Home
Query Results
"ENSGT00730000111141_0"
Found:
ENSGT00730000111141_0
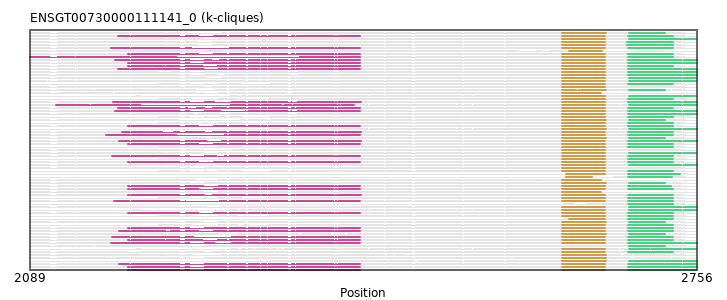
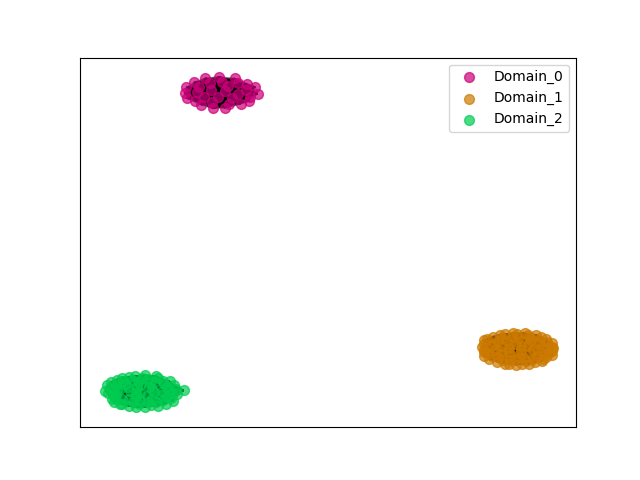
ENSHGLP00000014574 (view gene)
ENSGALP00000053471 (view gene)
ENSMGAP00000015381 (view gene)
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------RRGYSRSPDRRR
--SRSSERFS-SRSPKRLTDLDKAQLLEIAKANAAAMCAKSGVPLPPSLMPMLSQ-KK-D
DKASQKSSRDTLKELTEKCKKIAQST--DDVIVNKPHVSDEE-EEERPFYNHPFKLNEPK
PIFFNLSTPSIKPAPPPQPKNQVSLSKEFPVSSGSQHRKKEADSVYGEWVPVEK-GKEDS
--KDDVFPKP-SIEGVDITAAMNDRALAQIQFFESSDLLSVRRVITVTENRKIDAWAQSN
SIPGQFTGSTGAQILSSDELTNSGPQAWIRKDQFLRAAPV
PF01585 // ENSGT00730000111141_0_Domain_1 (2621, 2665)
TGGMGAQLMRKMGWREGEGL
GKNKEGSVEPIMVDFKTDRKGLVAENSGT00730000111141_0_Linker_1_2 (2665, 2688)
VGEKAQKRSGHYV-VMKDLSGKHPF00035 // ENSGT00730000111141_0_Domain_2 (2688, 2756)
PVSALMEICNKRR
WSPPEFVLVDDSGPDHRKHFIFKVRVNGNEYRPTFASLNK-KHAKATAATAALQAMGLVP
KESMVNTTMFRSASHR
ENSFDAP00000011600 (view gene)
ENSSARP00000011196 (view gene)
MATNIEQIFRSFVVSKFREIQQELSSGRSEGQLNGETNTPIEGNQAGDA-AASARNLPNE
EIVQKIEEVLSGVLDTELRHKPDMKEAS-RK-RCVSVQTDPTD-EIPSKKSKKHKKHKNK
KKKKK----KEKEKKYKRQPEETESKPKSHHDGNIDLESDSYLKYDSESSAMALEHPVRA
FGLSE------------------------TSEST-TVVLEPSMVSMETSEPHTLETLKPA
TKTAELSV----ASSS---VISEQSVAVTLEPSVTKIVDSFETAPVPTT----VVLKS-S
EPAVTIPMDYQMKSMLKSLESTPPEPSM-IL-TEPPVAKVIEPPETF-LVSSETPTEVHP
EPSTS-TMDFPEPPATEVQRLPEQPVEVPLEVADSSMARPQEMSELPKTTALEMPESS-V
ASAMELPGPPATSIPELQGPSVTPVLELPGPSAT-----------PVPELPGPLSSPVPE
LLGPPATA--APELPGPSVTSVPQLLQELPGLPATAMGLEPPQEVPEPPVMAQ----ELP
GMPAVTAAVELPGQPAVTVAMELTEQPVTTTELEQPMGMTAVEHPGQPEVTTATGLLGQP
EAAMVLELPGQSVATTALELSGQSSVTGVPELPGLSSATRALELAGQSVAAGALEL-PGQ
LMAAGA-LEYAGQSG-AAGALELL-GQPLATGVLELPG-QPGAPELPGQPVATVALEISV
-QSVVT-TELSTMTVSQSLEVPS-TTALE--SYSTVAQELPATLVGDATV---VDPLMAQ
ESHILASNTMETHMLASNTMDSQMLASNTMDSQMLASNTMDSQMLATSSMDSQMLATSSM
DTQMLATSTMD----------SQMLATSSMDSQMLATSS--MDSQMLATSSMDSQMLATS
SMDSQMLATSSMDSQMLATST--MDSQMLATSTMDSQMLATSSMDSQMLA--SGTMDSQM
LASGTMDAQMLASGTMDAQMLASSSQDSAMLGSKSPDPYRLAQDPYRLAQDPYRLGHEAY
RLGHEAYRLGQDPYRLGHDPYRLTPDPYRMSARPYRIAPRSYRIAPRPYRLAPRPLMLAS
RRSMMMSYAAERSMMSSY--ERSMMSYERSMMSPMAERSMMSAYERSMMSAYERSMMSPM
AERSMMSAYERSMMSAYERSMMSPMTDRSMMSMGADRSMMSSYSAADRYIVMSSYSAADR
SMXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXX-XXX
XXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXX
XXXXXXXXXXXXXXXTVPEQP-MEPESSVISTPVESAVAEEHE--SAERPVTYMVSDTPI
MSAEPTVLTTESSVISET-DNFNSMSASGHI-A-SEASLSLLEPTV-----------AVP
ESSQSTVELPGTAL------------------PELPAVTAQEPPAGTFPEPLAVA-----
--------------------------VSDPPAMNVPEPPTDAVLQPS--AMAEPEHVSIS
MPVVSAL------EPTVPILEPAVSVPQPARIVPE---------SSVSVQESNVTLSEPV
VPVLETSQVMSSETALVSTPMILESSVM------KGMHLLSG-DQNLAPEVVMQEMPSHS
DKE-PHSEGHLKTDLYENEPGINVDLNINNHLIAKETEQNTLSAASTGAAGEIGEEKILP
INETKQCTVLDTCPSVSEADIGGT-LSSAGPV-ALEPDA----VGISKGMEFATTSALSS
ISKYD-EVSLATQDTEHDMVISTSPSGGSEADIEGPLPAKDIHLDL--SNNFVSK-DTEG
PLPMKESDQNLAVALSPKEN-SGEDKEITLPNKEMLSDSGFSATIDDINEADLVRPLLPK
DMERLASLRAG-------IEGPLLPSEVDRDKS-AASPVVIGTQERASE-SSSEEKDDYE
IFVKVKDPHEKSKK-KNREKGEKEKK----------------------------------
-----------------------------------------------RDSSLRSRSKRSK
SSEHKSRK-------RTSESRSRARKRSSKSKS-HRSQTRSRSRSRRRRRSSRSRSKSRG
RRSVSKEKRKRSPKHRSKSRERKRKRSS-SRDNRKSVRARSRTPSRRSRSHTPSRRRRSR
SVGRRRSFSVSPSRRSRT-----PSRRSRTPSRRSRTPSRRSRTPSRRSSTGRHRTF---
---------------CNWINLRFV--VRRTGF-GSAVR-SRRPWI-PVRRRFSRSPIRRK
RSRSSE---RGRSPKRL--TDDKAQLLEIAKANAAAMCAKAGVPLPNLK--A-PP-PTIE
EKA--KSGGAT-IDLAEKCKQIAQSKEDDDVIVNKPHVSDEE-DEEPPFYHHPFKLSEPK
PIFFNLNIAA---AKPTPPKSQVTLTKEFPVSSGSQHRKKEADSVYGEWVPVEKNGEENK
D-DDNVFSSNLPSEPVDISTAMSERALAQKRLSENAFDLEAM-SMLNRAQERIDAWAQLN
SIPGQFTGSTGVQVLTQEQLANTGAQAWIKKDQFLRAAPV
PF01585 // ENSGT00730000111141_0_Domain_1 (2621, 2665)
TGGMGAVLMRKMGWREGEGL
GKNKEGNKEPILVDFKTDRKGLVAENSGT00730000111141_0_Linker_1_2 (2665, 2688)
VGERAQKRSGNFSAAMKDLSGKHPF00035 // ENSGT00730000111141_0_Domain_2 (2688, 2724)
PVSALMEICNKRR
WQPPEFLLVHDSGPDHRKHFLFRVLRNGA-LTPNCMFFLN-RY-----------------
----------------
ENSCJAP00000054117 (view gene)
ENSEEUP00000002160 (view gene)
--------------------------GRNEGQLNGETNTPVEGNQAGDA-AASARNLPNE
EIVQKIEEVLSGVLDTELRYKPDMKEAS-RK-RCVSVQTDPTE-EVPAKKSKKHKKHKNK
KKKKK----KEKEKKYKRQPGESESKPKSHRDGNLHLESASFAKPDSEPSAAALEQPARA
FGLSE------------------------ASEFP-EAVLDPPAVSMEASEPPTSETQMPA
TQTAELSA----GS---TPVISEQSVAGTLEPCKTKVLDSFATVPVPSP---AVVLKSSP
EPVVAMSMEYQVKSVLKSFESTPPDPSK-VT-IEPPVAKVVEPSETL---VSEAPAEVRA
EPSTSTTMNFPESSAPEVLRWPEPPVAVPSETADSSMTCPQELLELPENTASELPESS-V
ASAVELPGPPATSMPESQGPPVTPALELPGPSAT-----------LVPESPGPPSTPVPE
LLGPPATA--VPELPGPSVTSVPQSPQELPGLSAPSEGLEPPQEVPEPPVMAQ----ELP
GLPVVTAAVGVPGQPAATVAMELTEQPVTTTELEQPVGMTTLEHPGQPEVTTATGLLGQP
EAAMVLELPGQSVATTALELSGQPSVTGV---PGLPSATRALELAGQSMAAGALEL-SGQ
LMAAGA-LEYAGQAG-AAGALELL-GQPLATGVLELPG-QPGAPELPGQPVATVALEISV
-QSVVT-TELSTMTVSQSLEVPS-TTALE--SYNTVAQELPTTLVGETSVTVGVDPLMAQ
ESHMLASNTMETHMLASSAMESQMLASTAMDSQMLASNTMDSQMLASSTMDSQMLATSSM
DSQMLATSTMESQMLATSSMESQMLAT----------SS--MDSQMLATSSMDSQMLATS
SMDSQMLATSSMDSQMLATSS--MDSQMLATSSMDSQMLATSSMDSQMLA--SGAMDSQM
LASGTMDAQMLASGTMDAQMLASSTQDSSMLGSKSPDLYRLAQDPYRLAQDPYRLGHDAY
RLGHEAYRLGQDPYRLGHDPYRLTPDPYRMSPRPYRIAPRSYRISPRPYRLAPRPLMLAS
RRSMMMSYAAERSMMSSY--ERSMMSYERSMMSPMAERSMMSAYERSMMSAYERSMMSPM
AERSMMSAYERSMMSTYERSMMSPMADRSMMSMGADRSMMSSYSAADRS-MMSSYSAADR
SMMSSYT-DRSMMSMATDSYTDSYTDTYAEAYMVPPLPPEEPPTMPPLPPEEPPMTPPLP
PEEPPDGSALPAEQSVLTAENTWPSEVPALPSEESVLLTEPTVSQSEISESSAVPVNYSV
TASEPSVLE-SEAAVTVPEPP-QEPESLVTSTPIESAVVGEEHESVPERAATYMMSATPI
VSAEPTMLTPESSVMSETAGDYESMKTSEHV-ASEL-AVSPPESAE-----------TIA
EPPQRAAELPP----------------V--AVSEVAVVAPPEPSAGTXXXXXXXXXXXXX
X-------------------------XXXXXXXXXXXXXX-XXXX----XXXXXXXXXXX
XXXXXXX------X---XXXXXXXXXXX-XXXXXXX---------XXXXXXXXXXXXXXX
XXXXXXXXXXXXXXXXXXXXXXXXXXXXXXSSVIKGTNLLSG-DQNLSPETDQE---VSH
PDEGPHAEGPQKNDCYESEHLVTTDLSVNNHLVADVVVEH-----NSVFASEIGEEKVLP
TDETQQCTVLDTCPIASEVHAEETPPPPAGPL-ALELDA----VGTSQGIEFATTSALSL
VSKYDE---VTTQDNEHDMVISTSPSGGSEADIEGPLPAKDIHLDLPAN-DFLSR-DAEG
PLALKEVDQSLAVTLSPKES-SGEDKDVPVPERESVPESGFPATIDDINEADLVRPLLPK
DMERLASLRAG-------IKGPLLASEVERDKSAXXXXXXXXXXXXXXX-XXXXXXXXXX
XXXXXXXXXXXXXXXXXXXXXXXXXX----------------------------------
-----------------------------------------------XXXXXXXXXXXXX
XXXXXXXX-------XXXXXXXXXXXXX-XXXX-XXXXXXXXXXXXXXXXX-X-XXXXXX
XXXXXXXXXXXXXXXXXXXXXXXXXXXX-XXXXXXXXXXXXXXXXXXXXXXXXXXXXXXX
XXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXX-------------------XXXXXXX
XXXXX--XXXXXXXXXXXXXXXXX--XXXXXXXXXXXX-XXXXXX-XXXXXXXXXXXXXX
XXXXXX---XXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXX-XX-XXXX
XXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXX-XXXXXXXXXXXXXXXXX
XXXXXXXXXX---XXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXX
XXXXXXXXXXXXXXPVDISTAMSERALAQKRLSENAFDLEAM-SMLNRAQERIDAWAQLN
SIPGQFTGSTGVQVLTQEQLANTGAQAWIKKDQFLRAAPV
PF01585 // ENSGT00730000111141_0_Domain_1 (2621, 2665)
TGGMGAVLMRKMGWR-GEGL
GKNKEGNKEPILVDFKTDRKGLVAENSGT00730000111141_0_Linker_1_2 (2665, 2688)
VGERAQKRSGNFSAAMKDLSGKHPF00035 // ENSGT00730000111141_0_Domain_2 (2688, 2724)
PVSALMEICNKRR
WQPPEFLLVHDSGPDHRKHFLFRVLRNGA-LTPNCMFFLN-RY-----------------
----------------
ENSMICP00000030960 (view gene)
ENSMPUP00000001279 (view gene)
MATNIEQIFRSFVVSKFREIQQELSSGRSEGQLNGETNTPNEGNQAGDA-AASARSLPNE
EIVQKIEEVLSGVLDTELRYKPDLKEAS-RKSKCVSVQTDPTD-EIPTKKSKKHKKHKNK
KKKKK----KEKEKKYKRQPEESESKAKSHHDGNIDLESDSFLKFDSEPSAVALEHPVRA
FGLSE------------------------TSESP-AVVLEPPVVSMEVSEAHTLETLKPA
TKTAELSV----ASTSTISVQSEQSVAVTLESSMTKILDSFTTAPVPTT---TVMLKS-P
EPAVTMSMEYQMKPMLKSLETTPPEQSKV-M-LEPPVAKGLEPSETL-VLLSEIPTEVHP
EPSTSTTMDFPESSATEGVRLPEQPVEVPSEIADSSMTRPQELPELPKTTALEQPESS-V
ASVMELPGPPATSKPELQGPPVTPVLELPGPSAT-----------PLPELPGPLSTPVPE
