFPGVAGPPGITGFPGFIGSRGDKGAPGR---------------------------
---AGLYGEIGPTGDF---GDIGDTINLPGRPGLKGEQGTAGIP-GLKGFFGEKGTEGDI
GFPGITGVTGVQGPPGLKGQTGFPGLTGPPGSQGEPGRIGLPG--GKGDDGWPGTSGLP-
--------------------------------GFPGLRGIRGLHGLPGTKGFPGSPGADI
HG----------D---PGFPGPPGDRGDPGDANTLPGPVGVPGQKGQQGAP---------
------------------------------------------------------------
----------GYRGPPGPPGSAALPGSKGDAGNPGAPGTPGTKGWAGDS
PF01391 // ENSGT00900000140804_1_Domain_20 (2030, 2108)
GPQGRPGVFGL
PGEKG---------PRGEQGFMGNTGPA-----GAVGDRGPKGPKGDPGFPG-------A
PGIVGAPGIAGIPQKIAIPPGTVGPQGRRGPPGAPGEMGPQGPPGEP-------------
---------------------------------------GFRGAPGKAGPQGRGGVSAVP
GFRGDEGPVGHQGPIGQE----GAPGRPGSPGLPGMPGRSVSIGY
PF01413 // ENSGT00900000140804_1_Domain_23 (2266, 2371)
LLVKHSQTDQEPMCP
VGMNKLWSGYSLLYFEGQEKAHNQDLGLAGSCLARFSTMPFLYCNPGDVCYYASRNDKSY
WLSTTAPLPMMPVAEDE--IKPYISRCSVCENSGT00900000140804_1_Linker_23_25 (2371, 2376)
EAPAVPF01413 // ENSGT00900000140804_1_Domain_25 (2376, 2504)
AIAVHSQ--DVSIPHCPAGWRSLWI
GYSFLMHTAAGDEGGGQS----LVSPGSCLEDFRAT---PFIECNG-GRGTCHYYANKYS
--FWLTTIPEQSFQGSPSA-----DTLKAGLIRTHISRCQVCMKNL--------------
--------------
ENSPCOP00000017693 (view gene)
--------------MGRDR-RAASGPALQRWLLLGAVTVGVLAQSVLAGVKKLDVPCGGR
DCS-GGCQCYPEKGGRGQPGPVGPQGYNGPPGLQGFPGLQGRKGDKGERGAPGITGPKGD
VGARGVSGFP
PF01391 // ENSGT00900000140804_1_Domain_0 (131, 190)
GADGIP---GHPGQGGPRGRPGYDGCNGTRGDAGPQGPYGSDGFPGPPGP
QGPKGQKGEPYALSREDRDRYRGESGEPGLVGYQGPPGRPGPVGQMGPVGAPGRPGPPGP
PGPKGQPGNRGLGF--YGEKGEKGDVGQPGPNGIPSDT-IHPIVGPTRTTILPDQYK
PF01391 // ENSGT00900000140804_1_Domain_2 (298, 433)
GEK
GS---EGEPGIKGI----SVKGEEGIVGFSGARGSPGF---DGEKGSPGQ----------
---------------------------------------------------------KGS
RGLDGFQGPD-GPRGPKGEAGDRGPPGLPAYSPHP-------------------------
-----------------------------------------SLAKGARGDPGFPGAYGEP
GSRGEPGDPGPPGL---P--------------------------GTSIIDED-EKRGLPG
EMGPKGFAGDPGVPARYP
PF01391 // ENSGT00900000140804_1_Domain_5 (619, 747)
GPPGADGKPGLQGRPGPAGEPGPD------------------
-GFLFGLKGIEGRVGFPGPSGFPGTRGQKGWKG---------------------------
-----------------------EAGENSGT00900000140804_1_Linker_5_6 (747, 760)
DCRCAE-DDEFVRPF01391 // ENSGT00900000140804_1_Domain_6 (760, 1015)
GVPGLPGLKGFPGVNGEPGRK
GEPGDPGQHGIPGFPGFKGAPGNVGV----------------------------------
------------------------------------------------------------
------------------------------------------------------------
-----------------------------------------PGP---KG--AKGDSRTIT
TKGERGQPGIPGVPGIKGDDGLPGREGLDGFPGL-----PGP-PGDGIKGPPGDQGYSGA
PGTKGVPGERGPPGLGLPGPKGERGFPGDAGLPGPSGFPGPPGPPGTPGQIDCD---TGV
KRPIGEDRQEAV--QPGCI--GGPK-----------------------GLPGLPGPP---
G----------------PTGAKGLRGIPGFPGADGGPGLKGFPGDPGREGFPGPP-GFVG
PRGSKGAVGLPGPDGPPGPIGLPGPAGPPGDRGLPGEVLGAQPGPRGDAGLPGHPGLKGL
PGDRGPPGFRGS-----QGMPGMPGLKGQSGFPGPSGQPGLPGPQGLHGF-PGNPGQEGP
LGQPGAP-GL--------------------GGLPGDRGEPGDTGVPGPVGMKGFSGDRGD
AGLSGERGHPGSPGFKGMVGMPGTPGLK-------
PF01391 // ENSGT00900000140804_1_Domain_13 (1476, 1538)
GDRGSPGMDGFQGMPGLKGRPGLPG
SKGEAGFFGI----PGLKGLAGEPGVKGSRGDPGPPGENSGT00900000140804_1_Linker_13_14 (1538, 1551)
PPPIILPG-MKDIPF01391 // ENSGT00900000140804_1_Domain_14 (1551, 1619)
KGEKGDEGPM
GLKGYLGLK-------------GTQGMPGIPGLSGIP-GLPGRPGYIKGA-KGDIGVPGV
PGLPGLPGVAGPPGITGFPGFTGSRGDKGAPGS---------------------------
---AGLYGEIGPTGDF---GDIGDTVDLPGRPGLKGERGSAGTP-GLKGFFGEKGIEGSI
GFPGIPGVPGVQGPPGLKGQTGFPGLTGLQGPQGEPGRIGPPG--SKGDDGWPGVPGLP-
---------------------------------FPGSRGISGLHGLPGTKGLPGSPGADI
YG----------D---PGFQGSPGDRGDPGEANTLPGPVGAPGQKGEQGAP---------
--------------------------GARGPAGSPGLQGFPGI-TPPSNISGSPGDKGA-
---PGIFGLEGYRGPPGPPGPAALPGGKGDAGNPGAPGVPGTKGWVGDP
PF01391 // ENSGT00900000140804_1_Domain_20 (2030, 2108)
GPQGRPGVFGL
PGEKG---------PRGEQGLMGNTGAT-----GTVGDRGPRGPKGDQGFPG-------A
PGPVGSPGIAGVPPRIAVQPGIVGPQGRRGLPGAQGEMGPQGPPGDP-------------
---------------------------------------GLRGEPGKAGPQGRGGVSAVP
GFRGDQGPVGHQGPTGQE----GEPGRPGSPGLPGMPGRSVSIGY
PF01413 // ENSGT00900000140804_1_Domain_23 (2266, 2371)
LLVKHSQTDQEPMCP
VGMNRLWSGYSLLYFEGQEKAHNQDLGLAGSCLARFSTMPFLYCNPGDVCYYASRNDKSY
WLSTTAPLPMMPVAEDD--IKPYISRCSVCENSGT00900000140804_1_Linker_23_25 (2371, 2376)
EAPAVPF01413 // ENSGT00900000140804_1_Domain_25 (2376, 2504)
AIAVHSQ--DVSIPHCPPGWRSLWI
GYSFLMHTAAGDEGGGQS----LVSPGSCLEDFRAT---PFIECNG-GRGTCHYYANKYS
--FWLTTIPEQSFQGSPSA-----DTLKAGLIRTHISRCQVCMKNL--------------
--------------
ENSSSCP00000010191 (view gene)
--------------MDRDP-RAASGPALRRWLLLGTVMVGLLAQSVLAGVKKLDVPCGGR
DCS-GGCQCYPEKGGR
PF01391 // ENSGT00900000140804_1_Domain_0 (77, 132)
GQPGPVGPQGYTGPPGLQGFPGLQGRKGDKGERGAPGITGPKGD
VGARGVSGFPGADGIP---GHPGQGGPRGRPGSDGCNGTVGDTGYAGPVGPDGFLGPPGP
QGPKGQKGEPYALSREDRDKYR
PF01391 // ENSGT00900000140804_1_Domain_1 (203, 251)
GEPGEPGLVGFQGPPGRPGPVGQMGPVGAPGRPGPPGP
PGPKGQPGNRENSGT00900000140804_1_Linker_1_2 (251, 322)
GLGF--YGEKGEKGDVGQPGPNGIPSDN-HHPIIGPTRETIYLDQYKGEK
GS---EGEPGRKGI----SLKPF01391 // ENSGT00900000140804_1_Domain_2 (322, 448)
GEEGIMGFSGSRGVPGF---DGEKGSPGQ----------
---------------------------------------------------------KGS
RGLDGYEGPD-GYPGPKGERGDPGPPGAPAYSPHP-------------------------
-----------------------------------------SLAKGARGEPGFPGALGEP
GARGEPGDPGPPGL---P--------------------------GTSVRDED-EKRGLPG
EMGPKGYAGEPGAPALYPGPPGADGKPGLRGPPGPPGPPGPD------------------
-DFLFGLKGAKGSMGYPGPSGFPGARGQKGWKG---------------------------
-----------------------DAGDC-KCAEDDQFVG
PF01391 // ENSGT00900000140804_1_Domain_6 (760, 1015)
GLPGPPGPKGFPGINGEPGRK
GSQGDPGQHGIPGFPGFKGAPGDAGP----------------------------------
------------------------------------------------------------
------------------------------------------------------------
-----------------------------------------PGP---KG--MKGDSRTIT
TKGERGQPGVPGVPGLRGDDGAPGRDGLDGFPGL-----PGP-PGDGIKGPPGDAGHPGV
PGTKGLAGDRGPPGLGLPGPKGERGFPGDDGLPGPPGFPGPPGPPGPPGQIDCD---SGV
KRPIGADGQEVI--QPGCV--GGPK-----------------------GSPGQPGPP---
G----------------PPGAKGLRGIPGPSGADGAP
PF01391 // ENSGT00900000140804_1_Domain_10 (1238, 1297)
GLKGFPGDPGREGFPGPP-GFVG
PRGSKGAVGPPGLDGLPGPSGLPGPVGPPGDKGLPGEVLGAQPGSRGDPGLPGHPGLKGP
PGERGAPGFRGS-----EGMPGMPGLKGQPGFPGPSGQPGLPGPPGQHGF-PGAPGREGP
LGPPGAP-GF--------------------GGLPGDRGDPGDTGVPGPVGMKGLSGDRGD
PGLLGERGHPGSPGFKGVAGMPGAPGPK-------GTRGSPGMHGFQGMLGLKGSPGLPG
SKGEAGFFGV----PGLKGLPGEPGVKGSRGDPGLPGPPPTILPG-MKDIKGEKGDEGPM
GLKGYLGLK-------------GIPGMPGIPGLSGVP-GLPGKPGHVK
PF01391 // ENSGT00900000140804_1_Domain_15 (1609, 1694)
GA-KGDTGAPGV
PGSPGFPGLPGPPGIIGFPGFTGSRGDKGSPGR---------------------------
---AGLYGEIGATGDF---GDIGDTIDLPGSPGLKGERGTAGVP-GLKGFFGEKGTVGDV
GFPGITGLAGVQGPPGLKGQTGFPGLTGLQGPQGDPGRAGVPG--AKGELGWPGNVGLP-
--------------------------------GLPGIRGISGLHGLPGTKGFPGSPGADV
HG----------D---PGFPGPAGDRGDPGEANTLPGPAGAPGQKGERGAP---------
--------------------------GERGPVGNPGLQGFPGI-TPPSNVSGLPGDTGA-
---PGIFGPEGYRGPPGPPGPAALPGSKGDEGLPGIPGNPGGKGWVGDP
PF01391 // ENSGT00900000140804_1_Domain_20 (2030, 2108)
GPQGRPGVFGL
PGEKG---------PRGEQGFMGNTGAT-----GSVGDRGPKGPKGDRGLPG------PP
-GAVGAPGIVGIPQRIAVQRGPVGPQGRRGPPGAQGEMGPQGPPGEP-------------
---------------------------------------GFRGAPGKAGPQGRGGVSAVP
GFRGDQGPVGQQGPVGQE----GEPGRPGSPGLPGMPGRSVSIGY
PF01413 // ENSGT00900000140804_1_Domain_23 (2266, 2371)
LLVKHSQTDQEPMCP
VGMNKLWSGYSLLYFEGQEKAHNQDLGLAGSCLARFSTMPFLYCNPGDICHYASRNDKSY
WLSTTAPLPMMPVAEEE--IRPYISRCSVCENSGT00900000140804_1_Linker_23_25 (2371, 2376)
EAPAVPF01413 // ENSGT00900000140804_1_Domain_25 (2376, 2504)
AIAVHSQ--DVSIPHCPAGWRSLWI
GYSFLMHTAAGDEGGGQS----LVSPGSCLEDFRAT---PFIECNG-ARGTCHYYANKYS
--FWLTTIPEQSFQSSPSA-----DTLKAGLIRTHISRCQVCMKNL--------------
--------------
{"ENSSBOP00000011138": [["Domain_0", 131, 190, "PF01391"], ["Linker_0_1", 190, 203], ["Domain_1", 203, 251, "PF01391"], ["Linker_1_2", 251, 322], ["Domain_2", 322, 447, "PF01391"], ["Domain_5", 619, 747, "PF01391"], ["Linker_5_6", 747, 760], ["Domain_6", 760, 1015, "PF01391"], ["Domain_10", 1240, 1297, "PF01391"], ["Domain_17", 1764, 1855, "PF01391"], ["Domain_19", 1972, 2034, "PF01391"], ["Domain_22", 2200, 2258, "PF01391"], ["Linker_22_23", 2258, 2266], ["Domain_23", 2266, 2371, "PF01413"], ["Linker_23_25", 2371, 2376], ["Domain_25", 2376, 2504, "PF01413"]], "ENSPEMP00000017918": [["Domain_0", 131, 189, "PF01391"], ["Linker_0_1", 189, 203], ["Domain_1", 203, 251, "PF01391"], ["Linker_1_2", 251, 322], ["Domain_2", 322, 449, "PF01391"], ["Domain_5", 619, 747, "PF01391"], ["Linker_5_6", 747, 760], ["Domain_6", 760, 1015, "PF01391"], ["Domain_13", 1476, 1538, "PF01391"], ["Domain_15", 1613, 1694, "PF01391"], ["Domain_20", 2033, 2111, "PF01391"], ["Domain_23", 2266, 2371, "PF01413"], ["Linker_23_25", 2371, 2376], ["Domain_25", 2376, 2504, "PF01413"]], "ENSRBIP00000027815": [["Domain_0", 131, 189, "PF01391"], ["Domain_2", 322, 448, "PF01391"], ["Domain_5", 619, 747, "PF01391"], ["Linker_5_6", 747, 760], ["Domain_6", 760, 1014, "PF01391"], ["Domain_12", 1418, 1482, "PF01391"], ["Domain_15", 1608, 1694, "PF01391"], ["Domain_20", 2030, 2108, "PF01391"], ["Domain_23", 2266, 2371, "PF01413"], ["Linker_23_25", 2371, 2376], ["Domain_25", 2376, 2504, "PF01413"]], "ENSFCAP00000013714": [["Domain_0", 77, 133, "PF01391"], ["Domain_1", 203, 251, "PF01391"], ["Linker_1_2", 251, 321], ["Domain_2", 321, 448, "PF01391"], ["Domain_5", 619, 747, "PF01391"], ["Linker_5_6", 747, 760], ["Domain_6", 760, 1015, "PF01391"], ["Domain_13", 1476, 1538, "PF01391"], ["Domain_17", 1710, 1765, "PF01391"], ["Domain_20", 2030, 2108, "PF01391"], ["Domain_23", 2266, 2371, "PF01413"], ["Linker_23_25", 2371, 2376], ["Domain_25", 2376, 2504, "PF01413"]], "ENSCJAP00000026472": [["Domain_0", 131, 190, "PF01391"], ["Domain_2", 321, 447, "PF01391"], ["Domain_5", 619, 747, "PF01391"], ["Linker_5_6", 747, 760], ["Domain_6", 760, 1015, "PF01391"], ["Domain_10", 1238, 1297, "PF01391"], ["Domain_17", 1710, 1769, "PF01391"], ["Domain_19", 1972, 2034, "PF01391"], ["Domain_23", 2266, 2371, "PF01413"], ["Linker_23_25", 2371, 2376], ["Domain_25", 2376, 2504, "PF01413"]], "ENSPPAP00000016802": [["Domain_0", 131, 190, "PF01391"], ["Linker_0_1", 190, 203], ["Domain_1", 203, 251, "PF01391"], ["Linker_1_2", 251, 322], ["Domain_2", 322, 447, "PF01391"], ["Domain_5", 619, 747, "PF01391"], ["Linker_5_6", 747, 760], ["Domain_6", 760, 1015, "PF01391"], ["Domain_12", 1348, 1427, "PF01391"], ["Domain_14", 1551, 1619, "PF01391"], ["Domain_20", 2030, 2108, "PF01391"], ["Domain_23", 2266, 2371, "PF01413"], ["Linker_23_25", 2371, 2376], ["Domain_25", 2376, 2504, "PF01413"]], "ENSMODP00000005712": [["Domain_0", 74, 132, "PF01391"], ["Domain_1", 203, 251, "PF01391"], ["Linker_1_2", 251, 322], ["Domain_2", 322, 448, "PF01391"], ["Linker_2_3", 448, 526], ["Domain_3", 526, 614, "PF01391"], ["Linker_3_5", 614, 619], ["Domain_5", 619, 747, "PF01391"], ["Linker_5_6", 747, 760], ["Domain_6", 760, 1015, "PF01391"], ["Linker_6_7", 1015, 1026], ["Domain_7", 1026, 1091, "PF01391"], ["Domain_9", 1071, 1129, "PF01391"], ["Domain_13", 1476, 1538, "PF01391"], ["Domain_17", 1794, 1892, "PF01391"], ["Domain_19", 1975, 2034, "PF01391"], ["Domain_21", 2120, 2227, "PF01391"], ["Linker_21_23", 2227, 2265], ["Domain_23", 2265, 2372, "PF01413"], ["Linker_23_25", 2372, 2375], ["Domain_25", 2375, 2505, "PF01413"]], "ENSGGOP00000012281": [["Domain_0", 77, 132, "PF01391"], ["Domain_1", 203, 251, "PF01391"], ["Linker_1_2", 251, 322], ["Domain_2", 322, 447, "PF01391"], ["Domain_5", 619, 747, "PF01391"], ["Linker_5_6", 747, 760], ["Domain_6", 760, 1015, "PF01391"], ["Domain_9", 1070, 1129, "PF01391"], ["Linker_9_10", 1129, 1238], ["Domain_10", 1238, 1297, "PF01391"], ["Domain_17", 1710, 1769, "PF01391"], ["Domain_19", 1972, 2034, "PF01391"], ["Domain_23", 2266, 2371, "PF01413"], ["Linker_23_25", 2371, 2376], ["Domain_25", 2376, 2504, "PF01413"]], "ENSDNOP00000013857": [["Domain_0", 