LLGPPATA--VPELPGPSATSVPQLSQELPGLPAPSVGLEPPQEVPEPPVMAQ----ELP
GLPAVTAAVELPGQPAVTVAMELTEQPVTTSELEQPVGMTAVEHPGQPEVTTATGLLGQP
EAAMVLELPGQPVATTALELPGQPSVTGVPELPGLPSATRALELSGQPVATGALEL-PGQ
LMATGA-LEFSGQSG-AAGALELL-GQPLATGVLELPG-QPGAPELPGQPVATVALEISV
-QSVVT-TELSTMTVSQSLEVPS-TTALE--SYNTVAQELPTTLVGETSVTVGVDPLMAQ
ESHMLASNTMETHMLASNTMDSQMLASNTMDSQMLASNTMDSQMLATSSMDSQMLATSSM
DSQMLATSSMDSQMLATSSMDSQMLATSSMDSQMLATSS--MDSQMLATSS---------
-MDSQMLATSSMDSQMLATST--MDSQMLATSTMDSQMLATSSMDSQMLA--SGTMDSQM
LASGTMDAQMLASGSMDAQMLASSTQDSAMLGSKSPDPYRLAQDPYRLAQDPYRLGHDPY
RLGHDAYRLGQDPYRLGHDPYRLTPDPYRMSPRPYRIAPRSYRIAPRPYRLAPRPLMLAS
RRSMMMSYAAERSMMSSY--ERSMMSYERSMMSPMAERSMMSAYERSMMSAYERSMMSPM
AERSMMSAYERSMMSAYERSMMSPMADRSMMSMSADRSMMSSYSAADRS-MMSSYSAADR
SMMSSYTADRSMMSMAADSYTDSYTDTYTEAYMVPPLPPEEPPTMPPLPPEEPPMTPPLP
PEEPPEGPALPTEQSALTAENTWPTEVPALPPEESVSLSEPSVSQSEISEPSALPANYSV
SASDPSALA-SEAAVTVPDPA-LEPESSVTSTPVESAVVAEEHQVVPERAVTYMVSETTI
MSAEPTVLASESSVMSETAETFDSMKASGHA-S--EVSLSLLDPAA-----------ANP
EASQSTL----------------ELPAM--AVSELPAVAVPEPPTGTVPEPPAAAAVLEP
PAVAIP--------------DPTAVAVSDPIAVAVPDTP--AEGVPETLALAESEHVTTP
VPVVSAL------EPTVPVLEPAVSVPQSNTVV-SE--------PSVPVQEPTVIISEPA
VTVSEQTQVIPTEMVLESTPMILESS------VVKGVNLLSS-DQNTAPEIGMQEIPMHS
DEE-PHAEGHLKNDPYESEHGTDTDLKTDNHFVAQEMEHNTASAASTGAVGEIGEGNILS
IGETKQCTVLDTCSSVSEAD-GGT-LSSAGPL-ALEPDA----VGTSKGIEFAIASALGS
VSNYDAEVSLTAQDTEHDMIISTSPSGGSEADIEGPLPAKDIHLDLPSN-NFINK-DAEG
PLSIKDCDQTLAVALSPKES-SGEDKEIPLPTKEIQSDSGFSATIDDINEADLVRPLLPK
DMERLTSLRAG-------IEGPLLPSEVERDKS-AASPVVISIPERASE-SSSEEKDDYE
IFVKVKDTHEKSKKNKNRDKGEKEKK----------------------------------
-----------------------------------------------RDSSLRSRSKRSK
SSEHKSRK-------RTSESRSRARKRSSKSKS-HRSQTRSRSRSRRRRRSSRSRSKSRG
RRSVSKEKRKRSPKHRSKSRERKRKRSS-SRDNRKTVRARSRTPSRRSRSHTPSRRRRSR
SVGR-RSFSISPSRRSRT-----PSRRSRTPSRRSRTPSRRS--------------RTPS
RRSRT--PSRRSRTPSRRRRSRSV--VRRRSFSISPVR-LRRSRT-PLRRRFSRSPIRRK
RSRSSE---RGRS-PKRLTDLNKAQLLEIAKANAAAMCAKAGVPLPPNLKPA-PP-PTIE
EKVAKKSGGATIEELTEKCKQIAQSKEDDDVIVNKPHVSDEE-EEEPPFYHHPFKLSEPK
PIFFNLNIAA---AKPTPPKSQVTLTKEFPVSSGSQHRKKEADSVYGEWVPVEKNGEENK
D-DDNVFSSNLPSEPVDISTAMSERALAQKRLSENAFDLEAM-SMLNRAQERIDAWAQLN
SIPGQFTGSTGVQVLTQEQLANTGAQAWIKKDQFLRAAPV
PF01585 // ENSGT00730000111141_0_Domain_1 (2621, 2664)
TGGMGAVLMRKMGWREGEGL
GKNKEGNKEPILVDFKTDRKGLVENSGT00730000111141_0_Linker_1_2 (2664, 2688)
AVGERAQKRSGNFSAAMKDLSGKHPF00035 // ENSGT00730000111141_0_Domain_2 (2688, 2755)
PVSALMEICNKRR
WQPPEFLLVHDSGPDHRKHFLFRVLINGSAYQPSFASPNK-KHAKATAATVVLQAMGLVP
KDLMANATCFRSASRR
ENSPCOP00000008444 (view gene)
ENSMNEP00000015129 (view gene)
ENSBTAP00000011175 (view gene)
MATNIEQIFRSFVVSKFREIQQELSSGRSEGQLNGETNTPIEGNQAGDA-AASARNLPNE
DIVQKIEEVLSGVLDTELRYKPDLKEAS-RKSRCVSVQTDPTD-EIPTKKSKKHKKHKNK
KKKKK----KEKEKKYKRQSEESESKPKSHHDGNI--ESDSFLKFDSEPSEMALEHSVRA
FGLYD------------------------TSESP-AVVLEPPVVSMEVSEPHILETLTPA
TKHTELSV----ASTSVVSMPSEQSVAVMLEPSTTKILDSFATAPVPTT---TVVLKS-S
EPVVTVSVECQMKSVQKSLESTPPEPSKIML-LEPPVAKVLELSETL--VSSETPTEVHP
EPSTSTAMDFPESSATEVLRLPEQPVEGPSEIADSSMTRPQEMLELPKTTALELQESS-V
ASVMELPGPPATSMPELQGPPVTPVLELPGPSAT-----------PAPELPGPLSTPVPE
LLGPPATA--VPELAGPSVTSVPQLSQELSGLPAPSMGLEPPQEVPEPPVMAQ----ELP
GLPAVTTAVELPGQPVVTVAMELTEQPVTTTELEQPVGMTAVEHPGQPEVTTATGLLGQP
EAAMVLELPGQPVATTALELSGQPSVTGVPELPGLPSATRALELSGQPVATGALEL-PGQ
LMAAGA-LEFSGQSG-AAGALELL-GQPLATGVLELPG-QPGAPELPGQPVATVALEISV
-QSVVT-TELSTMTVSQSLEVPS-TTALE--SYNT-------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
----------ERSMMSSY--ERSMMSYERSMMSPMAERSMMSAYERSMMSAYERSMMSPM
AERSMMSAYERSMMSAYERSMMSPMADRSMMSMGADRSMMSSYSAADRS-MMSSYSAADR
SMMSSYTADRSMMSMAADSYTDSYTDTYTEAYMVPPLPPEEPPTMPPLPPEEPPMTPPLP
PEEPPEGPVLAAEQSALTAENTWSAEVPALAPEESVSLPEPPVSQSEIAEPLAVTANYSV
SASEPSVLA-SEADVTVPEPQ-LEPESSVMSTPVESAAGAEEHEMVAERPVTYMVSETPT
TSAEPTVLTSEPSVISETAETYDSMRASGQT-A-SEVSMSLLEPAV-----------PIA
EPSQSTVD----------------LPAM--AVPEHPAGIVSETSTVAVPEPQAEA-----
----------VLDPPAAAVQDPPAVAVLDPP----------AEAVPEPLALSEPEHVTVA
VPVVSTL------EPTVPILEPVVSILQPNVIVSE---------PSSCVQESTVTISEPP
VTVSEQTQVIPAETVVESTAVILESS------VIKGMNLLSG-DQSIASEIGLQEIAMHS
EEE-PRAEGLLKSGSNETENCISTDLNINNHLIAQEMECSTVSAASTGAIGEMGEEAVLP
TSETKQCTVLDTCPSVSETELGGT-LSSVGPL-VLESEA----VGTGKDLEFGTASALSS
VSKYDGEVSLTTQDTEHDMVISTSPSGGSEADIEGPLPAQDIHPDLSN--NFINK-DAEG
SLPVQESDQMLAAAISPKES-SGEDKEVSLTTKEILSDSGFPASIDDINEADLVRPLLPK
DMERLTNLRAG-------IEGPLLPSEVERDKS-AASPVVISIPEKASE-SSSEEKDDYE
IFVKVKDTHEKSKKNKNRDKGEKEKK----------------------------------
-----------------------------------------------RDSSLRSRSKRSK
SSEHKSRK-------RTSESRSRARKRSSKSKS-HRSQTRSRSRSRRRRRSSRSRSKSRG
RRSVSKEKRKRSPKHRSKSRERKRKRSS-SRDNRKTGRARSRTPSRRSRSHTPSRRRRSR
SGGR-RSFSISPSRRSRT-----PSRRSRTPSRRSRTPSRRS--------------RTPS
RRSRT--PSRRSRTPSRRRRSRSV--VRRRSFSISPVR-LRRSRT-PLRRRFSRSPIRRK
RSRSSE---RGRS-PKRLTDLNKAQLLEIAKANAAAMCAKAGVPLPPNLKPA-PP-PTIE
EKVAKKSGGATIEELTEKCKQIAQSKEDDDVIVNKPHVSDEE-EEEPPFYHHPFKLSEPK
PIFFNLNIAA---AKPTPPKSQVTLTKEFPVSSGSQHRKKEADSVYGEWVPVEKNGEENK
D-DDNVFSSNLPSEPVDISTAMSERALAQKRLSENAFDLEAM-SMLNRAQERIDAWAQLN
SIPGQFTGSTGVQVLTQEQLANTGAQAWIKKDQFLRAAPV
PF01585 // ENSGT00730000111141_0_Domain_1 (2621, 2664)
TGGMGAVLMRKMGWKEGEGL
GKNKEGNKEPILVDFKTDRKGLVENSGT00730000111141_0_Linker_1_2 (2664, 2688)
AVGERAQKRSGNFSAAMKDLSGKHPF00035 // ENSGT00730000111141_0_Domain_2 (2688, 2755)
PVSALMEICNKRR
WQPPEFLLVHDSGPDHRKHFLFRVLINGSAYQPSFASPNK-KHAKATAATVVLQAMGLVP
KDLMANATCFRSASRR
ENSOPRP00000004675 (view gene)
MAADIEQIFRSFVVSKFREIQQELSXXXXXXXXXXXXXXXXXXXQAGDAAAASARNLPNE
EIVQKIEEVLSGVLDTELRSKPEMKETS-RKSRCVSVQTDPTE-EVPAKKSKKHKKHKNK
KKKKK----KEKEKKYKRQPEQSQSKPKSHNDGNVDLESDSFLKFDAESSAVALE-PVRA
FGRSE------------------------SSESP-AVLLEPPLVAMEVSEPHILETLKPA
TKTAELSV----ASTSVISEPSEQPMAVMLEPSTSKSLDSCATALLPTS---TVALKS-S
EPVAAMSLEYHKQCLLKSLESASTESSKIILAXXXXXXXXXXXXXX--------------
-------------------------------------------XXXXXXXXXXXXXXX-X
XXALELPGPPATSMPELQRPPLQ--LEIPGPS------------TPVPELPGPLSTPVSE
MSGPPVTS--VPELPGPSVTPLPQMSQEFSGLSAPSVCFETPQEVSEPPAVAQ----ELP
GQPAVSSAVELPGQPAVTVAMELTEQPVTTTELEHPVGMTAVEHPGHPEVTTAAGLLGQP
EAAMVLELPGQPVATTALELSGQASVTGVPELPGLPSATRALELSGQPVATGALEL-PGQ
LMATGA-LEYTGQPG-AAGALELL-GQPLATGVLELPG-QPGAPELPGQPVATVALEISV
-QSVVTTSELSTMTVSQSLEVPS-TTALE--SYNTVAQELPTTLVGETSVTVGVDPLMAQ
DSHMLASNTMETHMLASNTMESQMLASNTMDSQMLASNTMDSQMLATSSMDSQMLATSTM
DSQMLATSTMDSQMLATSSMDSQMLATSSMDSQML------------ATSSMDSQMLATS
SMDSQMLATSSMDSQMLATST--MDSQMLATSTMDSQMLATSSMDSQMLA--SGTMDSQM
LASGTMDAQMLASGTMDAQMLASSTQDSAMLDSKSPDPYRLAQDPYRLAQDPYRLGHDPY
RLGHEAYRLGQDPYRLGHDPYRLTPDPYRMSPRPYRIAPRSYRIAPRPYRLAPRPLMLAS
RRSMMMSYAAERSMMSSY--ERSMMSYERSMMSPMAERSMMSAYERSMMSAYERSMMSPM
AERSMMSAYERSMMSAYERSMMSPMADRSMMSMSADRSMMSSYSAADRS-MMSSYSAADR
SMMSSYT-DRSMMSMAADSYADSYTDTYTEAYMVPPLPPEEPPTMPPLPPEEPPMTPPLP
PEEPPEGTALPAEDSALAAESTWPTEVPALPPEEAVSQPEPPVSQSEVAESSAVSAHYSV
SASESSVLA-SEAVMTVPEPP-LEPESSIVSTPIEPAVVA-EHEVVPERPVTETVM---M
-SVE--QAVSEPPVISEAVEHSDSMRALE--HVASESSVSLLAPAV-----------TAP
EP-ENIMELPALAVSEAAPVAVQEPPATVVAVPELPAKSVLEPPPGAE------------
-----------------------------------SESMAV--TVIAPPAVTEPEHVTIP
VS-ASVL------DPTVPVMESAVSVLQPAMIVSQ---------PSVSVQETTVTISESS
VTISEQTQIISTEMVLESTPVILESSGMSAPLAVKGMHLLSG-DQSLVPEIGMQKMMTLH
SSEQPHAEGHLKSDAYANECSVNSDLNTSSHLIMKEMEHGTVSAVDT---VHASEEKILP
ISETEQCTVLG--AGVTEADVGRT-LSNTGPF-AMEPDAV---G-TSKGVDYVTPCAVSS
LSKYDVDAPLAVQDTEHDMIISTSPSGGSEADIEGPLPAKDIHLDLSSS-NIV-K-DREE
QLPGKESEQALAVAGSPKQS-SGEEKEIPLPTKDT-ADSAFPATIEDIXXXXXXXXXXXX
XXXXXXXXXXX-------XXXXXXXXXXXXXXX-XXXXXXXSTAERASE-SSSEELDDYE
IFVKVKDTHEKSKKNKNRDKGEKEKK----------------------------------
-----------------------------------------------RDSSLRSRSKRSK
SSEHKSRK-------RTSESRSRARKRSSKSKS-HRSQTRSRSRSRRRRRSSRSRSKSRG
RRSVSKEKRKRSPKHRSKSRERKRKRSS-SRDNRKTVRARSRTPSRRSRSHTPSRRRRSR
SVGRRRSFSVSPNRRSRT-----PSRRS--------------RTPSRRSRTPSRRSRTPS