77, 132, "PF01391"], ["Domain_1", 203, 251, "PF01391"], ["Linker_1_2", 251, 321], ["Domain_2", 321, 448, "PF01391"], ["Domain_5", 619, 747, "PF01391"], ["Linker_5_6", 747, 760], ["Domain_6", 760, 1015, "PF01391"], ["Domain_10", 1163, 1258, "PF01391"], ["Domain_15", 1613, 1703, "PF01391"], ["Domain_19", 1991, 2058, "PF01391"], ["Linker_19_21", 2058, 2121], ["Domain_21", 2121, 2216, "PF01391"], ["Linker_21_23", 2216, 2265], ["Domain_23", 2265, 2372, "PF01413"], ["Linker_23_25", 2372, 2375], ["Domain_25", 2375, 2505, "PF01413"]], "ENSRROP00000029286": [["Domain_0", 131, 189, "PF01391"], ["Domain_2", 322, 447, "PF01391"], ["Domain_5", 619, 747, "PF01391"], ["Linker_5_6", 747, 760], ["Domain_6", 760, 1014, "PF01391"], ["Domain_13", 1476, 1538, "PF01391"], ["Domain_15", 1608, 1694, "PF01391"], ["Domain_19", 1972, 2034, "PF01391"], ["Domain_23", 2266, 2371, "PF01413"], ["Linker_23_25", 2371, 2376], ["Domain_25", 2376, 2504, "PF01413"]], "ENSNGAP00000013591": [["Domain_0", 131, 189, "PF01391"], ["Linker_0_1", 189, 203], ["Domain_1", 203, 251, "PF01391"], ["Linker_1_2", 251, 322], ["Domain_2", 322, 449, "PF01391"], ["Domain_6", 760, 1015, "PF01391"], ["Linker_6_7", 1015, 1022], ["Domain_7", 1022, 1079, "PF01391"], ["Domain_9", 1067, 1127, "PF01391"], ["Linker_9_10", 1127, 1241], ["Domain_10", 1241, 1297, "PF01391"], ["Domain_15", 1613, 1702, "PF01391"], ["Domain_20", 2030, 2108, "PF01391"], ["Domain_23", 2266, 2371, "PF01413"], ["Linker_23_25", 2371, 2376], ["Domain_25", 2376, 2504, "PF01413"]], "ENSTGUP00000009053": [["Domain_23", 2265, 2372, "PF01413"], ["Linker_23_25", 2372, 2375], ["Domain_25", 2375, 2505, "PF01413"]], "ENSTSYP00000000129": [["Domain_0", 83, 142, "PF01391"], ["Domain_1", 203, 251, "PF01391"], ["Linker_1_2", 251, 322], ["Domain_2", 322, 448, "PF01391"], ["Domain_6", 762, 1015, "PF01391"], ["Domain_12", 1418, 1482, "PF01391"], ["Domain_15", 1613, 1700, "PF01391"], ["Domain_19", 1974, 2034, "PF01391"], ["Domain_23", 2266, 2371, "PF01413"], ["Linker_23_25", 2371, 2376], ["Domain_25", 2376, 2504, "PF01413"]], "ENSCPOP00000009528": [["Domain_0", 77, 132, "PF01391"], ["Domain_5", 619, 696, "PF01391"], ["Domain_13", 1476, 1538, "PF01391"], ["Domain_17", 1765, 1857, "PF01391"], ["Domain_19", 1991, 2061, "PF01391"], ["Domain_23", 2309, 2371, "PF01413"], ["Linker_23_25", 2371, 2376], ["Domain_25", 2376, 2504, "PF01413"]], "ENSHGLP00100016468": [["Domain_0", 77, 133, "PF01391"], ["Domain_5", 619, 747, "PF01391"], ["Linker_5_6", 747, 760], ["Domain_6", 760, 1015, "PF01391"], ["Domain_10", 1163, 1258, "PF01391"], ["Domain_14", 1550, 1606, "PF01391"], ["Domain_20", 2030, 2108, "PF01391"], ["Domain_23", 2266, 2371, "PF01413"], ["Linker_23_25", 2371, 2376], ["Domain_25", 2376, 2504, "PF01413"]], "ENSTGUP00000015565": [["Domain_12", 1413, 1469, "PF01391"]], "ENSFALP00000004596": [["Domain_12", 1412, 1467, "PF01391"], ["Domain_17", 1765, 1854, "PF01391"], ["Linker_17_18", 1854, 1896], ["Domain_18", 1896, 1988, "PF01391"], ["Domain_22", 2200, 2259, "PF01391"], ["Linker_22_23", 2259, 2265], ["Domain_23", 2265, 2372, "PF01413"], ["Linker_23_25", 2372, 2375], ["Domain_25", 2375, 2505, "PF01413"]], "ENSACAP00000013906": [["Domain_0", 116, 176, "PF01391"], ["Linker_0_1", 176, 201], ["Domain_1", 201, 251, "PF01391"], ["Linker_1_2", 251, 321], ["Domain_2", 321, 448, "PF01391"], ["Domain_5", 619, 746, "PF01391"], ["Linker_5_6", 746, 757], ["Domain_6", 757, 1015, "PF01391"], ["Domain_9", 1069, 1128, "PF01391"], ["Domain_12", 1354, 1427, "PF01391"], ["Domain_17", 1709, 1767, "PF01391"], ["Domain_20", 2030, 2108, "PF01391"], ["Linker_20_21", 2108, 2120], ["Domain_21", 2120, 2220, "PF01391"], ["Linker_21_23", 2220, 2265], ["Domain_23", 2265, 2372, "PF01413"], ["Linker_23_25", 2372, 2375], ["Domain_25", 2375, 2504, "PF01413"]], "ENSAMEP00000017981": [["Domain_0", 77, 136, "PF01391"], ["Domain_1", 206, 266, "PF01391"], ["Linker_1_2", 266, 320], ["Domain_2", 320, 449, "PF01391"], ["Linker_2_3", 449, 526], ["Domain_3", 526, 585, "PF01391"], ["Linker_3_4", 585, 591], ["Domain_4", 591, 617, "PF01391"], ["Linker_4_5", 617, 619], ["Domain_5", 619, 748, "PF01391"], ["Linker_5_6", 748, 760], ["Domain_6", 760, 1019, "PF01391"], ["Linker_6_7", 1019, 1022], ["Domain_7", 1022, 1065, "PF01391"], ["Linker_7_9", 1065, 1070], ["Domain_9", 1070, 1128, "PF01391"], ["Domain_12", 1389, 1468, "PF01391"], ["Domain_17", 1716, 1776, "PF01391"], ["Domain_18", 1906, 2005, "PF01391"], ["Domain_21", 2133, 2248, "PF01391"], ["Linker_21_23", 2248, 2264], ["Domain_23", 2264, 2373, "PF01413"], ["Linker_23_25", 2373, 2374], ["Domain_25", 2374, 2505, "PF01413"]], "ENSGALP00000027157": [["Domain_0", 129, 189, "PF01391"], ["Linker_0_1", 189, 202], ["Domain_1", 202, 251, "PF01391"], ["Linker_1_2", 251, 321], ["Domain_2", 321, 445, "PF01391"], ["Linker_2_3", 445, 526], ["Domain_3", 526, 614, "PF01391"], ["Linker_3_5", 614, 619], ["Domain_5", 619, 746, "PF01391"], ["Linker_5_6", 746, 759], ["Domain_6", 759, 1015, "PF01391"], ["Linker_6_7", 1015, 1023], ["Domain_7", 1023, 1089, "PF01391"], ["Domain_9", 1073, 1130, "PF01391"], ["Domain_11", 1303, 1364, "PF01391"], ["Domain_17", 1768, 1856, "PF01391"], ["Domain_20", 2030, 2108, "PF01391"], ["Linker_20_22", 2108, 2203], ["Domain_22", 2203, 2259, "PF01391"], ["Linker_22_23", 2259, 2266], ["Domain_23", 2266, 2371, "PF01413"], ["Linker_23_25", 2371, 2376], ["Domain_25", 2376, 2504, "PF01413"]], "ENSMAUP00000023031": [["Domain_0", 73, 119, "PF01391"], ["Domain_2", 334, 447, "PF01391"], ["Domain_6", 760, 1015, "PF01391"], ["Domain_9", 1069, 1127, "PF01391"], ["Linker_9_10", 1127, 1244], ["Domain_10", 1244, 1297, "PF01391"], ["Domain_14", 1566, 1643, "PF01391"], ["Domain_20", 2033, 2111, "PF01391"], ["Domain_23", 2266, 2308, "PF01413"]], "ENSMEUP00000012988": [["Domain_0", 77, 136, "PF01391"], ["Domain_1", 202, 244, "PF01391"], ["Domain_3", 529, 585, "PF01391"], ["Domain_6", 