RRSRT--PSRRSRTPSRRRRSRSV--VRRRSFSISPVR-LRRSRT-PLRRRFSRSPIRRK
RSRSSE---RGRS-PKRLTDLDKAQLLEIAKANAAAMCAKAGVPLPPNLKPA-PP-PTIE
EKVAKKSGGATIEELTEKCKQIAQSKEDDDVIVNKPHVSDEE-EEEPPFYHHPFKLSEPK
PIFFNLNIAA---AKPTPPKSQVTLTKEFPVSSGSQHRKKEADSVYGEWVPVEKNGEENK
D-DDNVFSSNLPSEPVDISTAMSERALAQKRLSENAFDLEAM-SMLNRAQERXDAWAQLN
SIPGQFTGSTGVQI-TQEQLANTGAQAWIKKDQFLRAAPV
PF01585 // ENSGT00730000111141_0_Domain_1 (2621, 2665)
TGGMGAVLMRKMGWREGEGL
GKNKEGNKEPILVDFKTDRKGLVAENSGT00730000111141_0_Linker_1_2 (2665, 2688)
VGERAQKRSGNFSAAMKDLSGKHPF00035 // ENSGT00730000111141_0_Domain_2 (2688, 2724)
PVSALMEICNKRR
WQPPEFLLVHDSGPDHRKHFLFRVLRNGA-LTPNCMFFLN-RY-----------------
----------------
ENSPTRP00000086465 (view gene)
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
-----------------------------------------MPGLTV
PF01585 // ENSGT00730000111141_0_Domain_1 (2628, 2664)
LMRKMGWREGEGL
GKNKEGNKEPILVDFKTDRKGLVENSGT00730000111141_0_Linker_1_2 (2664, 2687)
AVGERARKRSGNFSAAMKDLPGKPF14709 // ENSGT00730000111141_0_Domain_2 (2687, 2732)
HPVSALMEICNKRR
WQPPEFLLVHDSGPDHRKRFLFRVLRSGSPYQPSCMFFLN-RD-----------------
----------------
ENSHGLP00100020224 (view gene)
------------------------------------------------------------
-----------------------LKEAS-RKSRCVSVQTDPTD-EIPSKKSKKHKKHKNK
KKKKK----KEKEKKFKRQPEESESKPKSHHDGNIDLESDSFLKFDSEPSAVALEHPIRA
FGLSE------------------------TSESS-TVVLEPSISSMEVSEPHILETLKPA
TKTAELSI----GSTSVISE---QSMAATLEPSVTKIVESFATAPVPTT----VVLKS-S
EPVATMSVEYQMKSVLKSLEHIAPEPSKIM-LAEPLVAKELEPSETL-VIVSETSSEVHP
EPSTSTTVDFPESSATEVLRLPEHPV--PSEIADPSVTRPQELLELSKTSAVELPESS-V
ASVLELPGPPATSMPELQGPPVTPVLELPGPSA-----------TPVSELSGPLSTPVPE
LPGPPATA--VPELAGPSVTPVPQLSQELPGPPAPSMGLEPPQEVPEPPVMVH----ELP
GLPAVSATVELPGQPAVTVAMELTEQPVTTTEFEQPVGMTAVEHPGHPEVTATTGLLGQP
EATMVLELPGQPVAATALELSGPPSVAGVPELPGLPSATRALELSGQSVAAGALEL-PGQ
LMATGA-LEFSGQSG-AAGALELL-GQPLATGVLELPG-QSGAPELPGQPVATVALEISV
-QSVVTTSELSTMTMSQSLEVPS-TTALE--SYNTVAQELPTTLVGETSVTVGVDPLMAQ
ESHMLASNTMETHMLASNTMDSQMLASNTMDSQMLASNTMDSQMLASSTMDSQMLATSSM
DSQMLATSSMDSQMLATSSMDSQMSTSTSMDSQMLAS----------ATSSMDSQMLATS
SMDSQMLATSSMDSQMLATSS--MDSQMLATSSMDSQMLATSSMDSQMLA--SGTMDSQM
LASGTMDAQMLASGTMDAQMLASSTQDSAMLDSKSPDPYRLAQDPYRLAQDAYRLGHDPY
RLGHDAYRLGQDPYRLGHDPYRLTPDPYRMSPRPYRIAPRSYRIAPRPYRLAPRPLMLAS
RRSMMMSYAAERSMMSSY--ERSMMSYERSMMSPMAERSMMSAYERSMMSAYERSMMSPM
AERSMMSAYERSMMSAYERSMMSPMADRSMMSMGADRSMMSSYSAADRS-MMSSYSAADR
SMMSSYTADRSMMSMAADSYTDSYTDTYTEAYMVPPLPPEEPPTMPPLPPEEPPMTPPLP
PEEPPEGPALTMEQSALTAENTWPAEMPALPPEESVSQPEPPVSQSEISEPLAVPTNYSV
SESESSVLV-SEAVVTITEPP-QEPESSVISTPVESAVVA-EYEIVPERPVTCIVSETPI
-SPEPAVLTSEPPVMTETIETLDSTQFEG--HVSSETSMSLIDPTV-----------TFP
EPAENIQELPAMAVPEPPAVAIPESPAV----------GVPESPAMAVPESPAVADPES-
-------PA-------LAIPELPAMVVPELPAMPKPEPPAV--AVFELPSVAEPEHVTVP
VSAVSAL------DPAVSVLEPAVSVLQPTVIVSE---------PCISVQQYSVTVLEPA
VTVSEQTQIISTEMVLESIPVIAESSVSS-SHVMKGMNLLSG-DQNLAPEIGTQET-LLY
SGEETHAEGHLKSDSYESEHGLNVDFNINNHLIAKAMESNM-STASTDSVSGIGEEKILP
INETNKCTVLDTCLGVSQADGGGS-LSSASPL-ALEPDTP---G-TNKGVEFVMATAITT
INKYD-DVSLTTQDTEHDMVISTSPGGVGEADIEGPLPAKDIHFDLPSN-NLVSK-DIEE
PLFVKESDQVLPIALSPKES-G--EDEVPLPSKEIVSDSGFCANIDEINEADLVRPLLPK
DMERLASLRAG-------IEGPLLPSEAERDKS-AASPVVISIPERASE-SSSEEKDDYE
IFVKVKDTHEKSKKNKNRDKGEKEKK----------------------------------
-----------------------------------------------RDSSLRSRSKRSK
SSEHKSRK-------RTSESRSRARKRSSKSKS-HRSQTRSRSRSRRRRRSSRSRSKSRG
RRSVSKEKRKRSPKHRSKSRERKRKRSS-SRDNRKTGRARSRTPSRRSRSHTPSRRRRSR
SVGRRRSFSISPSRXXXX-----XXXXXXXXX----XXXXXXXXXXXRSRTPSRRSRT--
-------PSRRSRTPSRRRRSRSV--VRRRSFSISPVR-LRRSRT-PLRRRFSRSPIRRK
RSRSSE---R-----ERLTDLDKAQLLEIAKANAAAMCAKAGVPLPPNLKPA-PP-PTIE
EKVAKKSGGATIEELTEKCKQIAQSKEDDDVIVNKPHVSDEE-EEEPPFYHHPFKLSEPK
PIFFNLNIAA---AKPTPPKSQVTLTKEFPVSSGSQHRKKEADSVYGEWVPVEKNGEENK
D-DDNVFSSNLPSEPVDISTAMSERALAQKRLSENAFDLEAM-SMLNRAQERIDAWAQLN
SIPGQFTGSTGVQVLTQEQLANTGAQAWIKKDQFLRAAPV
PF01585 // ENSGT00730000111141_0_Domain_1 (2621, 2664)
TGGMGAVLMRKMGWREGEGL
GKNKEGNKEPILVDFKTDRKGLVENSGT00730000111141_0_Linker_1_2 (2664, 2687)
AVGERAQKRSGNFSAAMKDLSGKPF14709 // ENSGT00730000111141_0_Domain_2 (2687, 2756)
HPVSALMEICNKRR
WQPPEFLLVHDSGPDHRKHFLFRVLINGSAYQPSFASPNK-KHAKATAATVVLQAMGLVP
KDLMANATCFRSASRR
ENSCAFP00000040685 (view gene)
MATNIEQIFRSFVVSKFREIQQELSSGRSEGQLNGETNTPNEGNQAGDA-AASARSLPNE
EIVQKIEEVLSGVLDTELRYKPDLKEAS-RKSRCVSVQTDPTD-EIPTKKSKKHKKHKNK
KKKKK----KEKEKKYKRQPEESESKVKSHHDGNIDLESDSFLKFDSEPSAMALEHPVRA
FGLSE------------------------TSESP-AVVLEPPVVSMEISESHTLETLKPA
TKTAELSV----ASTSVISVQSEQSVALTLEPSMTKILDSFTTAPVPTT---TVVLKS-P
EPVVTMSMEYQMKPVLKSLETIPPEQSQII--LEPPVAKGLESSETL-VVSPEIPTEVHP
EPSTSTTMDFPESSATEGLPLPEQPIEVPSEIADSSMTRPQELLELPKTTALELPESS-V
ASVMELPGPPATSKPELQGPPVTPVLELPGPSAT-----------PLPELPGPLSTPVPE
LLGPPATA--VPELPGPSATSVPQLSQELPGLPAPSVGLEPPQEVPEPPVMAQ----ELP
GLPAVTAAVELPGQPAVTVAMELTEQPVTTSELEQPVGMTTVEHPGPPEVTTATGLLGQP
EAAMVLELPGQPVATTALELPGQPSVTGVPELPGLPSATRALELSGQPVATGALEL-PGQ
LMATGA-LEFSGQSG-AAGALELL-GQPLATGVLELPG-QPGAPELPGQPVATVALEISV
-QSVVT-TELSTMTVSQSLEVPS-TTALE--SYNTVAQELPTTLVGETSVTVGVDPLMAQ
ESHMLASNTMETHMLASNTMDSQMLASNTMDSQMLASNTMDSQMLASSTMDSQMLATSSM
DSQMLATSSMDSQMLATSSMDSQMLATSSMDSQMLATSS--MDSQMLATSSMDSQM----
------LATSSMDSQMLATST--MDSQMLATSTMDSQMLATSSMDSQMLA--SGTMDSQM
LASGTMDAQMLASGTMDAQMLASSTQDSAMLGSKSPDPYRLAQDPYRLAQDPYRLGHDPY
RLGHDAYRLGQDPYRLGHDPYRLTPDPYRMSPRPYRIAPRSYRIAPRPYRLAPRPLMLAS
RRSMMMSYAAERSMMSSY--ERSMMSYERSMMSPMAERSMMSAYERSMMSAYERSMMSPM
AERSMMSAYERSMMSAYERSMMSPMADRSMMSMGADRSMMSSYSAADRS-MMSSYSAADR
SMMSSYTADRSMMSMAADSYTDSYTDTYTEAYMVPPLPPEEPPTMPPLPPEEPPMTPPLP
PEEPPEGPALPTEQSALTAENTWSTEVPALPPEESVSLSEPSVSQSEMSEPSALPANYSV
SASDPSVLA-SEAAVTVPEPP-LEPESSVTATPVESTVVAEEHQIVPERAVTYMVSETPI
MTAEPTVLTSEPSVMSETAETFDSMKASGHV-A-SEVSLSLLEPAA-----------TNP
EPSQNTL---------------E-LPAM--TISELPAVAVPQPPTGTVPEPPTVA-VLET
PAVAIPDPTAVA------VSDPVAVAVPDPPVEAVPETL----------ALAESEHVTIP
VPV-SAL------EPTVPVLEPVVSVPQ-PNTVVSE-------P-SVSVQESTVIISEPA
VTVSEQTQVIPDEMVLESTPMILEPT------VIKGVSILPG-DQNLAPEIGIQEIPMHA
DEE-PHAEGHLKNDPYESENGTNIELNVNNHLVAKEMEHNTVSAASTGAVDEIGEGNILS
ISETKQCTVLDTCSSVSEADGGT--LSSAGPL-ALEPDA----MGTSKGIEFATESALSS
VNKYDVEVSLTTQDTEHDMIISTSPSGGSEADIEGPLPAKDIHLDLPSN-NFMSK-DAEG
PLPIKECDQTLAVALSPKES-SGEDKEVALPTKEILSDSGFSANIDDINEADLVRPLLPK
DMERLTSLRAG-------IEGPLLASEVERDKS-AASPVVISIPERASE-SSSEEKDDYE
IFVKVKDTHEKSKKNKNRDKGEKEKK----------------------------------
-----------------------------------------------RDSSLRSRSKRSK
SSEHKSRK-------RTSESRSRARKRSSKSKS-HRSQTRSRSRSRRRRRSSRSRSKSRG
RRSVSKEKRKRSPKHRSKSRERKRKRSS-SRDNRKTVRARSRTPSRRSRSHTPSRRRRSR
SVGR-RSFSISPSRRSRT-----PSRRSRTPSRRSR--------------TPSRRSRTPS
RRSRT--PSRRSRTPSRRRRSRSV--VRRRSFSISPVR-LRRSRT-PLRRRFSRSPIRRK
RSRSSE---RGRS-PKRLTDLNKAQLLEIAKANAAAMCAKAGVPLPPNLKPA-PP-PTIE
EKVAKKSGGATIEELTEKCKQIAQSKEDDDVIVNKPHVSDEE-EEEPPFYHHPFKLSEPK
PIFFNLNIAA---AKPTPPKSQVTLTKEFPVSSGSQHRKKEADSVYGEWVPVEKNGEENK
D-DDNVFSSNLPSEPVDISTAMSERALAQKRLSENAFDLEAM-SMLNRAQERIDAWAQLN
SIPGQFTGSTGVQVLTQEQLANTGAQAWIKKDQFLRAAPV
PF01585 // ENSGT00730000111141_0_Domain_1 (2621, 2664)
TGGMGAVLMRKMGWREGEGL
GKNKEGNKEPILVDFKTDRKGLVENSGT00730000111141_0_Linker_1_2 (2664, 2686)
AVGERAQKRSGNFSAAMKDLSGPF14709 // ENSGT00730000111141_0_Domain_2 (2686, 2732)
KHPVSALMEICNKRR
WQPPEFLLVHDSGPDHRKHFLFRVLRNGSPYQPNCMFFLN-RY-----------------
----------------
ENSOARP00000013974 (view gene)
MATNIEQIFRSFVVSKFREIQQELSSGRSEGQLNGETNTPIEGNQAGDA-AASARSLPNE
DIVQKIEEVLSGVLDTELRYKPDLKEAS-RKSRCVSVQTDPTD-EIPTKKSKKHKKHKNK
KKKKK----KEKEKKYKRQSEESESQPKSHHDGNI--ESDSFLKFDSEPSEMALEHSVRA
FGLYD------------------------TSESP-AVVLEPPVVSMEVSEPHILETLPPA
TKHTELSV----ASTSVVSVQSEQSVAVMLEPSTTKILDSFATAPVPTT---TVVLKS-S
EPVVTVSVECQMKSVQKSLESTPPEPSKIML-LEPPVAKVLELSETL--VSSETPTEVHP
EPSTSTTMDFPESSATEVLRLPEQPVEGPSEIADSSMTRPQEMLELPKTTALELQESS-V
ASVMELPGPPATSMPELQGPPVTPVLELPGPSAT-----------PAPELPGPLSTPVPE
LLGPPATA--VPELAGPSVTSVPQLSQELSGLPAPSMGLEPPQEVPEPPVMAQ----ELP
GLPAVTTAVELPGQPVVTVAMELTEQPVTTTELEQPVGMTAVEHPGQPEVTTATGLLGQP
EAAMVLELPGQPVATTALELSGQPSVTGVPELPGLPSATRALELSGQPVATGALEL-PGQ
LMAAGA-LEFSGQSG-AAGALELL-GQPLATGVLELPG-QPGAPELPGQPVATVALEISV
-QSVVT-TELSTMTVSQSLEVPS-TTALE--SYNTVAQELPTTLVGETSVTVGVDPLMAQ
ESHMLASNTMETHMLASNTMDSQMLASNTMDSQMLASNTMDSQMLASSTMDSQMLATSSM
DSQMLATSSMDSQMLATSSMDSQ-------------------------------------
---------------MLATSS--MDSQMLATSTMDSQMLATSSMDSQMLA--SGTMDSQM
LASGSMDAQMLASGTMDAQMLASSTQDSAMLGSKSPDPYRLAQDPYRLAQDPYRLGHDPY
RLGHDAYRLGQDPYRLGHDPYRLTPDPYRMSPRPYRIAPRSYRIAPRPYRLAPRPLMLAS
RRSMMMSYAAERSMMSSY--ERSMMSYERSMMSPMAERSMMSAYERSMMSAYERSMMSPM
AERSMMSAYERSMMSAYERSMMSPMADRSMMSMGADRSMMSSYSAADRS-MMSSYSAADR
SMMSSYTADRSMMSMAADSYTDSYTDTYTEAYMVPPLPPEEPPTMPPLPPEEPPMTPPLP
PEEPPEGPVLATEQSALTAENTWSAEVPALPPEESVSLPEPPVSQSEIAEPLAVTANYSV
SASEPSVLA-SEADVTVPEPP-LEPESSVMSAPVESAAGAEEHEIVAERPVTYMVSETPT
TSAEPTVLTSEPSVMSDIAETYDSMRASGQT-A-PEVSVCLLEAAV-----------PVG
EPSQSTV----------------DLPAM--AVTELPSVAVPEHPAGNVSETSTMA-----
----VPEPQA------EALQDPPAVAVLDRL----------AEAVPEPLALSEPEHVTVA
V--------ASTLEPTVPILEPAVSILQPNVIV-SE--------PSGCVQESTVTISEPP
VTVSEQTQVIPTETVVESTPVILESS------TIKGMNLLSG-DQNIASEIGLQDIAMHS
DEE-PRAEGLLKSGSNETENCINTDLNINNHLIAQEVERSTVSAASTGAVGEIGEETILP
TGETKQCTVLDTCPSVGETELGGT-LSSVGPL-VLESEA----VGTGKDLEFGTTSALSS
VSKYDGEVSLTTQDTEHDMVISTSPSGGSEADIEGPLPAKDIHPDLSN--NFINK-DAEG
SLPVQESDQTLAAAVSPKES-SGEDKEVCLTTKEILSDPGFPASIDDINEADLVRPLLPK
DMERLTNLRAG-------IEGPLLSSEVERDKS-AASPVVISIPERASE-SSSEEKDDYE
IFVKVKDTHEKGKKNKNRDKGEKEKK----------------------------------
-----------------------------------------------RDSSLRSRSKRSK
SSEHKSRK-------RTSESRSRARKRSSKSKS-HRSQTRSRSRSRRRRRSSRSRSKSRG
RRSVSKEKRKRSPKHRSKSRERKRKRSS-SRDNRKTGRARSRTPSRRSRSHTPSRRRRSR
SGGR-RSFSRPPSRGSRT-----PSRRSRTPSRR---------------------SRTPS
RRSRT--PSRRSRTPSRRRRSRSV--VRRRSFSISPVR-LRRSRT-PLRRRFSRSPIRRK
RSRSSE---RGRS-PKRLTDLNKAQLLEIAKANAAAMCAKAGVPLPPNLKPA-PP-PTIE
EKVAKKSGGATIEELTEKCKQIAQSKEDDDVIVNKPHVSDEE-EEEPPFYHHPFKLSEPK
PIFFNLNIAA---AKPTPPKSQVTLTKEFPVSSGSQHRKKEADSVYGEWVPVEKNGEENK
D-DDNVFSSNLPSEPVDISTAMSERALAQKRLSENAFDLEAM-SMLNRAQERIDAWAQLN
SIPGQFTGSTGVQVLTQEQLANTGAQAWIKKDQFLRAAPV
PF01585 // ENSGT00730000111141_0_Domain_1 (2621, 2664)
TGGMGAVLMRKMGWKEGEGL
GKNKEGNKEPILVDFKTDRKGLVENSGT00730000111141_0_Linker_1_2 (2664, 2688)
AVGERAQKRSGNFSAAMKDLSGKHPF00035 // ENSGT00730000111141_0_Domain_2 (2688, 2755)
PVSALMEICNKRR
WQPPEFLLVHDSGPDHRKHFLFRVLINGSAYQPSFASPNK-KHAKATAATVVLQAMGLVP
KDLMANATCFRSASRR
ENSSTOP00000016835 (view gene)
ENSCAPP00000017326 (view gene)
-----------------------------------------------D-AASSARSLPNE
EIVQKIEEVLSGVLDTELRYKPDLKEAS-RKSRCVSVQTDPTD-EIPSKKSKKHKKHKNK
KKKKK----KEKEKKYKRQPEESESKPKSHHDGNIDLESDSFLKFDSEPSAMALEHPXXX
XXXXX------------------------XXXXX-XXXXXXXXXXXXXXXXX----XXXX
XXXXXXXX----XXXXXXXXXXXXXXXVMLEPSVTKIVDSFATTLVPTTVPATVVVKS-P
EPVATMSVEYQMKSALKSSEHTSLEPSHIM-LAEPPVAKELEPSETH-VIALETSTEVYP
EPSISTTKDFPESSATEVLRLPEHSI--PSEIADPAMTRPQDLLELSKTSALELPVSS-V
ASALELPGPPATSMPELQGPPVTPMLELPGPSA-----------TPVSELSGPLSTPVPE
LPGPPATA--VPELPGPSVTPVPQLSQELPGPLAPSVGLEPPQEVPEPPVMAQ----ELP
GLPAVSATVELPGQPAITVAMELTEQPVTTTEFEQPVGMTAVEHPGHSEVTAATGLLGQP
EAAMVLELPGQPVATTALELSGQPSVTGVPELPGLSSATRALELSGQSV-AGALEL-PGQ
LMAAGA-LEFSGQSG-AAGALELL-GQPLATGVLELPG-QPGAPELPGQPVATVALEISV
-QSVVTTSELSTMTVSQSLEVPS-TTALE--SYNTVAQELPTTLVGETSVTVGVDPLMAQ
ESRMLASNTMETHMLASNTMDSQMLASNTMDSQMLASNSMDSQMLASSTMDSQMLASSTM
DSQMLATSSMDSQMLATSSMDSQMLATSSMDSQML------------ATSSMDSQMLATS
SMDSQMLATSSMDSQMLATSS--MDSQMLATSSMDSQMLATSSMDSQMLA--SGTMDSQM
LASGTMDAQMLASGTMDAQMLASSTQDSAMLDSKSPDPYRLAQDPYRLAQDAYRLGHDPY
RLGHDAYRLGQDPYRLGHDPYRLTPDPYRMSPRPYRIAPRSYRIAPRPYRLAPRPLMLAS
RRSMMMSYAAERSMMSSY--ERSMMSYERSMMSPMAERSMMSAYERSMMSAYERSMMSPM
AERSMMSAYERSMMSAYERSMMSPMADRSMMSMGADRSMMSSYSAADRS-MMSSYSAADR
SMMSSYTADRSMMSMAADSYTDSYTDTYTEAYMVPPLPPEEPPTMPPLPPEEPPMTPPLP
PEEPPEGSALPTEQSALTAENTWATEMPALPPEENVSHPEPPVGQSEISEPSAVPTNYSV
SGSESSMLV-SESVVTIPEPP-LEPESSVTSTPVESAVVA-EHEIVTERPVACVVSETAV
-MAEATLLTSEPPVVTETIETSESMKFER--HVASEASVSLLEAAV-----------TVP
EPAENIQ-----------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
-----------------------------------------ELSSVAINEADLVRPLLPK
DMERLTSLRAG-------IEGPLLPSEVERDKS-AASPVVISIPERASE-SSSEEKDDYE
IFVKVKDTHEKSKKNKNRDKGEKEKK----------------------------------
-----------------------------------------------RDSSLRSRSKRSK
SSEHKSRK-------RTSESRSRARKRSSKSKS-HRSQTRSRSRSRRRRRSSRSRSKSRG
RRSVSKEKRKRSPKHRSKSRERKRKRSS-SRDNRKTGRVRSRTPSRRSRSHTPSRRRRSR
SVGRRRSFSISPSRRSRT-----PSRRS--------------RTPSRRSRTPSRRSRTPS
---------RRSRTPSRRRRSRSV--VRRRSFSISPVR-LRRSRT-PLRRRFSRSPIRRK
RSRSSE---R-----ERLTDLDKAQLLEIAKANAAAMCAKAGVPLPPNLKPA-PP-PTIE
EKVAKKSGGATIEELTEKCKQIAQSKEDDDVIVNKPHVSDEE-EEEPPFYHHPFKLSEPK
PIFFNLNIAA---AKPTPPKSQVTLTKEFPVSSGSQHRKKEADSVYGEWVPVEKNGEENK
D-DDNVFSSNLPSEPVDISTAMSERALAQKRLSENAFDLEAM-SMLNRAQERVCGTL---
--------------IEQEQ--------LKTWDQFLRAXXXXXXXXXX
PF01585 // ENSGT00730000111141_0_Domain_1 (2628, 2661)
LLRKMGWREGEGL
GKNKEGNKEPILVDFKTDRKGNK-------------------------------FFLEIR
WQPPEFLLVHDSGPDHRKHFLFR-------------------------------------
----------------
ENSCAFP00000013423 (view gene)
MATNIEQIFRSFVVSKFREIQQELSSGRSEGQLNGETNTPNEGNQAGDA-AASARSLPNE
EIVQKIEEVLSGVLDTELRYKPDLKEAS-RKSRCVSVQTDPTD-EIPTKKSKKHKKHKNK
KKKKK----KEKEKKYKRQPEESESKVKSHHDGNIDLESDSFLKFDSEPSAMALEHPVRA
FGLSE------------------------TSESP-AVVLEPPVVSMEISESHTLETLKPA
TKTAELSV----ASTSVISVQSEQSVALTLEPSMTKILDSFTTAPVPTT---TVVLKS-P
EPVVTMSMEYQMKPVLKSLETIPPEQSQII--LEPPVAKGLESSETL-VVSPEIPTEVHP
EPSTSTTMDFPESSATEGLPLPEQPIEVPSEIADSSMTRPQELLELPKTTALELPESS-V
ASVMELPGPPATSKPELQGPPVTPVLELPGPSAT-----------PLPELPGPLSTPVPE
LLGPPATA--VPELPGPSATSVPQLSQELPGLPAPSVGLEPPQEVPEPPVMAQ----ELP
GLPAVTAAVELPGQPAVTVAMELTEQPVTTSELEQPVGMTTVEHPGPPEVTTATGLLGQP
EAAMVLELPGQPVATTALELPGQPSVTGVPELPGLPSATRALELSGQPVATGALEL-PGQ
LMATGA-LEFSGQSG-AAGALELL-GQPLATGVLELPG-QPGAPELPGQPVATVALEISV
-QSVVT-TELSTMTVSQSLEVPS-TTALE--SYNTVAQELPTTLVGETSVTVGVDPLMAQ
ESHMLASNTMETHMLASNTMDSQMLASNTMDSQMLASNTMDSQMLASSTMDSQMLATSSM
DSQMLATSSMDSQMLATSSMDSQMLATSSMDSQMLATSS--MDSQMLATSSMDSQM----
------LATSSMDSQMLATST--MDSQMLATSTMDSQMLATSSMDSQMLA--SGTMDSQM
LASGTMDAQMLASGTMDAQMLASSTQDSAMLGSKSPDPYRLAQDPYRLAQDPYRLGHDPY
RLGHDAYRLGQDPYRLGHDPYRLTPDPYRMSPRPYRIAPRSYRIAPRPYRLAPRPLMLAS
RRSMMMSYAAERSMMSSY--ERSMMSYERSMMSPMAERSMMSAYERSMMSAYERSMMSPM
AERSMMSAYERSMMSAYERSMMSPMADRSMMSMGADRSMMSSYSAADRS-MMSSYSAADR
SMMSSYTADRSMMSMAADSYTDSYTDTYTEAYMVPPLPPEEPPTMPPLPPEEPPMTPPLP
PEEPPEGPALPTEQSALTAENTWSTEVPALPPEESVSLSEPSVSQSEMSEPSALPANYSV
SASDPSVLA-SEAAVTVPEPP-LEPESSVTATPVESTVVAEEHQIVPERAVTYMVSETPI
MTAEPTVLTSEPSVMSETAETFDSMKASGHV-A-SEVSLSLLEPAA-----------TNP
EPSQNTL---------------E-LPAM--TISELPAVAVPQPPTGTVPEPPTVA-VLET
PAVAIPDPTAVA------VSDPVAVAVPDPPVEAVPETL----------ALAESEHVTIP
VPV-SAL------EPTVPVLEPVVSVPQ-PNTVVSE-------P-SVSVQESTVIISEPA
VTVSEQTQVIPDEMVLESTPMILEPT------VIKGVSILPG-DQNLAPEIGIQEIPMHA
DEE-PHAEGHLKNDPYESENGTNIELNVNNHLVAKEMEHNTVSAASTGAVDEIGEGNILS
ISETKQCTVLDTCSSVSEADGGT--LSSAGPL-ALEPDA----MGTSKGIEFATESALSS
VNKYDVEVSLTTQDTEHDMIISTSPSGGSEADIEGPLPAKDIHLDLPSN-NFMSK-DAEG
PLPIKECDQTLAVALSPKES-SGEDKEVALPTKEILSDSGFSANIDDINEADLVRPLLPK
DMERLTSLRAG-------IEGPLLASEVERDKS-AASPVVISIPERASE-SSSEEKDDYE
IFVKVKDTHEKSKKNKNRDKGEKEKK----------------------------------
-----------------------------------------------RDSSLRSRSKRSK
SSEHKSRK-------RTSESRSRARKRSSKSKS-HRSQTRSRSRSRRRRRSSRSRSKSRG
RRSVSKEKRKRSPKHRSKSRERKRKRSS-SRDNRKTVRARSRTPSRRSRSHTPSRRRRSR
SVGR-RSFSISPSRRSRT-----PSRRSRTPSRRSR--------------TPSRRSRTPS
RRSRT--PSRRSRTPSRRRRSRSV--VRRRSFSISPVR-LRRSRT-PLRRRFSRSPIRRK
RSRSSE---RGRS-PKRLTDLNKAQLLEIAKANAAAMCAKAGVPLPPNLKPA-PP-PTIE
EKVAKKSGGATIEELTEKCKQIAQSKEDDDVIVNKPHVSDEE-EEEPPFYHHPFKLSEPK