758, 1016, "PF01391"], ["Linker_6_7", 1016, 1022], ["Domain_7", 1022, 1061, "PF01391"], ["Linker_7_9", 1061, 1074], ["Domain_9", 1074, 1131, "PF01391"], ["Linker_9_10", 1131, 1192], ["Domain_10", 1192, 1271, "PF01391"], ["Domain_14", 1561, 1636, "PF01391"], ["Domain_18", 1896, 1968, "PF01391"], ["Domain_21", 2139, 2254, "PF01391"], ["Linker_21_23", 2254, 2264], ["Domain_23", 2264, 2373, "PF01413"], ["Linker_23_25", 2373, 2374], ["Domain_25", 2374, 2505, "PF01413"]], "ENSMGAP00000015755": [["Domain_0", 146, 190, "PF01391"], ["Linker_0_1", 190, 202], ["Domain_1", 202, 251, "PF01391"], ["Linker_1_2", 251, 320], ["Domain_2", 320, 449, "PF01391"], ["Linker_2_3", 449, 526], ["Domain_3", 526, 585, "PF01391"], ["Domain_5", 619, 748, "PF01391"], ["Linker_5_6", 748, 758], ["Domain_6", 758, 1016, "PF01391"], ["Linker_6_7", 1016, 1023], ["Domain_7", 1023, 1047, "PF01391"], ["Linker_7_9", 1047, 1073], ["Domain_9", 1073, 1131, "PF01391"], ["Linker_9_10", 1131, 1192], ["Domain_10", 1192, 1271, "PF01391"], ["Domain_14", 1551, 1604, "PF01391"], ["Domain_18", 1906, 2005, "PF01391"], ["Domain_22", 2200, 2263, "PF01391"], ["Linker_22_23", 2263, 2264], ["Domain_23", 2264, 2373, "PF01413"], ["Linker_23_25", 2373, 2374], ["Domain_25", 2374, 2505, "PF01413"]], "ENSOARP00000006991": [["Domain_0", 130, 189, "PF01391"], ["Linker_0_1", 189, 203], ["Domain_1", 203, 251, "PF01391"], ["Domain_6", 760, 1015, "PF01391"], ["Domain_11", 1303, 1364, "PF01391"], ["Domain_14", 1551, 1605, "PF01391"], ["Domain_19", 1972, 2034, "PF01391"], ["Domain_21", 2120, 2221, "PF01391"], ["Linker_21_23", 2221, 2265], ["Domain_23", 2265, 2330, "PF01413"], ["Domain_25", 2415, 2505, "PF01413"]], "ENSAPLP00000009420": [["Domain_0", 89, 151, "PF01391"], ["Linker_0_1", 151, 202], ["Domain_1", 202, 251, "PF01391"], ["Linker_1_2", 251, 317], ["Domain_2", 317, 446, "PF01391"], ["Linker_2_3", 446, 526], ["Domain_3", 526, 586, "PF01391"], ["Domain_5", 619, 748, "PF01391"], ["Linker_5_6", 748, 758], ["Domain_6", 758, 1016, "PF01391"], ["Linker_6_7", 1016, 1022], ["Domain_7", 1022, 1067, "PF01391"], ["Linker_7_9", 1067, 1073], ["Domain_9", 1073, 1131, "PF01391"], ["Domain_13", 1460, 1530, "PF01391"], ["Domain_15", 1613, 1702, "PF01391"], ["Domain_19", 1991, 2059, "PF01391"], ["Linker_19_21", 2059, 2139], ["Domain_21", 2139, 2254, "PF01391"], ["Linker_21_23", 2254, 2264], ["Domain_23", 2264, 2373, "PF01413"], ["Linker_23_25", 2373, 2374], ["Domain_25", 2374, 2505, "PF01413"]], "ENSNLEP00000009690": [["Domain_0", 77, 132, "PF01391"], ["Domain_1", 203, 251, "PF01391"], ["Linker_1_2", 251, 322], ["Domain_2", 322, 447, "PF01391"], ["Domain_5", 619, 746, "PF01391"], ["Linker_5_6", 746, 760], ["Domain_6", 760, 1015, "PF01391"], ["Domain_12", 1418, 1482, "PF01391"], ["Domain_17", 1710, 1765, "PF01391"], ["Domain_19", 1972, 2034, "PF01391"], ["Domain_23", 2308, 2371, "PF01413"], ["Linker_23_25", 2371, 2376], ["Domain_25", 2376, 2504, "PF01413"]], "ENSSTOP00000019851": [["Domain_0", 131, 190, "PF01391"], ["Linker_0_1", 190, 203], ["Domain_1", 203, 251, "PF01391"], ["Linker_1_2", 251, 297], ["Domain_2", 297, 425, "PF01391"], ["Domain_6", 760, 1015, "PF01391"], ["Domain_13", 1466, 1535, "PF01391"], ["Domain_17", 1750, 1837, "PF01391"], ["Domain_20", 2033, 2111, "PF01391"], ["Domain_23", 2266, 2371, "PF01413"], ["Linker_23_25", 2371, 2376], ["Domain_25", 2376, 2504, "PF01413"]], "ENSJJAP00000009510": [["Domain_0", 131, 189, "PF01391"], ["Domain_2", 322, 449, "PF01391"], ["Domain_6", 760, 1015, "PF01391"], ["Domain_9", 1069, 1127, "PF01391"], ["Domain_12", 1418, 1482, "PF01391"], ["Domain_17", 1764, 1854, "PF01391"], ["Domain_20", 2030, 2108, "PF01391"], ["Domain_23", 2266, 2371, "PF01413"], ["Linker_23_25", 2371, 2376], ["Domain_25", 2376, 2504, "PF01413"]], "ENSCCAP00000035349": [["Domain_0", 77, 134, "PF01391"], ["Domain_1", 203, 251, "PF01391"], ["Linker_1_2", 251, 322], ["Domain_2", 322, 447, "PF01391"], ["Domain_5", 619, 747, "PF01391"], ["Linker_5_6", 747, 760], ["Domain_6", 760, 1015, "PF01391"], ["Domain_13", 1476, 1538, "PF01391"], ["Domain_17", 1764, 1855, "PF01391"], ["Domain_19", 1972, 2034, "PF01391"], ["Domain_23", 2266, 2371, "PF01413"], ["Linker_23_25", 2371, 2376], ["Domain_25", 2376, 2504, "PF01413"]], "MGP_CAROLIEiJ_P0083735": [["Domain_0", 131, 189, "PF01391"], ["Linker_0_1", 189, 203], ["Domain_1", 203, 251, "PF01391"], ["Linker_1_2", 251, 321], ["Domain_2", 321, 448, "PF01391"], ["Domain_6", 760, 1014, "PF01391"], ["Domain_13", 1479, 1539, "PF01391"], ["Domain_15", 1613, 1701, "PF01391"], ["Domain_20", 2033, 2111, "PF01391"], ["Domain_23", 2266, 2371, "PF01413"], ["Linker_23_25", 2371, 2376], ["Domain_25", 2376, 2504, "PF01413"]], "ENSMPUP00000010555": [["Domain_0", 131, 190, "PF01391"], ["Linker_0_1", 190, 203], ["Domain_1", 203, 247, "PF01391"], ["Domain_3", 526, 613, "PF01391"], ["Linker_3_5", 613, 619], ["Domain_5", 619, 747, "PF01391"], ["Linker_5_6", 747, 760], ["Domain_6", 760, 1015, "PF01391"], ["Domain_11", 1303, 1364, "PF01391"], ["Domain_17", 1710, 1765, "PF01391"], ["Domain_19", 1973, 2034, "PF01391"], ["Domain_21", 2120, 2220, "PF01391"], ["Linker_21_23", 2220, 2265], ["Domain_23", 2265, 2372, "PF01413"], ["Linker_23_25", 2372, 2375], ["Domain_25", 2375, 2505, "PF01413"]], "ENSOANP00000024611": [["Domain_0", 128, 189, "PF01391"], ["Linker_0_1", 189, 203], ["Domain_1", 203, 251, "PF01391"], ["Domain_5", 619, 747, "PF01391"], ["Linker_5_6", 747, 760], ["Domain_6", 760, 1014, "PF01391"], ["Linker_6_7", 1014, 1023], ["Domain_7", 1023, 1085, "PF01391"], ["Domain_9", 1073, 1130, "PF01391"], ["Domain_11", 1302, 1363, "PF01391"], ["Domain_17", 1710, 1765, "PF01391"], ["Domain_22", 2203, 2259, "PF01391"], ["Linker_22_23", 2259, 2265], ["Domain_23", 2265, 2372, "PF01413"], ["Linker_23_25", 2372, 2375], ["Domain_25", 2375, 2505, "PF01413"]], "ENSHGLP00000002972": [["Domain_0", 77, 133, "PF01391"], ["Domain_2", 322, 