PIFFNLNIAA---AKPTPPKSQVTLTKEFPVSSGSQHRKKEADSVYGEWVPVEKNGEENK
D-DDNVFSSNLPSEPVDISTAMSERALAQKRLSENAFDLEAM-SMLNRAQERIDAWAQLN
SIPGQFTGSTGVQVLTQEQLANTGAQAWIKKDQFLRAAPV
PF01585 // ENSGT00730000111141_0_Domain_1 (2621, 2664)
TGGMGAVLMRKMGWREGEGL
GKNKEGNKEPILVDFKTDRKGLVENSGT00730000111141_0_Linker_1_2 (2664, 2686)
AVGERAQKRSGNFSAAMKDLSGPF14709 // ENSGT00730000111141_0_Domain_2 (2686, 2732)
KHPVSALMEICNKRR
WQPPEFLLVHDSGPDHRKHFLFRVLRNGSPYQPNCMFFLN-RY-----------------
----------------
ENSLAFP00000013694 (view gene)
MATNIEQIFRSFVVSKFREIQQELSSGRNEGQLNGETSTPIEGNQAGDS-AASARSLPNE
EIVQKIEEVLSGVLDTELRYKPDLKEAS-RKSRCVSVQTDPTD-EIPSKKSKKH------
-------------KKYKRQPEEPESKPKSHHDGNIDIESDSFLKFDSEPSVMGLEHPVRA
LGLSE------------------------TGDSP-AVVLEPPVVAMEVSEPHNFETLKPA
TKTAELQV----ESTSVISEQSEHSMAVMLEPSTAKILDCFPAAPVPTP---ALVLKS-P
EPVVTMSMEYQMTPVLKPLESTSPEPSKIVL-VEPPVAKMLESAETL-VESSETSAEVHP
EPSTSTTTGFPESSAPDVLRLTEQPVEVPLESADSSITRPQELLELPKATALELPESS-V
ASVMELQGPPATSMPEMQGPSVTPALELPGLSAT-----------LVPELPGPPSSPVPE
LPGPPATV--VSELPGPSVTPVPQLLQELPGLPASSVGLEPPQEVPEPPVMAQ----ELP
GLPVVTAAVELPGQPVVTVAMELTEQPVTTTELEQPLGMTTMEHPGQAEVTTATGLLRQP
EAAMVLELPGQPVATTALELPGQSSVTGVPELPGLPSATRALELSGQPVATGALEL-SGQ
LMAAGA-LEFSGQSG-AAGALELL-GQPLATGVLELPG-QPGAPELPGQPVATVALEISV
-QSVVTTSELSTMTVSQSLEVPS-TTALE--SYNTVAQELPTTLVGETSVTVGVDPLMAQ
ESHMLASNTMETHMLASNTMDSQMLASNTMDSQMLASNTMDSQMLASSTMDSQMLATSSM
DSQMLATSSMDSQMLATSSMDSQMLAT---------------------------------
-------------------SS--MDSQMLATSSMDSQMLATSSMDSQMLA--SGTMDSQM
LASGTMDAQMLASGTMDAQMLASSTQDSAMLDSKSPDAYRLAQDPYRLAQDPYRLGHDPY
RLGHDAYRLGQDPYRLGHDPYRLTPDPYRMSPRPYRIAPRSYRIATRPYRLAPRPLMLAS
RRSMMMSYAAERSMMSSY--ERSMMSYERSMMSPMAERSMMSAYERSMMSAYERSMMSPM
AERSMMSAYERSMMSAYERSMMSPM-DRSMMSMGADRSMMSSYSAADRS-MMSSYSAADR
SMMSSYTADRSMMSMAADSYTDSYTDTYTEAYMVPPLPPEEPPTMPPLPPEEPPMTPPLP
PEEPPEGPALTTEQSALTAESTWPTELPALPPEESVSLPEPPVSQSEISEPLAVPANYSV
SASEPTVLA-AEAIVGVAEPP-SEPVSSVTSTPVQSAEVA-EHEMIAETPVTSVVSETPI
MSAEPTMFTSQPALTPEMAETFDSLRASGHA-A-SEVSMSPLEPAV-----------TIP
ESTQSTS---------------E-LPAM--VVSELPSTVVPEPPAMAVPEPPAMA-VPEL
PAMAVTESSAVAVPEPLVAAGPEPLAVSEVLVVAVSESP----------AVIEPEHLPVP
VPVVSAL------EPTVPIVDPAVSVLQ-PAMIASE-------P-SVSVQESTLTVSEPA
VTVSAQTQVISTELVLESTSVILESSVMSSTHIIKGMNLISG-DSNLAPEIGMQEITMYS
GEE-PPAEGHLKSDCYGSEQGVSIDLNVNSHLITKEMEHNTASAASTGGVGEIGEEKILP
VNETEQCTVLVTCPGVSEAEVGGS-LSSVDPL-ALEPDA----KGTSEGIEFATASALSS
VNKYDVEVSLTTQDTEHDMVISTSPSGGSEADIEGPLPAKDIHFHVPSNHNLVGE-DAEG
PLPMKESDQAVAVALSPKES-SGEDKEVSLHSEEILPDTGFSANIEDINEADLVRPLLPK
DMERLTSLRAG-------IEGPLLPSEVERDKS-AASPGVIGLPERASE-SSSEEKDDYE
IFVKVKDTHEKSKKNKNRDKGEKEKK----------------------------------
-----------------------------------------------RDSSLRSRSKRSK
SSEHKSRK-------RTSESRSRARKRSSKSKS-HRSQTRSRSRSRRRRRSSRSRSKSRG
RRSVSKEKRKRSPKHRSKSRERKRKRSS-SRDNRKTVRARSRTPSRRSRSHTPSRRRRSR
SVGRRRSFSVSPSRRSR-------------------------------------------
------------------RRSMCT--GVRRSFSISPVR-LRRSRT-P-RRRFSRSPIRRK
RSRSSE---RGRS-PKRLTDLDKAQLLEIAKANAAAMCAKAGVPLPPNLKPA-PP-PSIE
EKVAKKSGGATIEELTEKCKQIAQSKEDDDVIVNKPHVSDEE-EEEPPFYHHPFKLSEPK
PIFFNLNIAA---AKPTPPKSQVTLTKEFPVSSGSQHRKKEADSVYGEWVPVEKNGEENK
DDDDNVFSSNLPSEPVDISTAMSERALAQKRLSENAFDLEAM-SMLNRAQERIDAWAQLN
SIPGQFTGSTGVQVLTQEQLANTGAQAWIKKDQFLRAAPV
PF01585 // ENSGT00730000111141_0_Domain_1 (2621, 2665)
TGGMGAVLMRKMGWREGEGL
GKNKEGNKEPILVDFKTDRKGLVAENSGT00730000111141_0_Linker_1_2 (2665, 2688)
VGERAQKRSGNFSAAMKDLSGKHPF00035 // ENSGT00730000111141_0_Domain_2 (2688, 2724)
PVSALMEICNKRR
WQPPEFLLVHDSGPDHRKHFLFRVLRNGSPYQPNCMFFLN-RY-----------------
----------------
ENSCLAP00000017154 (view gene)
MAADIEQVFRSFVVSKFREIQQELSSGRSEGQLNGETNTPIEGNQAGD-AAASARSLPNE
EIVQKIEEVLSGVLDTELRYKPDLKEAS-RKSRCVSVQTDPTD-EIPSKKSKKHKKHKNK
KKKKK----KEKEKKYKRQPEESESKLKSHHDGNIDLESDSFLKFDSEPSSMALEHPIKA
FGLSE------------------------TSESS-AVVLEPPVSSMELSESHILETLKPA
TKTAEIPV----GSTSVISE---QSMAVMLEPSVTKIVDSFATAPVPTT----VVLKS-S
EPLATMSMEYQMKSALKSLEHTSPEPSKIML-AEPPVAKELQPSETR-VIASETSTEVHP
EPSTSTTKDCPESSASEVLRLPEHPV--PSEIADPSMTRPQDLLELSKTSALELPESS-V
ASALELPGPPATSMPELQGPPVTPVLELPGPSA-----------TPVSELSGPLSTPVPE
LPGPPATA--VPELPGPSVTPVPQLSQELPGPLAPSMGLEPPQEVPEPPVMAQ----ELP
GLSTVSATVELPGQPAVTVAMELTEQPVTTAEFEQPVGMTAVEHPGHPEVTAATGLLGQP
EAAMVLELPGQPVATTALELSGQPSVTGVPELPGLPSATRALELSGQSVATGALEL-PGQ
LMATGA-LEFSGQSG-AAGALELL-GQPLATGVLELPG-QPGAPELPGQPVATVALEISV
-QSVVTTSELSTMTVSQSLEVPS-TTALE--SYNTVAQELPTTLVGETSVTVGVDPLMAQ
ESHMLASNTMETHMLASNTMDSQMLASNTMDSQMLASNTMDSQMLASSTMDSQMLASSTM
DSQMLATSSMDSQMLA------------------------------------------TS
SMDSQMLATSSMDSQMLATTTSSMDSQMLATSSMDSQMLATSSMDSQMLA--SGTMDSQM
LASGTMDAQMLASGTMDAQMLASSTQDSGMLDSKSPDPYRLAQDPYRLAQDAYRLGHDPY
RLGHDAYRLGQDPYRLGHDPYRLTPDPYRMSPRPYRIAPRSYRIAPRPYRLAPRPLMLAS
RRSMMMSYAAERSMMSSY--ERSMMSYERSMMSPMAERSMMSAYERSMMSAYERSMMSPM
AERSMMSAYERSMMSAYERSMMSPMADRSMMSMGADRSMMSSYSAADRS-MMSSYSAADR
SMMSSYTADRSMMSMAADSYTDSYTDTYTEAYMVPPLPPEEPPTMPPLPPEEPPMTPPLP
PEEPPEGSALPTEQSALTAENTWSAEMPALPPEESVSHPEPPVSQSEILEPSAVP-DYSV
SGSEPSVLV-SEAVVTMPEPP-LELETSVTSTPVESAIVA-EHEIVPERPVTCMVSETPI
-LAEASVLTSESPVVTETMETLESMKFEG--HVASEASVPLVEPAV-----------TIP
EPAENIQEMPAMAVSESAVMAVSESIVVAVPAPELPAVAVPQAPAMAFPESPAVAAPEPP
-ATGKPEPP------------------------------AV--AVVEPPSVAEPELVSVP
VSAVSSL------EPTVPVLEPAVSVLQPAMIVSE---------PCVSVQQYTVTVSEPA
VTVSEQTQVISTEVAVESTPVIAESSLMS-SHVMKGMNSLSD-DQSLAPEIGMQEN-PLH
SGEEPHAEGHLKSDSHESEHGINVDLTINNHLIAKAMET---------------------
-------------------------------------------G-ASKGIEFVTTPAFTA
VNNY-DDISLTAQDTEHDMVISTSPSGVSEADIEGPFGE---------------------
--------------------------EVPLSAKEVVSDSGFCANIDEINEADLVRP----
---------AG-------IEGPLLPSEVERDKS-AASPVVISIPERASE-SSSEEKDDYE
IFVKVKDTHEKSKKNKNRDKGEKEKK----------------------------------
-----------------------------------------------RDSSLRSRSKRSK
SSEHKSRK-------RTSESRSRARKRSSKSKS-HRSQTRSRSRSRRRRRSSRSRSKSRG
RRSVSKEKRKRSPKHRSKSRERKRKRSS-SRDNRKTGRARSRTPSRRSRSHTPSRRR---
--------SISPSRRSRT-----PSRRS--------------RTPSRRSRTPSRRSRTPS
RRSRT--PSRRSRTPSRRRRSRSV--VRRRSFSISPVR-LRRSRT-PLRRRFSRSPIRRK
RSRSSE---R-----ERLTDLDKAQLLEIAKANAAAMCAKAGVPLPPNLKPA-PP-PTIE
EKVAKKSGGATIEELTEKCKQIAQSKEDDDVIVNKPHVSDEE-EEEPPFYHHPFKLSEPK
PIFFNLNIAA---AKPTPPKSQVTLTKEFPVSSGSQHRKKEADSVYGEWVPVEKNGEENK
D-DDNVFSSNLPSEPVDISTAMSERALAQKRLSENAFDLEAM-SMLNRAQERIDAWAQLN
SIPGQFTGSTGVQVLTQEQLANTGAQAWIKKDQFLRAAPV
PF01585 // ENSGT00730000111141_0_Domain_1 (2621, 2664)
TGGMGAVLMRKMGWREGEGL
GKNKEGNKEPILVDFKTDRKGLVENSGT00730000111141_0_Linker_1_2 (2664, 2687)
AVGERAQKRSGNFSAAMKDLSGKPF14709 // ENSGT00730000111141_0_Domain_2 (2687, 2756)
HPVSALMEICNKRR
WQPPEFLLVHDSGPDHRKHFLFRVLINGSAYQPSFASPNK-KHAKATAATVVLQAMGLVP
KDLMANATCFRSASRR
ENSODEP00000004818 (view gene)
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
--------------------------------------------------------MAAD
IEQVFRSFVVSKFREIQQ-----ELSSG--------------RSE-----------GQLN
GETNT--PVEGNQAGDAAAS----------------------------ARNLPNEEIVQK
IEEVLS---GVLD-TELRYKPDKAQLLEIAKANAAAMCAKAGVPLPPNLKPA-PP-PTIE
EKVAKKSGGATIEELTEKCKQIAQSKEDDDVIVNKPHVSDEE-EEEPPFYHHPFKLSEPK
PIFFNLNIAA---AKPTPPKSQVTLTKEFPVSSGSQHRKKEADSVYGEWVPVEKNGEENK
D-DDNVFSSNLPSEPVDISTAMSERALAQKRLSENAFDLEAM-SMLNRAQERIDAWAQLN
SIPGQFTGSTGVQVLTQEQLANTGAQAWIKKDQFLRAAPV
PF01585 // ENSGT00730000111141_0_Domain_1 (2621, 2664)
TGGMGAVLMRKMGWREGEGL
GKNKEGNKEPILVDFKTDRKGLVENSGT00730000111141_0_Linker_1_2 (2664, 2687)
AVGERAQKRSGNFSAAMKDLSGKPF14709 // ENSGT00730000111141_0_Domain_2 (2687, 2732)
HPVSALMEICNKRR
WQPPEFLLVHDSGPDHRKHFLFRVLRNGSPYQPNCMFFLN-RY-----------------
----------------
ENSCCAP00000024911 (view gene)
ENSSBOP00000007974 (view gene)
ENSXETP00000030802 (view gene)
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------RKSFSRSPVRRK
RSRSRDKSLDSSRSPKRLTDLDKAQLLEIAKANAAAMCAKAGVPLPPSLKPVIAPVTPVE
EKITLRSYGVTIQELTEKCKQIAQSKE-DDVIVNKPHDSDEEEEEERPFYNHPFKVSEHK