448, "PF01391"], ["Domain_5", 619, 747, "PF01391"], ["Linker_5_6", 747, 760], ["Domain_6", 760, 1015, "PF01391"], ["Linker_6_7", 1015, 1022], ["Domain_7", 1022, 1080, "PF01391"], ["Domain_9", 1070, 1130, "PF01391"], ["Linker_9_10", 1130, 1163], ["Domain_10", 1163, 1257, "PF01391"], ["Domain_17", 1710, 1858, "PF01391"], ["Domain_19", 1975, 2036, "PF01391"], ["Domain_23", 2266, 2371, "PF01413"], ["Linker_23_25", 2371, 2376], ["Domain_25", 2376, 2504, "PF01413"]], "ENSPVAP00000005802": [["Domain_0", 134, 196, "PF01391"], ["Linker_0_1", 196, 218], ["Domain_1", 218, 278, "PF01391"], ["Linker_1_2", 278, 320], ["Domain_2", 320, 449, "PF01391"], ["Linker_2_3", 449, 526], ["Domain_3", 526, 585, "PF01391"], ["Domain_5", 619, 748, "PF01391"], ["Linker_5_6", 748, 758], ["Domain_6", 758, 1016, "PF01391"], ["Linker_6_7", 1016, 1022], ["Domain_7", 1022, 1067, "PF01391"], ["Linker_7_9", 1067, 1073], ["Domain_9", 1073, 1131, "PF01391"], ["Domain_12", 1348, 1429, "PF01391"], ["Domain_15", 1613, 1705, "PF01391"], ["Domain_20", 2039, 2119, "PF01391"], ["Linker_20_21", 2119, 2136], ["Domain_21", 2136, 2251, "PF01391"], ["Linker_21_23", 2251, 2264], ["Domain_23", 2264, 2306, "PF01413"], ["Domain_25", 2393, 2505, "PF01413"]], "ENSBTAP00000005916": [["Domain_0", 130, 189, "PF01391"], ["Linker_0_1", 189, 203], ["Domain_1", 203, 251, "PF01391"], ["Linker_1_2", 251, 322], ["Domain_2", 322, 448, "PF01391"], ["Linker_2_3", 448, 526], ["Domain_3", 526, 612, "PF01391"], ["Domain_5", 595, 662, "PF01391"], ["Linker_5_6", 662, 760], ["Domain_6", 760, 1015, "PF01391"], ["Domain_13", 1476, 1538, "PF01391"], ["Domain_15", 1609, 1694, "PF01391"], ["Domain_20", 2031, 2108, "PF01391"], ["Linker_20_21", 2108, 2120], ["Domain_21", 2120, 2221, "PF01391"], ["Linker_21_23", 2221, 2265], ["Domain_23", 2265, 2372, "PF01413"], ["Linker_23_25", 2372, 2375], ["Domain_25", 2375, 2505, "PF01413"]], "ENSLAFP00000007563": [["Domain_0", 146, 193, "PF01391"], ["Linker_0_1", 193, 206], ["Domain_1", 206, 266, "PF01391"], ["Linker_1_2", 266, 325], ["Domain_2", 325, 455, "PF01391"], ["Linker_2_3", 455, 526], ["Domain_3", 526, 585, "PF01391"], ["Domain_5", 619, 748, "PF01391"], ["Linker_5_6", 748, 758], ["Domain_6", 758, 1016, "PF01391"], ["Linker_6_7", 1016, 1022], ["Domain_7", 1022, 1067, "PF01391"], ["Linker_7_9", 1067, 1073], ["Domain_9", 1073, 1131, "PF01391"], ["Linker_9_10", 1131, 1163], ["Domain_10", 1163, 1265, "PF01391"], ["Domain_17", 1768, 1862, "PF01391"], ["Linker_17_18", 1862, 1906], ["Domain_18", 1906, 2005, "PF01391"], ["Domain_21", 2119, 2205, "PF01391"], ["Linker_21_23", 2205, 2264], ["Domain_23", 2264, 2373, "PF01413"], ["Linker_23_25", 2373, 2374], ["Domain_25", 2374, 2505, "PF01413"]], "ENSCHIP00000032096": [["Domain_0", 74, 127, "PF01391"], ["Domain_6", 760, 1015, "PF01391"], ["Linker_6_7", 1015, 1023], ["Domain_7", 1023, 1082, "PF01391"], ["Domain_9", 1070, 1127, "PF01391"], ["Domain_13", 1476, 1538, "PF01391"], ["Domain_15", 1613, 1700, "PF01391"], ["Domain_19", 1972, 2034, "PF01391"], ["Domain_23", 2266, 2371, "PF01413"], ["Linker_23_25", 2371, 2376], ["Domain_25", 2376, 2504, "PF01413"]], "MGP_PahariEiJ_P0046588": [["Domain_0", 131, 189, "PF01391"], ["Linker_0_1", 189, 203], ["Domain_1", 203, 251, "PF01391"], ["Linker_1_2", 251, 321], ["Domain_2", 321, 448, "PF01391"], ["Domain_6", 760, 1015, "PF01391"], ["Domain_9", 1068, 1127, "PF01391"], ["Linker_9_10", 1127, 1241], ["Domain_10", 1241, 1297, "PF01391"], ["Domain_15", 1613, 1700, "PF01391"], ["Domain_20", 2033, 2111, "PF01391"], ["Domain_23", 2266, 2371, "PF01413"], ["Linker_23_25", 2371, 2376], ["Domain_25", 2376, 2504, "PF01413"]], "ENSRNOP00000054270": [["Domain_0", 77, 134, "PF01391"], ["Domain_2", 322, 449, "PF01391"], ["Domain_6", 759, 1015, "PF01391"], ["Domain_9", 1069, 1127, "PF01391"], ["Domain_13", 1479, 1539, "PF01391"], ["Domain_17", 1710, 1765, "PF01391"], ["Domain_20", 2033, 2111, "PF01391"], ["Domain_23", 2266, 2371, "PF01413"], ["Linker_23_25", 2371, 2376], ["Domain_25", 2376, 2504, "PF01413"]], "ENSPANP00000001006": [["Domain_0", 131, 189, "PF01391"], ["Domain_2", 322, 447, "PF01391"], ["Domain_5", 619, 747, "PF01391"], ["Linker_5_6", 747, 760], ["Domain_6", 760, 1015, "PF01391"], ["Domain_13", 1476, 1538, "PF01391"], ["Domain_17", 1765, 1857, "PF01391"], ["Domain_20", 2030, 2108, "PF01391"], ["Domain_23", 2266, 2371, "PF01413"], ["Linker_23_25", 2371, 2376], ["Domain_25", 2376, 2504, "PF01413"]], "ENSCGRP00000009304": [["Domain_6", 760, 1015, "PF01391"], ["Domain_9", 1070, 1129, "PF01391"], ["Domain_12", 1348, 1428, "PF01391"], ["Domain_14", 1551, 1619, "PF01391"], ["Domain_20", 2027, 2106, "PF01391"], ["Domain_23", 2266, 2371, "PF01413"], ["Linker_23_25", 2371, 2376], ["Domain_25", 2376, 2504, "PF01413"]], "ENSDORP00000009566": [["Domain_0", 77, 132, "PF01391"], ["Domain_13", 1466, 1535, "PF01391"], ["Domain_17", 1709, 1802, "PF01391"], ["Domain_19", 1992, 2047, "PF01391"], ["Domain_22", 2200, 2258, "PF01391"], ["Linker_22_23", 2258, 2266], ["Domain_23", 2266, 2371, "PF01413"], ["Linker_23_25", 2371, 2376], ["Domain_25", 2376, 2504, "PF01413"]], "ENSMFAP00000004048": [["Domain_0", 77, 132, "PF01391"], ["Domain_1", 203, 251, "PF01391"], ["Linker_1_2", 251, 322], ["Domain_2", 322, 447, "PF01391"], ["Domain_5", 619, 747, "PF01391"], ["Linker_5_6", 747, 760], ["Domain_6", 760, 1015, "PF01391"], ["Domain_12", 1348, 1428, "PF01391"], ["Domain_15", 1608, 1694, "PF01391"], ["Domain_20", 2030, 2108, "PF01391"], ["Domain_23", 2266, 2371, "PF01413"], ["Linker_23_25", 2371, 2376], ["Domain_25", 2376, 2504, "PF01413"]], "ENSTBEP00000004421": [["Domain_0", 77, 136, "PF01391"], ["Domain_2", 322, 422, "PF01391"], ["Domain_6", 799, 1064, "PF01391"], ["Linker_6_9", 1064, 1070], ["Domain_9", 1070, 1128, "PF01391"], ["Domain_11", 1275, 1333, "PF01391"], ["Domain_17", 1765, 1859, "PF01391"], ["Domain_23", 2288, 2373, "PF01413"], ["Linker_23_24", 2373, 2374], ["Domain_24", 2374, 2406, "PF01413"]], "ENSCHOP00000001082": [["Domain_0", 83, 145, "PF01391"], ["Domain_1", 206, 266, "PF01391"], ["Linker_1_2", 266, 298], ["Domain_2", 298, 434, "PF01391"], ["Domain_6", 758, 1016, "PF01391"], ["Linker_6_7", 1016, 1022], ["Domain_7", 1022, 1058, "PF01391"], ["Domain_10", 1247, 1297, "PF01391"], ["Domain_14", 1551, 1607, "PF01391"], ["Domain_20", 2027, 2107, "PF01391"], ["Linker_20_21", 2107, 2136], ["Domain_21", 2136, 2251, "PF01391"], ["Linker_21_23", 2251, 2264], ["Domain_23", 2264, 2373, "PF01413"], ["Linker_23_25", 2373, 2374], ["Domain_25", 2374, 2505, "PF01413"]], "ENSOGAP00000009095": [["Domain_0", 131, 190, "PF01391"], ["Linker_0_1", 190, 203], ["Domain_1", 203, 246, "PF01391"], ["Linker_1_2", 246, 297], ["Domain_2", 297, 425, "PF01391"], ["Domain_5", 619, 747, "PF01391"], ["Linker_5_6", 747, 760], ["Domain_6", 760, 1015, "PF01391"], ["Domain_10", 1163, 1257, "PF01391"], ["Domain_14", 1551, 1619, "PF01391"], ["Domain_18", 1945, 2002, "PF01391"], ["Domain_21", 2120, 2220, "PF01391"], ["Linker_21_23", 2220, 2265], ["Domain_23", 2265, 2372, "PF01413"], ["Linker_23_25", 2372, 2375], ["Domain_25", 2375, 2505, "PF01413"]], "ENSSHAP00000011227": [["Domain_0", 74, 132, "PF01391"], ["Domain_1", 203, 251, "PF01391"], ["Linker_1_2", 251, 322], ["Domain_2", 322, 448, "PF01391"], ["Domain_5", 596, 662, "PF01391"], ["Linker_5_6", 662, 759], ["Domain_6", 759, 1015, "PF01391"], ["Linker_6_7", 1015, 1022], ["Domain_7", 1022, 1079, "PF01391"], ["Domain_9", 1073, 1130, "PF01391"], ["Linker_9_10", 1130, 1188], ["Domain_10", 1188, 1257, "PF01391"], ["Domain_17", 1709, 1765, "PF01391"], ["Domain_19", 1991, 2058, "PF01391"], ["Linker_19_22", 2058, 2203], ["Domain_22", 2203, 2259, "PF01391"], ["Linker_22_23", 2259, 2265], ["Domain_23", 2265, 2372, "PF01413"], ["Linker_23_25", 2372, 2375], ["Domain_25", 2375, 2500, "PF01413"]], "ENSMOCP00000024736": [["Domain_0", 73, 143, "PF01391"], ["Domain_1", 203, 251, "PF01391"], ["Linker_1_2", 251, 322], ["Domain_2", 322, 448, "PF01391"], ["Domain_6", 760, 1014, "PF01391"], ["Domain_12", 1348, 1428, "PF01391"], ["Domain_15", 1609, 1694, "PF01391"], ["Domain_19", 1994, 2060, "PF01391"], ["Domain_23", 2266, 2371, "PF01413"], ["Linker_23_25", 2371, 2376], ["Domain_25", 2376, 2504, "PF01413"]], "ENSMMUP00000029330": [["Domain_0", 131, 189, "PF01391"], ["Linker_0_1", 189, 203], ["Domain_1", 203, 251, "PF01391"], ["Linker_1_2", 251, 322], ["Domain_2", 322, 447, "PF01391"], ["Domain_5", 619, 747, "PF01391"], ["Linker_5_6", 747, 760], ["Domain_6", 760, 1015, "PF01391"], ["Domain_12", 1348, 1428, "PF01391"], ["Domain_17", 1765, 1857, "PF01391"], ["Domain_20", 2030, 2108, "PF01391"], ["Domain_23", 2266, 2371, "PF01413"], ["Linker_23_25", 2371, 2376], ["Domain_25", 2376, 2504, "PF01413"]], "ENSCATP00000017946": [["Domain_0", 131, 189, "PF01391"], ["Domain_2", 322, 447, "PF01391"], ["Domain_5", 619, 747, "PF01391"], ["Linker_5_6", 747, 760], ["Domain_6", 760, 1015, "PF01391"], ["Domain_13", 1476, 1538, "PF01391"], ["Domain_17", 1765, 1848, "PF01391"], ["Domain_19", 1972, 2034, "PF01391"], ["Domain_23", 2266, 2371, "PF01413"], ["Linker_23_25", 2371, 2376], ["Domain_25", 2376, 2504, "PF01413"]], "ENSPSIP00000014136": [["Domain_0", 77, 134, "PF01391"], ["Domain_1", 203, 251, "PF01391"], ["Linker_1_2", 251, 298], ["Domain_2", 298, 419, "PF01391"], ["Domain_5", 596, 663, "PF01391"], ["Linker_5_6", 663, 760], ["Domain_6", 760, 1015, "PF01391"], ["Domain_12", 1418, 1487, "PF01391"], ["Domain_20", 2057, 2109, "PF01391"], ["Linker_20_22", 2109, 2200], ["Domain_22", 2200, 2259, "PF01391"], ["Linker_22_23", 2259, 2265], ["Domain_23", 2265, 2372, "PF01413"], ["Linker_23_25", 2372, 2375], ["Domain_25", 2375, 2505, "PF01413"]], "ENSXETP00000005650": [["Domain_0", 77, 136, "PF01391"], ["Domain_1", 203, 263, "PF01391"], ["Linker_1_2", 263, 310], ["Domain_2", 310, 440, "PF01391"], ["Linker_2_3", 440, 526], ["Domain_3", 526, 555, "PF01391"], ["Domain_5", 669, 783, "PF01391"], ["Linker_5_6", 783, 787], ["Domain_6", 787, 1046, "PF01391"], ["Linker_6_8", 1046, 1067], ["Domain_8", 1067, 1094, "PF01391"], ["Linker_8_9", 1094, 1095], ["Domain_9", 1095, 1125, "PF01391"], ["Domain_12", 1369, 1468, "PF01391"], ["Domain_14", 1551, 1602, "PF01391"], ["Domain_18", 1900, 2026, "PF01391"], ["Domain_21", 2139, 2254, "PF01391"], ["Linker_21_23", 2254, 2264], ["Domain_23", 2264, 2373, "PF01413"], ["Linker_23_25", 2373, 2374], ["Domain_25", 2374, 2505, "PF01413"]], "ENSPTRP00000058373": [["Domain_0", 131, 190, "PF01391"], ["Linker_0_1", 190, 203], ["Domain_1", 203, 251, "PF01391"], ["Linker_1_2", 251, 322], ["Domain_2", 322, 447, "PF01391"], ["Domain_5", 619, 747, "PF01391"], ["Linker_5_6", 747, 760], ["Domain_6", 760, 1015, "PF01391"], ["Domain_13", 1476, 1538, "PF01391"], ["Domain_17", 1765, 1856, "PF01391"], ["Domain_19", 1972, 2034, "PF01391"], ["Domain_23", 2266, 2371, "PF01413"], ["Linker_23_25", 2371, 2376], ["Domain_25", 2376, 2504, "PF01413"]], "ENSCLAP00000003695": [["Domain_0", 77, 132, "PF01391"], ["Domain_1", 203, 251, "PF01391"], ["Linker_1_2", 251, 321], ["Domain_2", 321, 448, "PF01391"], ["Domain_5", 619, 747, "PF01391"], ["Linker_5_6", 747, 763], ["Domain_6", 763, 1014, "PF01391"], ["Domain_12", 1418, 1482, "PF01391"], ["Domain_14", 1551, 1619, "PF01391"], ["Domain_19", 1975, 2036, "PF01391"], ["Domain_23", 2266, 2371, "PF01413"], ["Linker_23_25", 2371, 2376], ["Domain_25", 2376, 2504, "PF01413"]], "ENSETEP00000000138": [["Domain_0", 128, 190, "PF01391"], ["Linker_0_1", 190, 202], ["Domain_1", 202, 232, "PF01391"], ["Linker_1_2", 232, 320], ["Domain_2", 320, 449, "PF01391"], ["Linker_2_3", 449, 524], ["Domain_3", 524, 585, "PF01391"], ["Domain_6", 758, 1016, "PF01391"], ["Linker_6_7", 1016, 1023], ["Domain_7", 1023, 1087, "PF01391"], ["Domain_10", 1189, 1268, "PF01391"], ["Linker_10_11", 1268, 1303], ["Domain_11", 1303, 