PISFSLLNPSVK----PAPKNQVTLTKEFPVSSGSQHRKKEADKVYGEWVPVDKK-SEES
--KDDVFTNTGPSQPVDITSAMNERALAQTRLTGNPFDMEAL-IMLNRAQEQIDAWAQST
SLPGQFTGSTGAQVLSADEISNSGPQAWLKKDQFLKAAPV
PF01585 // ENSGT00730000111141_0_Domain_1 (2621, 2665)
TGGRGALLMRKMGWKEGKGL
GRHNEGNVDPILIDFKTDRKGLVAENSGT00730000111141_0_Linker_1_2 (2665, 2688)
DGEKASNKL-ALP-MMKDLSGKHPF00035 // ENSGT00730000111141_0_Domain_2 (2688, 2756)
PISALMELCNKKK
WSPPEFVLVDDTGPEHRKRFLFRVTVNGGVYQPNQPSVNK-KLAKATAAAAALQALGALP
KESMTSTANFRSASTS
MGP_SPRETEiJ_P0044192 (view gene)
ENSETEP00000015781 (view gene)
MATNIEQIFRSFVVSKFREIQQELSSGRNEGQLNGETSAPIEGNQAGDS-SASARSLPNE
EIVQKIEEVLSGVLDTELRCKPDLKEAS-RKSRCVSVQTDPTDEEVPAKKSKKHKKHKNK
KKKKK----KEKEKKYKRQPEESEAKLKSH-----DIESDSFLKFDSEPSAMGLAHPIRE
LGLPE------------------------TSDSP-SVAPEPSLIAMEVSEPHNLETLKQA
TKPAELSI----ASTSVISEQSEHSVTVTLEPYMTKMVDSFATASVPTT---TLVLKS-S
DPVVTMSAEYQMKSVLTSWESTSPEPSKIML-VEPPVAKVLEPTETV-VESSETPAEVYP
EPSTSTTLDFPESSGPEVLRLPEQPVEASMEIAESPMTRPKELLELPKTAALELPESSLV
ASAMELPGPPATSMPELQGHPVTPVLELPGPSAT-----------PVAELPGPPSTPVPE
LPGPPATV--VSELPGPSVTPVPQLLQELPGLPASSVGLEPPQEVPEPPVMAQ----ELP
GLPVVTAATELPGQPVVTVAMELTEQPVTTTELEQPMGMTTMEHPGQPEVTTATGLLGQS
EATMVLELPGQSVATTALELPGQPSVTGVPELPGLPSATRALELSGQPVATGALDL-SGQ
LMAAGA-LEFSGQSG-AAGALELL-GQPLATGVLELPG-QPGAPELPGQPVATVALEISV
-QSVVTTSELSTMTVSQSLEVPS-TTALE--SYNTVAQELPTTLVGETSVTVGVDPLMTQ
ESHMLASNTMETHMLASNTMDSQMLASNTMDSQMLASNTMDSQMASST------------
---------------------------MDMDFQMLATSS--MDFQM-ATSSMDSQMLATS
SMDSQMLATSSMDSQMLATSS--MDSQMLATSTMDSQMLATSSMDSQMLA--SGTMDSQM
LASGTMDAQMLASGTMDAQMLASSSQDSAMLDSKSPDPYRLAQDPYRLAQDPYRLGHDPY
RLGHDAYRLGQDPYRLGHDPYRLTPDPYRMSPRPYRIAPRSYRIAPRPYRLAPRPLMLAS
RRSMMMSYAAERSMMSSY--ERSMMSYERSMMSPMAERSMMSAYERSMMSAYERSMMSPM
AERSMMSAYERSMMSAYERSMMSPMADRSMMSMGADRSMMSSYSAADRS-MMSSYSAADR
SMMSSYTTDRSMMSM-ADSYTDAYADTYTEAYMVPPLPPEEPPTITPLPPEEPPMTPPXX
XXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXX-XXXXXX
XXXXXXXXXXXXXXXXXXXXX-XXXXXXXXXIAVQSAVVV-EHEIIPETSV--TMTETPI
VSAEPAMLTSEPPIMSETAETFESMSTAGHA-SEIC--VPILQSTV-----------TIP
ESAQSTQE----------------LPAM--PVTELPAMSVPESMAVAVREP---------
----------------LAVAVPEPLAVSEVLVVAVPEPP----------AMTDPEHVAIS
VPVVSAL------EPTVSMVEPAVSVLQPAMIVSE---------PTVSVQESTLAVSEPS
VTVSEQTQVISTKLVLETTSMILESSVMSSTHIIKGMNLISS-NSNLAPEIGMQGITMYA
GEE-SHAEGHLKSDTYGSEHGMGIDLNINN-LIRSEMEHTTVSAASTGV--KIGKEKNLP
VNETEQCTVL-ACPGVSEADVRTL---SSDP---LEHDA----KATSKGIEFATTSAFSS
DNKYDVKVSVTTQDSEHDMVISTSPSGGSEADIEGPLLAKDFHLELPTN-NFAS-EDAER
PLPMPESDQTMTVDLNPKEN--SEEKEVPLPAKEILPNSGFPTNIEDINEADLVRPLLPK
DMERLTNLRAA-------IEGPLLASEVERDRSVA-SP--VGVAERASE-SSSEEKDDYE
IFVKVKDTHEKSKK-KNRDKGDKDKK----------------------------------
-----------------------------------------------KDSSLRSRSKRSK
SSEHKSRK-------RTSESSRXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXX
XXXXXXXXXXX--XXXXXXXXXXXXXXX-XXXXXXXXXXXXXXXXXXXXXXXXXXXXXXX
XXXXXXXXXXXXXXXXXX-XX--XXXXXXXXXXX--X-XX-X--------------XXXX
XXXXX--XXXXXXXXXXXXXXXXX--XXXXXXXXXXXX-XXXXXX-XXXXXXXXXXXXXX
XXXXXX---XXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXX-XXXX
XXXXXXXXXXXXXXXXXKCKQIAQSKEDDDVIVNKPHVSDEE-EEEPPFYHHPFKLSEPK
PIFFNLNIAAA--AKPTPPKSQVTLTKEFPVSSGSQHRKKEADSVYGEWVPVEKNGEENK
DEDDNVFSSTLPSEPVDISTAMTERALAQKRLSENAFDLEAM-SMLNRAQERIDAWAQLN
SIPGQFTGSTGVQVLTQEQLANTGAQAWIKKDQFLRAAPV
PF01585 // ENSGT00730000111141_0_Domain_1 (2621, 2660)
TGGMGAVLMRKMGWREGEGL
GKNKEGNKEPILVDFKTDRK----------------------------------------
------------------------------------------------------------
----------------
ENSNLEP00000004882 (view gene)
ENSPTRP00000053981 (view gene)
ENSCATP00000011175 (view gene)
ENSPTRP00000072700 (view gene)
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
--------------------------------------------------------MATN
IEQIFRSFVVSKFREIQQ-----ELSSGRN-------------------------EGQLN
GETNT--PIEGNQAGD------AA--A--------------------SARSLPNEEIVQK
IEEVLS---GVLD-TELRYKPDKAQLLEIAKANAAAMCAKAGVPLPPNLKPA-PP-PTIE
EKVAKKSGGATIEELTEKCKQIAQSKEDDDVIVNKPHVSDEE-EEEPPFYHHPFKLSEPK
PIFFNLNIAA---AKPTPPKSQVTLTKEFPVSSGSQHRKKEADSVYGEWVPVEKNGEENK
D-DDNVFSSNLPSEPVDISTAMSERALAQKRLSENAFDLEAM-SMLNRAQERIDAWAQLN
SIPGQFTGSTGVQVLTQEQLANTGAQAWIKKDQFLRAAPV
PF01585 // ENSGT00730000111141_0_Domain_1 (2621, 2664)
TGGMGAVLMRKMGWREGEGL
GKNKEGNKEPILVDFKTDRKGLVENSGT00730000111141_0_Linker_1_2 (2664, 2687)
AVGERAQKRSGNFSAAMKDLSGKPF14709 // ENSGT00730000111141_0_Domain_2 (2687, 2732)
HPVSALMEICNKRR
WQPPEFLLVHDSGPDHRKHFLFRVLRNGSPYQPNCMFFLN-RY-----------------
----------------
ENSMODP00000037779 (view gene)
MATNIEQIFRSFVVSKFREIQQELSSGRVEGQLNGETNPPVEGNQSGDA-AACARSLPNE
EIVQKIEEVLSGVLDTELRYKPDLKEAS-RKSRCVSVQTDPTD-EIPTKKSKKHKKHKNK
KKKKKK--EKEKEKKYKRQSEESDSKLKSHHGGNVGLESDSYLKFDSEPSVMGLEHPVRA
LGLPKPPAVMLKSPT----------MTLGLNESP-AVALEPPTIAVEISEPHTLVALKPT
SVGELSELHAALVPEPFTLEQPETSVAVTLESSVKQTLESPAVMPVPTT---AVVQKS-S
EAYVTVSSEYNARSVTKHLEIPSSEQSKTTL-LEPPASKVLELSETSGVIASETPAEAYP
RPRSSITLELPESSVTEVLRLPEQSTTIPLLS-----------MEPPTTTVLELPKPP-A
ISVLEMPGPPTTPVPELPGPPTTPMLELPGPPMT-----------SAPELLGPPVTLVPE
LPGPPATLVQVPELPGPSATPVLELSQDLPGLPAPSMVSELPEEVPRQPTMALELPQELP
GLPVVTTAMELPGQPVATVALELAEQPVMTTEPEQPTVMPALELPGQPEVTTASGLPGQP
VATTAPELPGQPEV---PELRGLPSVTGVSELPGQPSATVAQELPGQSGA----------
----------LGQPV-ATVALELL-GQPVASELSGQPT-GAAALELPGQPVATVALEISS
-HSVMTTSDLTTMTVSQSMEVPS-TTALE--SYSTVAQELPTTLVGDTSVTVSVDPLMTP
ESHMLASNTMDSQMLASNTMDSQMLASNTMDSQMLASNTMDSQMLASNTMDSQMLASNSM
DSQMLASNSMDSQMLASNSMDSQMLATSTMDSQMLATST--MDSQMLATSTMDSQMLATS
TMDSQMLATSTMDSQMLATST--MDSQMLATSTMDSQMLATSTMDSQMLA--SSTMDSQM
LATSTMDSQMLASSTMDAQMLASSTPESSMLGPKSPDPYRLAQDPYRLAQDPYRLGHDPY
RLGHDPYRLGQDPYRLGHDPYRLAPDPYRMSPRPYRIAPRPYRIAPRPYRLAPRPLMLAS
RRSMMMSYAAERSMMSSYAAERSMMSYERSMMSPMAERSMMSAYERSMMSAYERSMMSPM
AERSMMSAYERSMM-AYERSMMSPMAERSMMSMGADRSMMSSYS--DRS-MMSSYS--DR
SMMSSYTADRSMMSMTADSYADSYTDSYTEAYMVPPLPPEEPPTMPPLPPEEPPMTPPLP
PEEPPEGPSLPVEQSALPVENTWATAGTALLSEEPALPPEPPVMQTEIPEST--MSDYSV
GAAEPSVLT-SEPAITISEPP-PEPESSMTSTSLESHMTQEHAIIGPQV--AVLSSGAVS
ETAIPTAESTESPIVTESVETFEPILASGPI-SDQM--MFLSEVAM-----------TVP
EPAEVAVPEPAEAVI-------PEPAEM--TVTGLTEVAVQGLAEVAVQGPAEMA-VQGS
--------------TEVMIQGPEEVTYQAPTEVAYQAPTEVAYQAPTEVAYQAPTEVAYQ
APTEVAYQAPTEVAYQAPTEEPATVVVQESAMVVPELTVALSEPAMVEVAKPEATVLEPF
VTVPEGTLV-----VPELTPVMMESTVMTSEPVIKRTDIATSAEPNLPQQSAIQEMFIQS
SEESNNEKRQLESHSYESIHHVNIILDVNSPLMTKETEHNTVSATS-PVVSEIGMEKFLP
GNETDRNTILTTCSSE--ANIVET-VSTTSSH-IHELDV----KGTTKEIELYAASVTSS
ISKADIEGPLPTQEIEHDMVISTSPSGGSEADIEGPLPANDIHHDILSANNLLSKNEPEA
SLPVKDTEHDVLIAISHKDGSSEAEKETPLPAKNIEHESVPAANICDLNDADLVRPLLPK
DMERVTNLRTD-------VEGPLPASGVERDIVTTASPIVISVPERASE-SSSEEKDDYE
IFVKVKDTHDKSKKTKSRDKGDKERK----------------------------------
-----------------------------------------------RDSSLRSRSKRSK
SSEHKSRR-------RTSESRSRARKRSSKSKS-HRSQTRSRSRSRRRRRSSRSRSKSRG
RRSLSKEKRKRSPKHRSKSRERKRKRSS-SRDNRKVARARSRTPSRRSRSHTPSRRRRSR
SVVRRRSFSISPVRRSRT-----PSRRSRTPSRRSRTPSR--------------------
----------RSRTPSRRRRSRSV--VRRRSFSISPVR-LRRSRT-PLRRRFSRSPIRRK
RSRSSE---RGRSPPKRLTDLDKAQLLEIAKANAAAMCAKAGVPLPPNLKPA-PP-PTVE
EKVAKKSGGATIEELTEKCKQIAQSKEDDDVIVNKPHVSDEE-EEERPFYHHPFKLNEPK
PIFFNLNIAA---AKPTPPKNQVTLTKEFPVSSGSQHRKKEADSVYGEWVPVEKNGEENK
D-DDNVFSSNLPTEPVDISTAMSERALAQKRLSENAFDLEAM-SMLNRAQERIDAWAQLN
SIPGQFTGSTGVQVLTQEQLANTGAQAWIKKDQFLRAAPV
PF01585 // ENSGT00730000111141_0_Domain_1 (2621, 2664)
TGGMGAVLMRKMGWREGEGL
GKNKEGNKEPILVDFKTDRKGLVAVGERAQKRSGNFSAAMKDLSGKHPVSALMEICNKRR
WQPPEFLLVHDSGPDHRKHFLFRVLRNGSPYQPNCMFFLN-RY-----------------
----------------
ENSPPYP00000012703 (view gene)
MATNIEQIFRSFVVSKFREIQQELSSGRNEGQLNGETNTPIEGNQAGDA-AASARSLPNE
EIVQKIEEVLSGVLDTELRYKPDLKEAS-RKSRCVSVQTDPTD-EIPTKKSKKHKKHKNK
KKKKK----KEKEKKYKRQPEESESKTKSHDDGNIDLESDSFLKFDSEPSAMALELPTRA
FGLSE------------------------TNESP-AVVLEPPVVSMEVSEPHILETLKPA
TKTAELSV----ASTSVISEQSEQSVAVMPEPSMTKILDSFATAPVPIT---TVVLKS-S