1331, "PF01391"], ["Domain_17", 1769, 1862, "PF01391"], ["Domain_20", 2039, 2119, "PF01391"], ["Linker_20_21", 2119, 2133], ["Domain_21", 2133, 2248, "PF01391"], ["Linker_21_23", 2248, 2264], ["Domain_23", 2264, 2373, "PF01413"], ["Linker_23_25", 2373, 2374], ["Domain_25", 2374, 2505, "PF01413"]], "ENSPPYP00000006266": [["Domain_0", 77, 132, "PF01391"], ["Domain_1", 203, 251, "PF01391"], ["Linker_1_2", 251, 298], ["Domain_2", 298, 449, "PF01391"], ["Domain_5", 619, 747, "PF01391"], ["Linker_5_6", 747, 760], ["Domain_6", 760, 1015, "PF01391"], ["Domain_13", 1476, 1538, "PF01391"], ["Linker_13_14", 1538, 1551], ["Domain_14", 1551, 1619, "PF01391"], ["Domain_19", 1972, 2034, "PF01391"], ["Domain_21", 2120, 2217, "PF01391"], ["Linker_21_23", 2217, 2265], ["Domain_23", 2265, 2372, "PF01413"], ["Linker_23_25", 2372, 2375], ["Domain_25", 2375, 2505, "PF01413"]], "ENSMLUP00000016067": [["Domain_0", 77, 136, "PF01391"], ["Domain_1", 202, 251, "PF01391"], ["Linker_1_2", 251, 320], ["Domain_2", 320, 449, "PF01391"], ["Linker_2_3", 449, 525], ["Domain_3", 525, 558, "PF01391"], ["Domain_5", 619, 748, "PF01391"], ["Linker_5_6", 748, 758], ["Domain_6", 758, 1016, "PF01391"], ["Linker_6_7", 1016, 1026], ["Domain_7", 1026, 1066, "PF01391"], ["Linker_7_9", 1066, 1073], ["Domain_9", 1073, 1131, "PF01391"], ["Linker_9_10", 1131, 1163], ["Domain_10", 1163, 1265, "PF01391"], ["Domain_14", 1589, 1651, "PF01391"], ["Domain_18", 1900, 1999, "PF01391"], ["Domain_21", 2121, 2232, "PF01391"], ["Linker_21_23", 2232, 2264], ["Domain_23", 2264, 2373, "PF01413"], ["Linker_23_25", 2373, 2374], ["Domain_25", 2374, 2505, "PF01413"]], "ENSMUSP00000033899": [["Domain_0", 131, 189, "PF01391"], ["Domain_2", 321, 448, "PF01391"], ["Domain_6", 760, 1014, "PF01391"], ["Domain_9", 1068, 1127, "PF01391"], ["Domain_12", 1418, 1482, "PF01391"], ["Domain_14", 1550, 1619, "PF01391"], ["Domain_19", 1978, 2035, "PF01391"], ["Domain_23", 2266, 2371, "PF01413"], ["Linker_23_25", 2371, 2376], ["Domain_25", 2376, 2504, "PF01413"]], "ENSTTRP00000005937": [["Domain_0", 134, 196, "PF01391"], ["Domain_2", 325, 455, "PF01391"], ["Linker_2_3", 455, 524], ["Domain_3", 524, 558, "PF01391"], ["Domain_5", 619, 748, "PF01391"], ["Linker_5_6", 748, 758], ["Domain_6", 758, 1016, "PF01391"], ["Linker_6_7", 1016, 1022], ["Domain_7", 1022, 1065, "PF01391"], ["Domain_9", 1096, 1128, "PF01391"], ["Domain_13", 1454, 1524, "PF01391"], ["Domain_16", 1621, 1651, "PF01391"], ["Linker_16_17", 1651, 1765], ["Domain_17", 1765, 1859, "PF01391"], ["Domain_19", 1972, 2035, "PF01391"], ["Domain_21", 2124, 2235, "PF01391"], ["Linker_21_23", 2235, 2264], ["Domain_23", 2264, 2373, "PF01413"], ["Linker_23_25", 2373, 2374], ["Domain_25", 2374, 2505, "PF01413"]], "ENSTGUP00000009048": [["Domain_7", 1016, 1095, "PF01391"], ["Domain_13", 1485, 1540, "PF01391"], ["Domain_14", 1537, 1652, "PF01391"]], "ENSCANP00000013008": [["Domain_0", 131, 189, "PF01391"], ["Linker_0_1", 189, 203], ["Domain_1", 203, 251, "PF01391"], ["Linker_1_2", 251, 322], ["Domain_2", 322, 447, "PF01391"], ["Domain_5", 619, 747, "PF01391"], ["Linker_5_6", 747, 760], ["Domain_6", 760, 1014, "PF01391"], ["Domain_12", 1348, 1427, "PF01391"], ["Domain_17", 1710, 1767, "PF01391"], ["Domain_19", 1972, 2034, "PF01391"], ["Domain_23", 2266, 2371, "PF01413"], ["Linker_23_25", 2371, 2376], ["Domain_25", 2376, 2504, "PF01413"]], "ENSECAP00000018971": [["Domain_0", 77, 134, "PF01391"], ["Domain_1", 203, 251, "PF01391"], ["Linker_1_2", 251, 322], ["Domain_2", 322, 448, "PF01391"], ["Linker_2_3", 448, 526], ["Domain_3", 526, 613, "PF01391"], ["Linker_3_5", 613, 619], ["Domain_5", 619, 747, "PF01391"], ["Linker_5_6", 747, 760], ["Domain_6", 760, 1015, "PF01391"], ["Domain_11", 1303, 1364, "PF01391"], ["Domain_15", 1613, 1701, "PF01391"], ["Domain_19", 1972, 2034, "PF01391"], ["Domain_21", 2120, 2220, "PF01391"], ["Linker_21_23", 2220, 2265], ["Domain_23", 2265, 2372, "PF01413"], ["Linker_23_25", 2372, 2375], ["Domain_25", 2375, 2505, "PF01413"]], "ENSOCUP00000011443": [["Domain_0", 131, 190, "PF01391"], ["Linker_0_1", 190, 212], ["Domain_1", 212, 271, "PF01391"], ["Linker_1_2", 271, 322], ["Domain_2", 322, 448, "PF01391"], ["Linker_2_3", 448, 526], ["Domain_3", 526, 613, "PF01391"], ["Linker_3_5", 613, 619], ["Domain_5", 619, 744, "PF01391"], ["Linker_5_6", 744, 760], ["Domain_6", 760, 1015, "PF01391"], ["Domain_10", 1238, 1297, "PF01391"], ["Domain_17", 1785, 1891, "PF01391"], ["Domain_20", 2030, 2108, "PF01391"], ["Domain_23", 2265, 2372, "PF01413"], ["Linker_23_25", 2372, 2375], ["Domain_25", 2375, 2505, "PF01413"]], "ENSFALP00000011009": [["Domain_0", 138, 189, "PF01391"], ["Linker_0_1", 189, 202], ["Domain_1", 202, 251, "PF01391"], ["Domain_5", 678, 797, "PF01391"], ["Linker_5_7", 797, 1014], ["Domain_7", 1014, 1065, "PF01391"], ["Linker_7_9", 1065, 1070], ["Domain_9", 1070, 1128, "PF01391"], ["Domain_13", 1460, 1534, "PF01391"]], "ENSCAPP00000010546": [["Domain_0", 77, 134, "PF01391"], ["Domain_6", 762, 1015, "PF01391"], ["Domain_10", 1223, 1275, "PF01391"], ["Domain_17", 1765, 1855, "PF01391"], ["Domain_19", 1992, 2058, "PF01391"], ["Domain_23", 2308, 2371, "PF01413"], ["Linker_23_25", 2371, 2376], ["Domain_25", 2376, 2504, "PF01413"]], "ENSOPRP00000012047": [["Domain_0", 83, 145, "PF01391"], ["Domain_1", 202, 237, "PF01391"], ["Domain_3", 527, 587, "PF01391"], ["Domain_6", 760, 1019, "PF01391"], ["Linker_6_7", 1019, 1023], ["Domain_7", 1023, 1058, "PF01391"], ["Domain_12", 1413, 1453, "PF01391"], ["Domain_14", 1558, 1605, "PF01391"], ["Domain_18", 1906, 2005, "PF01391"], ["Domain_21", 2139, 2254, "PF01391"], ["Linker_21_23", 2254, 2264], ["Domain_23", 2264, 2373, "PF01413"], ["Linker_23_24", 2373, 2374], ["Domain_24", 2374, 2406, "PF01413"]], "ENSMICP00000040801": [["Domain_0", 131, 190, "PF01391"], ["Linker_0_1", 190, 203], ["Domain_1", 203, 251, "PF01391"], ["Linker_1_2", 251, 