EPVVTMSVEYQMKSLLKSVESTSPEPSKIML-VEPPVAKVLEPSETL-VVSSETPTEVYP
EPSTSTTMDFPESSPIEALRLPEQPVDVPSEIADSSMTRPQELPELPKTTALELQESS-V
ASAMELPGPPATSMPELQGPPVTPVLELPGPSAT-----------PVPELPGPLSTPVPE
LLGPPATA--VPELPGPSVTPVPQLSQELPGLPAPSMGLEPPQEVPEPPVMAQ----ELP
GLPLVTAAVELPEQPAVTVAMELTEQPVTTTELEQPVGMTTVEHPGHPEVTTATGLLGQP
EATMVLELPGQPVATTALELPGQPSVTGVPELPGLPSATRALELSGQPVATGALEL-PGP
LMAAGA-LEFSGQSG-AAGALELL-GQPLATGVLELPG-QPGAPELPGQPVATVALEISV
-QSVVTTSELSTMTVSQSLEVPS-TTALE--SYNTVAQELPTTLVGETSVTVGVDPLMAP
ESHILASNTMETHILASNTMDSQMLASNTMDSQMLASNTMDSQMLASSTMDSQMLATSTM
DSQMLATSSMDSQMLATSTMDSQMLATSSMDSQMLATSS--MDS----------QMLATS
SMDSQMLATSSMDSQMLATST--MDSQMLATSTMDSQMLATSSMDSQMLA--SGTMDSQM
LASGTMDAQMLASGTMDAQMLASSTQDSAMLGSKSPDPYRLAQDPYRLAQDPYRLGHDPY
RLGHDAYRLGQDPYRLGHDPYRLTPDPYRMSPRPYRIAPRSYRIAPRPYRLAPRPLMLAS
RRSMMMSYAAERSMMSSY--ERSMMSYERSMMSPMAERSMMSAYERSMMSAYERSMMSPM
AERSMMSAYERSMMSAYERSMMSPMADRSMMSMGADRSMMSSYSAADRS-MMSSYSAADR
SMMSSYTADRSMMSMAADSYTDSYTDTYTEAYMVPPLPPEEPPTMPPLPPEEPPMTPPLP
PEEPPEGPALPTEQSALTAENTWPTEVPSLPSEESVSQPEPPVSQSEISQPSAVPTDYSV
SASDPSVLV-SEAAVTVPEPP-PEPESSITSTPVESAVVAEEHEVVPERPVTCMVSETPV
MPVEPTVLASEPPVMSETAETFDSMRASGHV-A-SEVSTSLLVPAV-----------TTP
VLAESILE----------------PPAM--AAPESSAMAVLESSAVTVLESSTVT-----
------------------VLESSTVTVLEPSVVTVPEPP----------VVAEPDYVTIP
VPVVSAL------EPSVPVLEPAMSVLQPSMIVSE---------PSVSVQESTVTVSEPA
VTVSEQTQVIPTEVAIESTPMILESSIMS-SHVMKGINLSSG-DQNLAPEIGMQEIALHS
GEE-PHAEGHLKSDSYESEHGINIDLNINNHLIAKEMEHNTVCAAGTSPVGEIGEEKILP
TSETKQCTVLDTYAGVSEADAGET-VSSTGPL-ALEPDA----TGTSKGIEFITASTLSS
VNKYDVDVSLTTQDTEHDMVISTSPSGGSEADIEGPLPAKDIHLDLPSNNNLVSK-DTEE
PLPVKESDQTLAALLSPKES-SGGEKEVPPPPKETLSDSGFSANIEDINEADLVRPLLPK
DMERLTSLRAG-------IEGPLLASDVGRDKS-AASPVVSSMPERASE-SSSEEKDDYE
IFVKVKDTHEKSKKNKNRDKGEKEKK----------------------------------
-----------------------------------------------RDSSLRSRSKRSK
SSEHKSRK-------RTSESRSRARKRSSKSKS-HRSQTRSRSRSRRRRRSSRSRSKSRG
RRSVSKEKRKRSPKHRSKSRERKRKRSS-SRDNRKTVRARSRTPSRRSRSHTPSRRRRSR
SVGRRRSFSISPSRRSRT-----PSRRSRTPSRRSRTPSRRSRTP--------------S
RRSRT--PSRRSRTPSRRRRSRSV--VRRRSFSISPVR-LRRSRT-PLRRRFSRSPIRRK
RSRSSE---RGRS-PKRLTDLDKAQLLEIAKANAAAMCAKAGVPLPPNLKPA-PP-PTIE
EKVAKKSGGATIEELTEKCKQIAQSKEDDDVIVNKPHVSDEE-EEEPPFYHHPFKLSEPK
PIFFNLNIAA---AKPTPPKSQVTLTKEFPVSSGSQHRKKEADSVYGEWVPVEKNGEENK
D-DDNVFSSNLPSEPVDISTAMSERALAQKRLSENAFDLEAM-SMLNRAQERIDAWAQLN
SIPGQFTGSTGVQVLTQEQLANTGAQAWIKKDQFLRAAPV
PF01585 // ENSGT00730000111141_0_Domain_1 (2621, 2664)
TGGMGAVLMRKMGWREGEGL
GKNKEGNKEPILVDFKTDRKGLVENSGT00730000111141_0_Linker_1_2 (2664, 2688)
AVGERAQKRSGNFSAAMKDLSGKHPF00035 // ENSGT00730000111141_0_Domain_2 (2688, 2755)
PVSALMEICNKRR
WQPPEFLLVHDSGPDHRKHFLFRVLINGSAYQPSFASPNK-KHAKATAATVVLQAMGLVP
KDLMANATCFRSASRR
ENSPCAP00000009555 (view gene)
MATNIEQIFRSFVVSKFREIQQELSSGRNEGQLNGETSTPVEGNQAGDA--ASARSLPNE
EIVQKIEEVLSGVLDTELRYKPDLKEAS-RKSRCVSVQTDPTD-EIPSKKSKKHKKHKNK
KKKKK----KEKEKKYKRQPEDSESKPKSQHTGNPDTEPGPFPQADLEPSVVTSEPLGRA
LGSPE------------------------TSTSP-AVLEPPAVVALEASEPRAVEAWPPA
A------------ATPVISELSEHSRAGMLEPSVTPILDSFETAPLPGP---VPVLRS-S
EPAVTLSMDYQVVLVGKPVESTSLEPSGIMW-TEPPEAEVFESAERL-VESSGTAVEGPA
EPGLSVAVGFPASSASEELREPELPVEAPLESADAPMTRPQASPG--------LPEAS-V
APVMELPGPPATSVPERQGPPVTPVLELPGPSAV-----------PVPEVPGPPATPVPE
VPGLPAAV--ESELPGPSVTPGPQLLQEVPWLPASP-ALEPPQ-EPEPPVMAQ-----GP
GPPVMTAAVXXXXXXXXXXXXXXXXXXXXXXXXXXXVGLTAMEHPGQPEVTVASGLLRQP
EAAMVLELPGQPVAATALELAGQSSVAGVPELSGLPSAARALELSGQPVAAGALEL-SGQ
LMATGA-LEFSGQSG-AAGALELL-GQPLATGVLELPG-QPGAPELPGQPVATVALEISV
-QSVVTTSELSTMTVSQSLEVPS-TTALE--SYNTVATELPTALVGETSVTVGVDPLMAQ
ESHMLASNTMDSQMLASSSMDSQMLASNTMDSQMLASSTMDSQMLATSSMDSQMLATSSM
DSQMLA--------------------TSSMDSQMLATSS--MDSQMLATSSMDSQMLATS
SMDSQMLATSSMDSQMLATSS--MDSQMLATSTMDSQMLATSSMDSQMLA--SGTMDSQM
LASGTMDAQMLASGTMDAQMLASTTQDSAMLDSKSPDAYRLAQDPYRLAQDPYRLGHDPY
RLGHDAYRLGQDPYRLGHDPYRLTPDPYRMSPRPYRIAPRSYRIAPRPYRLAPRPLMLAS
RRSMMMSYAAERSMMSSY--ERSMMSYERSMMSPMAERSMMSAYERSMMSAYERSMMSPM
AERSMMSAYERSMMSAYERSMMSPM-DRSMMSMGADRSMMSSYSAADRS-MMSSYSAADR
SMMSSYTADRSMMSMAADSYTDSYSDTYTEAYMVPPLPPEEPPTMPPLPPEEPPMTPPLP
PEEPPEGSALTAEQSTLTAEDTWPAEVPVLSSEESAQLPELPASQSEISEPLAVPAGYAV
SVSEPSLVA-AGAAVAVPEPP-PEPESSVTPAPVPSAVAA-ADEMVTEAPVTYALPEVPV
MSAEPAVLTSGPPAVPKMADALDSLSASGGA-A-SAASMSLLEPAVTAAGSAQGPSEL--
--VAMA--------V-------QEPPAV--VVPALPVMAVTESSAVAVPET---------
-------------------------AVSEVLVVAVSEPP----------AVTEPAHLPVP
VPVVSAL------EPMVSAGEPAVAVLPPPMVPAE---------PSVSVEETSLMVSEPT
VTVSAQTQVISTELVLESTSVILESSVMSSTHIIKGMNLISG-DSHLASEIGMQEITMYS
HEE-PRAEGHLKRDSYGSDQGVSRDLHVSDHLMAAMTEHGPVPAASTGAVGAVGEEGGFP
VSEPEQSTVLAAGPGVGEAEGAGALP--SDPL-ALEAEADTLAVEAGGSAELAMAAAFTA
VDKYDGEVSLTAQDTEHDMVISSSPSGGSEADIEGPLPAKDIHLDLPSHHNLVGK-DAEG
PLPVTESGQAVAVALSPQES-SGECKEEPLPPEGVLPESGLPTSIEDINEADLVRPLLPK
DMERLTSLRAG-------IEGPLLPSEVERDRA-AASPGGIGLPERASE-SSSEEKDDYE
IFVKVKDTHEKSKKNKNRDKGDKEKK----------------------------------
-----------------------------------------------RDSSLRSRSKRSK
SSEHKSRK-------RTSESRSRARKRSSKSKS-HRSQTRSRSRSRRRRRSSRSRSKSRG
RRSVSKEKRKRSPKHRSKSRERKRKRSS-SRDNRKTVRARSRTPSRRSRSHTPSRRRRSR
SVGRRRSFSVSPSRRSRT-----PSRRSRTPSRRSRTPSRRS--------------RTPS
RRSRT--PSRRSRTPSRRRRSRSV--VRRRSFSISPVR-LRRSRT-PLRRRFSRSPIRRK
RSRSSE---RGRS-PKRLTDLDKAQLLEIAKANAAAMCAKAGVPLPPNLKPA-PP-PSLE
EKVAKKSGGATIEELTEKCKQIAQSKEDDDVIVNKPHVSDEE-EEEPPFYHHPFKLSEPK
PIFFNLNIAA---AKPTPPKSQVTLTKEFPVSSGSQHRKKEADSVYGEWVPVEKNGEESK
D-DDNVFSSNLPSEPVDISTAMSERALAQKRLSENAFDLEAM-SMLNRAQERIDAWAQLN
SIPGQFTGSTGVQVLTQEQLANTGAQAWIKKDQFLRAAPV
PF01585 // ENSGT00730000111141_0_Domain_1 (2621, 2660)
TGGMGAVLMRKMGWREGEGL
GKNKEGNKEPILVDFKTDRK----------------------------------------
------------------------------------------------------------
----------------
ENSMLEP00000021708 (view gene)
ENSGGOP00000010230 (view gene)
ENSFCAP00000027852 (view gene)
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
--------------------------------------------------------MATN
IEQIFRSFVVSKFREIQQ-----ELSSGRN-------------------------EGQLN
GETNT--PNEGNQAGD------AA--A--------------------SARSLPNEEIVQK
IEEVLS---GVLD-TELRYKPDKAQLLEIAKANAAAMCAKAGVPLPPNLKPA-PP-PTIE
EKVAKKSGGATIEELTEKCKQIAQSKEDDDVIVNKPHVSDEE-EEEPPFYHHPFKLSEPK
PIFFNLNIAA---AKPTPPKSQVTLTKEFPVSSGSQHRKKEADSVYGEWVPVEKNGEENK
D-DDNVFSSNLPSEPVDISTAMSERALAQKRLSENAFDLEAM-SMLNRAQERIDAWAQLN
SIPGQFTGSTGVQVLTQEQLANTGAQAWIKKDQFLRAAPV
PF01585 // ENSGT00730000111141_0_Domain_1 (2621, 2664)
TGGMGAVLMRKMGWREGEGL
GKNKEGNKEPILVDFKTDRKGLVENSGT00730000111141_0_Linker_1_2 (2664, 2687)
AVGERAQKRSGNFSAAMKDLSGKPF14709 // ENSGT00730000111141_0_Domain_2 (2687, 2732)
HPVSALMEICNKRR
WQPPEFLLVHDSGPDHRKHFLFRVLRNGSPYQPNCMFFLN-RY-----------------
----------------
ENSCSAP00000006065 (view gene)
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
-MMSSYTADRSMMSMAADSYTDSYTDTYTEAYMVPPLPPEEPPTMPPLPPEEPPMTPPLP
PEEPPEGPALPTEQSALTAENTWPAEVPSLPSEESVSQPEPPVSQSEISEPSAVPTDYSM
SASDPSVLV-SEATVTVPEPP-PEPESSITSTPVESAVVAEEHEVVPERPVTCMVPETPT
MSAEPTVLASEPPVLSETAETFESMRASGHV-A-SEVSTSLLEPAV-----------TSP
VLAESILE----------------PPAM--AVPESSAVAVLESSAVTVLESSTVT-----
------------------VLESSTVTVLEPSVVTVPEPP----------VVAEPDYITIP
VPVVSAL------EPSVPVLEPAVSVLQPSMIVSE---------PSVSVQESTVTVSEPA
VTVSEQTQVIPTEVAIESTPMILESSIMS-SHVMKGINLPSG-DQNLAPEIGMQEIPLHS
GEE-PHAEVHLKSDSYESEHGINIDLNINNHLIAKEMEHNTVCAASTSPVGEIGEEKILP
TSETKQCTVLVTYPGVSEADAGET-LSSTGPL-ALEPDA----TGTSKGIEFITTSTLSS
VNKYDIDVSLTTQDTEHDMVISTSPSGGSEADIEGPLPAKDIHLDLPSNNNLVSK-DTEE
PLPVKESDQTLAALLSPKES-SGGEKEVPPPPKETLSDSGFSANIEDINEADLVRPLLPK
DMERLTSLRAG-------IEGPLLASDVGRDKS-AASPVVSSMPERASE-SSSEEKDDYE
IFVKVKDTHEKSKKNKNRDKGEKEKK----------------------------------
-----------------------------------------------RDSSLRSRSKRSK
SSEHKSRK-------RTSESRSRARKRSSKSKS-HRSQTRSRSRSRRRRRSSRSRSKSRG
RRSVSKEKRKRSPKHRSKSRERKRKRSS-SRDNRKTVRARSRTPSRRSRSHTPSRRRRSR