321], ["Domain_2", 321, 447, "PF01391"], ["Domain_6", 760, 1015, "PF01391"], ["Domain_12", 1418, 1482, "PF01391"], ["Domain_17", 1765, 1857, "PF01391"], ["Domain_20", 2030, 2108, "PF01391"], ["Domain_23", 2266, 2371, "PF01413"], ["Linker_23_25", 2371, 2376], ["Domain_25", 2376, 2504, "PF01413"]], "ENSP00000353654": [["Domain_0", 77, 132, "PF01391"], ["Domain_1", 203, 251, "PF01391"], ["Linker_1_2", 251, 322], ["Domain_2", 322, 447, "PF01391"], ["Domain_5", 619, 747, "PF01391"], ["Linker_5_6", 747, 760], ["Domain_6", 760, 1015, "PF01391"], ["Domain_10", 1163, 1261, "PF01391"], ["Domain_17", 1765, 1857, "PF01391"], ["Domain_19", 1972, 2034, "PF01391"], ["Domain_23", 2266, 2371, "PF01413"], ["Linker_23_25", 2371, 2376], ["Domain_25", 2376, 2504, "PF01413"]], "ENSCGRP00001022430": [["Domain_0", 77, 132, "PF01391"], ["Domain_1", 203, 251, "PF01391"], ["Linker_1_2", 251, 322], ["Domain_2", 322, 448, "PF01391"], ["Domain_6", 760, 1015, "PF01391"], ["Domain_13", 1476, 1538, "PF01391"], ["Domain_17", 1764, 1855, "PF01391"], ["Domain_20", 2027, 2106, "PF01391"], ["Domain_23", 2266, 2371, "PF01413"], ["Linker_23_25", 2371, 2376], ["Domain_25", 2376, 2504, "PF01413"]], "ENSCSAP00000013500": [["Domain_0", 131, 189, "PF01391"], ["Domain_2", 322, 447, "PF01391"], ["Linker_2_3", 447, 525], ["Domain_3", 525, 612, "PF01391"], ["Domain_5", 596, 662, "PF01391"], ["Linker_5_6", 662, 760], ["Domain_6", 760, 1015, "PF01391"], ["Domain_13", 1476, 1538, "PF01391"], ["Domain_15", 1608, 1694, "PF01391"], ["Domain_20", 2030, 2108, "PF01391"], ["Linker_20_22", 2108, 2203], ["Domain_22", 2203, 2259, "PF01391"], ["Linker_22_23", 2259, 2265], ["Domain_23", 2265, 2372, "PF01413"], ["Linker_23_24", 2372, 2375], ["Domain_24", 2375, 2409, "PF01413"]], "ENSMNEP00000004318": [["Domain_0", 131, 189, "PF01391"], ["Linker_0_1", 189, 203], ["Domain_1", 203, 251, "PF01391"], ["Linker_1_2", 251, 322], ["Domain_2", 322, 447, "PF01391"], ["Domain_5", 619, 747, "PF01391"], ["Linker_5_6", 747, 760], ["Domain_6", 760, 1015, "PF01391"], ["Domain_13", 1476, 1538, "PF01391"], ["Domain_17", 1710, 1765, "PF01391"], ["Domain_20", 2030, 2108, "PF01391"], ["Domain_23", 2266, 2371, "PF01413"], ["Linker_23_24", 2371, 2376], ["Domain_24", 2376, 2409, "PF01413"]], "ENSPCAP00000014473": [["Domain_0", 132, 193, "PF01391"], ["Linker_0_1", 193, 202], ["Domain_1", 202, 251, "PF01391"], ["Linker_1_2", 251, 334], ["Domain_2", 334, 449, "PF01391"], ["Domain_5", 619, 663, "PF01391"], ["Domain_9", 1073, 1131, "PF01391"], ["Domain_13", 1476, 1539, "PF01391"], ["Linker_13_14", 1539, 1555], ["Domain_14", 1555, 1605, "PF01391"], ["Domain_18", 1900, 1999, "PF01391"], ["Domain_22", 2199, 2254, "PF01391"], ["Linker_22_23", 2254, 2264], ["Domain_23", 2264, 2306, "PF01413"], ["Domain_25", 2405, 2505, "PF01413"]], "ENSODEP00000016445": [["Domain_0", 131, 190, "PF01391"], ["Linker_0_1", 190, 203], ["Domain_1", 203, 251, "PF01391"], ["Linker_1_2", 251, 322], ["Domain_2", 322, 448, "PF01391"], ["Domain_5", 619, 747, "PF01391"], ["Linker_5_6", 747, 762], ["Domain_6", 762, 1014, "PF01391"], ["Domain_13", 1476, 1538, "PF01391"], ["Domain_17", 1765, 1858, "PF01391"], ["Domain_20", 2030, 2108, "PF01391"], ["Domain_23", 2266, 2371, "PF01413"], ["Linker_23_25", 2371, 2376], ["Domain_25", 2376, 2504, "PF01413"]], "ENSANAP00000018465": [["Domain_0", 131, 189, "PF01391"], ["Domain_5", 619, 747, "PF01391"], ["Linker_5_6", 747, 760], ["Domain_6", 760, 1015, "PF01391"], ["Domain_9", 1070, 1129, "PF01391"], ["Domain_13", 1476, 1537, "PF01391"], ["Domain_15", 1609, 1694, "PF01391"], ["Domain_20", 2030, 2108, "PF01391"], ["Domain_23", 2266, 2371, "PF01413"], ["Linker_23_25", 2371, 2376], ["Domain_25", 2376, 2504, "PF01413"]], "ENSCAFP00000009112": [["Domain_0", 83, 143, "PF01391"], ["Domain_1", 203, 251, "PF01391"], ["Linker_1_2", 251, 298], ["Domain_2", 298, 436, "PF01391"], ["Linker_2_3", 436, 526], ["Domain_3", 526, 613, "PF01391"], ["Domain_5", 593, 662, "PF01391"], ["Linker_5_6", 662, 760], ["Domain_6", 760, 1015, "PF01391"], ["Linker_6_7", 1015, 1026], ["Domain_7", 1026, 1092, "PF01391"], ["Domain_9", 1070, 1127, "PF01391"], ["Domain_13", 1467, 1535, "PF01391"], ["Domain_15", 1613, 1703, "PF01391"], ["Domain_20", 2033, 2111, "PF01391"], ["Linker_20_21", 2111, 2120], ["Domain_21", 2120, 2221, "PF01391"], ["Linker_21_23", 2221, 2265], ["Domain_23", 2265, 2372, "PF01413"], ["Linker_23_25", 2372, 2375], ["Domain_25", 2375, 2505, "PF01413"]], "ENSMLEP00000037549": [["Domain_0", 131, 189, "PF01391"], ["Linker_0_1", 189, 203], ["Domain_1", 203, 251, "PF01391"], ["Linker_1_2", 251, 321], ["Domain_2", 321, 447, "PF01391"], ["Domain_5", 619, 747, "PF01391"], ["Linker_5_6", 747, 760], ["Domain_6", 760, 1015, "PF01391"], ["Linker_6_7", 1015, 1023], ["Domain_7", 1023, 1082, "PF01391"], ["Domain_9", 1070, 1129, "PF01391"], ["Domain_12", 1348, 1428, "PF01391"], ["Domain_15", 1608, 1694, "PF01391"], ["Domain_20", 2030, 2108, "PF01391"], ["Domain_23", 2266, 2371, "PF01413"], ["Linker_23_25", 2371, 2376], ["Domain_25", 2376, 2504, "PF01413"]], "ENSPCOP00000017693": [["Domain_0", 131, 190, "PF01391"], ["Domain_2", 298, 433, "PF01391"], ["Domain_5", 619, 747, "PF01391"], ["Linker_5_6", 747, 760], ["Domain_6", 760, 1015, "PF01391"], ["Domain_13", 1476, 1538, "PF01391"], ["Linker_13_14", 1538, 1551], ["Domain_14", 1551, 1619, "PF01391"], ["Domain_20", 2030, 2108, "PF01391"], ["Domain_23", 2266, 2371, "PF01413"], ["Linker_23_25", 2371, 2376], ["Domain_25", 2376, 2504, "PF01413"]], "ENSSSCP00000010191": [["Domain_0", 77, 132, "PF01391"], ["Domain_1", 203, 251, "PF01391"], ["Linker_1_2", 251, 322], ["Domain_2", 322, 448, "PF01391"], ["Domain_6", 760, 1015, "PF01391"], ["Domain_10", 1238, 1297, "PF01391"], ["Domain_15", 1609, 1694, "PF01391"], ["Domain_20", 2030, 2108, "PF01391"], ["Domain_23", 2266, 2371, "PF01413"], ["Linker_23_25", 2371, 2376], ["Domain_25", 2376, 2504, "PF01413"]]}