SVGRRRSFSISPSRRSRT-----PSRRSRTPSRRSRTPSRRSRTP--------------S
RRSRT--PSRRSRTPSRRRRSRSV--VRRRSFSISPVR-LRRSRT-PLRRRFSRSPIRRK
RSRSSE---RGRS-PKRLTDLDKAQLLEIAKANAAAMCAKAGVPLPPNLKPA-PP-PTIE
EKVAKKSGGATIEELTEKCKQIAQSKEDDDVIVNKPHVSDEE-EEEPPFYHHPFKLSEPK
PIFFNLNIAA---AKPTPPKSQVTLTKEFPVSSGSQHRKKEADSVYGEWVPVEKNGEENK
D-DDNVFSSNLPSEPVDISTAMSERALAQKRLSENAFDLEAM-SMLNRAQERIDAWAQLN
SIPGQFTGSTGVQVLTQEQLANTGAQAWIKKDQFLRAAPV
PF01585 // ENSGT00730000111141_0_Domain_1 (2621, 2664)
TGGMGAVLMRKMGWREGEGL
GKNKEGNKEPILVDFKTDRKGLVENSGT00730000111141_0_Linker_1_2 (2664, 2688)
AVGERAQKRSGNFSAAMKDLSGKHPF00035 // ENSGT00730000111141_0_Domain_2 (2688, 2755)
PVSALMEICNKRR
WQPPEFLLVHDSGPDHRKHFLFRVLINGSAYQPSFASPNK-KHAKATAATVVLQAMGLVP
KDLMANATCFRSASRR
ENSPEMP00000028357 (view gene)
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
--------------------------------------------------------MAAD
IEQVFRSFVVSKFREIQQ-----ELSSG--------------RSE-----------GQLN
GETNA--PIEGSQAGDTAAS----------------------------TRSLPNEEIVQK
IEEVLS---GVLD-TELRYKPDKAQLLEIAKANAAAMCAKAGVPLPPNLKPA-PP-PTIE
EKVAKKSGGATIEELTEKCKQIAQSKEDDDVIVNKPHVSDEE-EEEPPFYHHPFKLSEPK
PIFFNLNIAA---AKPTPPKSQVTLTKEFPVSSGSQHRKKEADSVYGEWVPVEKNGEENK
D-DDNVFSSSLPSEPVDISTAMSERALAQKRLSENAFDLEAM-SMLNRAQERIDAWAQLN
SIPGQFTGSTGVQVLTQEQLANTGAQAWIKKDQFLRAAPV
PF01585 // ENSGT00730000111141_0_Domain_1 (2621, 2664)
TGGMGAVLMRKMGWREGEGL
GKNKEGNKEPILVDFKTDRKGLVENSGT00730000111141_0_Linker_1_2 (2664, 2687)
AVGERAQKRSGNFSAAMKDLSGKPF14709 // ENSGT00730000111141_0_Domain_2 (2687, 2732)
HPVSALMEICNKRR
WQPPEFLLVHDSGPDHRKHFLFRVLRNGSPYQPNCMFFLN-RY-----------------
----------------
ENSRNOP00000002769 (view gene)
MGP_CAROLIEiJ_P0042291 (view gene)
ENSOGAP00000016216 (view gene)
MATNIEQIFRSFVVSKFREIQQELSSGRNEGQLNGETNTPIEGNQAGDA-AASARSLPNE
EIVQKIEEVLSGVLDTELRCKPDMKEAS-RKSRCVSVQTDPTD-EIPTKKSKKHKKHKNK
KKKKK----KEKEKKYKRQPEESESKLKSHHDGNIDLESDSFLKVDSESPAMALEHPMRP
FVPSE------------------------TSESP-AVVLEPPVVSMQVSEPQILETLKSA
AKTTELSV----ASTS---VISEQSVTVMLEPSVTKILDSSATAPVSTT---TVVLKS-S
EPVVTM-SEYQMKSVLKSLESTSPEPSKIML-VEPPVAKLLEPSETL-VVSSETPTEVYP
EPSTSTTMDFPESSATEAQRLPEQPVEVPPEIADSSKTRPQELLELPKTTALELPESS-V
ASAMELPGPPATSMLELQGPSVTPVLELPGPSAT-----------PVSELPGPLSTPVPE
LPGPPATA--VPELLGPSVTPVQQLSQELPGLPAPSMGLEPPQEVPEPPVMAQ----ELP
GLPMVTATVELPEQPAVTVAMELTEQPVTTTELEQPVGMTTVEHPGHPEVTTATELLGQP
EATMVLELPGQPVATTALELPGQPSVTGVPELPGLPSATRALELSGQPVATGALEL-PGQ
LMATGA-LEFSGQSG-AAGALELL-GQPLATGVLELPG-QPGAPELPGQPVATVALEISV
-QSVVTTSELSTMTVSQSLEVPS-TTALE--SYNTVAQELPTTLVGETSVTVGVDPLMAQ
ESHMLASNTMETHMLASNTLDSQMLASNTMDSQMLASNTMDSQMLASSTMDSQMLATSSM
DSQMLATSSMDSQMLATSTMDSQMLATSSMDSQ------------MLATSSMDSQMLATS
SMDSQMLATSSMDSQMLATST--MDSQMLASSTMDSQMLASSSMDSQMLA--SGTMDSQM
LASGTMDAQMLASGTMDAQMLASSTQD-SMLGSKSPDPYRLAQDPYRLAQDPYRLGHDPY
RLGHDAYRLGQDPYRLGHDPYRLTPDPYRMSPRPYRIAPRSYRIAPRPYRLAPRPLMLAS
RRSMMMSYAAERSMMSSY--ERSMMSYERSMMSPMAERSMMSAYERSMMSAYERSMMSPM
AERSMMSAYERSMMSAYERSMMSPMADRSMMSMGADRSMMSSYSAADRS-MMSSYSAADR
SMMSSYTADRSMMSMAG-DYTDSYTDTYTEAYMVPPLPPEEPPTMPPLPPEEPPMTPPLP
PEEPPEGPALPTEQSALTAENTWPTEVPSLPPEESVSQPEPPVSQSEISESSALPANYSV
SATDPSVLA-SEAAVTVPEPL-LDPESSVTSTPVESTVIAAEHEIIPERPMTCVVSETPI
MSAEPSMLTSEPPLTSETAETFDSMRASGHV-A-SEVSMSLLEPAV-----------TIP
EAAESSILES---------------SAM--AVP--------ESPAMVVPEPPSALAFPEP
PAVAIPE----------------------ATVMAVPEPP----------VVAEPEHVTIP
VPVVPAL------EPTVPILEPVVSALQPAIIISE---------PPVSTQESSVTISESA
VNVSEQTQVISTEMVIESAPLILESSAMSSAHVMKGMNLLPD-DQNLDPEVGMQEMPLHS
GDE--PHEGYQKSDYYESEHGINTDVNINDHLIAKEMEHNTVFAVSTGPVGEIGEEKILP
TDETKQCPVLDTYSGVSEADAGET-LSSTDPL-ALELDA----VGTSKGIEFITASTLSS
VNKYDVEVSLATQDSEHDMVISNSPSGGSEADIEGPLPAKDIHLDLPSSNNLVSA---EE
PLPVKESDQTSAVALSPKET-SGVEKEAPFPPIEI-ADSGLSANIEAINEADLVRPLLPK
DMERLTSLRAG-------IEGPLLASEVERDKS-AASPVIISLPERASE-SSSEEKDDYE
IFVKVKDTHEKSKKNKNRDKGEKEKK----------------------------------
-----------------------------------------------RDSSLRSRSKRSK
SSEHKSRK-------RTSESRSRARKRSSKSKS-HRSQTRSRSRSRRRRRSSRSRSKSRG
RRSVSKEKRKRSPKHRSKSRERKRKRSS-SRDNRKTVRARSRTPSRRSRSHTPSRRRRSR
SVGRRRSXXXXXXXXXXXXXXXXXXXXXXTPSRR---------------------SRTPS
RRSRT--PSRRSRTPSRRRRSRSG--ARRRSFSISPVR-LRRSRT-PLRRRFSRSPIRRK
RSRSSE---RGRS-PKRLTDLDKAQLLEIAKANAAAMCAKAGVPLPPNLKPA-PP-PTIE
EKVAKKSGGATIEELTEKCKQIAQSKEDDDVIVNKPHVSDEE-EEEPPFYHHPFKLNEPK
PIFFNLNIAA---AKPTPPKSQVTLTKEFPVSSGSQHRKKEADSVYGEWVPVEKNGEENK
D-DDNVFSSNLPSEPVDISTAMSERALAQKRLSENAFDLEAM-SMLNRAQERIDAWAQLN
SIPGQFTGSTGVQVLTQEQLANTGAQAWIKKDQFLRAAPV
PF01585 // ENSGT00730000111141_0_Domain_1 (2621, 2660)
TGGMGAVLMRKMGWREGEGL
GKNKEGNKEPILVDFKTDRK----------------------------------------
------------------------------------------------------------
----------------
ENSCGRP00001015682 (view gene)
ENSDNOP00000022750 (view gene)
MATNIEQIFRSFVVSKFREIQQELSSGRSEGQLNGETNTPIEGNQAGDA-AASARNLPNE
EIVQKIEEVLSGVLDTELRYKPDLKEAS-RKSKCVSVQTDPTD-EIPTKKSKKHKKHKNK
KKKKK----KEKEKKYKKQPEDSESKPKSHHDGNIDLESDSFLKFDSEPSAMALEHPVRA
FDLPE------------------------ISESP-AFMLEPPAIAMVVSEPHIFETLNPA
TKTADLSV----ASTSVISEHSDESVAVTLEPSMTKILDSFPTAPVPAT---AAVLKS-S
EPIVT-SVDYQMKAVLKPLESTSPETSK-ML-VEPPVAKVLEPSETF-VVSSETSTEVHP
EPSTSTAVGFPESSVTEVLRLPEQPVEVPPEIADSSMTRPQELLELPQTTALEVPESS-V
ASAMELPGPPSTSMLELQGPPVTPVLELPGPSAT-----------LVPELPGAPSTPVPE
LPGPPATA--VPELPGPSVTPVPQLPQELPGLPSPSMGLEPPQEVPEPPVMAQ----ELP
VLPVVTAAVELPGQPAVTVAMELTEQPVTTTELEQPLGMTAVEHPGQPEVTTASGLLGQP
EATMVLELPGQPVATTALELPGQPSVTGVLELPGLPSATRALELSGQPVATGALEL-PGQ
LMAAGA-LEYSGQSG-AAGALELL-GQPLATGVLELPG-QPGAPELPGQPVATVALEISV
-QSVVTTSELSTMTVSQSLEVPS-TTALE--SYNTVAQELPTTLVGETSVTVGVDPLMAQ
ESHMLASNTMETHMLASNTMDSQMLASNTMDSQMLASNTMDSQMLASSTMDSQMLASSTM
DSQMLASSSMDSQMLATSTMDSQMLATSSMDSQMLATSS--MDSQMLA----------TS
SMDSQMLATSSMDSQMLATST--MDSQMLATSTMDSQMLATSSMDSQMLA--SGTMDSQM
LASGTMDAQMLASGTMDAQMLASSSQDSAMLGSKSPDPYRLAQDPYRLAQDAYRLGHDPY
RLGHDAYRLGQDPYRLGHDPYRLTPDPYRMSPRPYRIAPRSYRIAPRPYRLAPRPLMLAS
RRSMMMSYAAERSMMSSY--ERSMMSYERSMMSPMAERSMMSAYERSMMSAYERSMMSPM
AERSMMSAYERSMMSAYERSMMSPMADRSMMSMGADRSMMSSYSAADRS-MMSSYSAADR
SMMSSYTADRSMMSMAADSYTDSYTDTYTEAYMVPPLPPEEPPTMPPLPPEEPPMTPPLP
PEEPPEGPALPTEQSALATENTWPTEVPALPPEESVTLPEPPVSQSESSEPSVVPANYSV
STSEPSMLV-SEATVAVPEPP-LEPESSVILA--ESAVVPEEREIVLET----PMSETPI
MSAETTMLPSDPPIMSEMAEPLDSMQASGQA-V-SQLTISLQEPAV-----------AIP
EPVQSTVEQPATAIS-------E-----------PPVVVALEPPALAVPEPPAVAALVPP
-VVAVSE-------------------SPEPPAGAVPEIPAVALLEPR--AMAESER--VI
VPVVSALE-----PPTLHILEPAVSVLQPAMIVS-E--------PSVSVQESTVTISEPA
VTVSKQIQIISTEMVLESTPMILESSVMATSHIIRGVNLISG-HPNLAPEVSMQETPMHL
REK--SHDGHLKSDSYESECGINIDLNINNHLIAKEIEHNTVSAASTGVFGETGEEKSLP
VSETEQCTVLVTCSGVSGTDVEGT-LSSTGPF-ALESDG----KGT---IELTTAPAFSS
VNKYDVEVSLATQDTEHDMVISTSPSGGSEADIEGPLPAKDIHLDLPSNNNLISR-DAEG
PLPIRESDQAMAVL-SPKE--SGEDKEVPPPTEEVRPDSGFPTNIEDINEADLVRPLLPK
DMERLANLRAG-------IEGPLLPSEVERDKS-AASPVVVSIAERASE-SSSEEKDDYE
IFVKVKDTHEKSKKNKNRDKGEKEKK----------------------------------
-----------------------------------------------RDSSLRSRSKRSK
SSEHK-RK-------RTSESRSRARKRSSKSKS-HRSQTRSRSRSRRRRRSSRSRSKSRG
RRSVSKEKRKRSPKHRSKSRERKRKRSS-SRDNRKTVRARSRTPSRRSRSHTPSRRRRSR
SVGRRRSFSISPSRRSRT-----PSRRSRTPSRRSRTPSRRSRTPSRRSRTPSRRSRTPS
RRSRT--PSRRSRTPSRRRRSRSV--VRRRSFSISPVR-LRRSRT-PLRRRFSRSPIRRK
RSRSSE---RGRS-PKRLTDLDKAQLLEIAKANAAAMCAKAGVPLPPNLKPA-PP-PSIE
EKVAKKSGGATIEELTEKCKQIAQSKEDDDVIVNKPHVSDEE-DEEPPFYHHPFKLSEPK
PIFFNLNIAA---AKPTPPKSQVTLTKEFPVSSGSQHRKKEADSVYGEWVPVEKNGEENK
D-DDNVFSSNLPSEPVDISTAMSERALAQKRLSENAFDLEAM-SMLNRAQERIDAWAQLN
SIPGQFTGSTGVQVLTQEQLANTGAQAWIKKDQFLRAAPV
PF01585 // ENSGT00730000111141_0_Domain_1 (2621, 2664)
TGGMGAVLMRKMGWREGEGL
GKNKEGNKEPILVDFKTDRKGLVENSGT00730000111141_0_Linker_1_2 (2664, 2688)
AVGERAQKRSGNFSAAMKDLSGKHPF00035 // ENSGT00730000111141_0_Domain_2 (2688, 2755)
PVSALMEICNKRR
WQPPEFLLVHDSGPDHRKHFLFRVLINGSAYQPSFASPNK-KHAKATAATVVLQAMGLVP
KDLMANATCFRSASRR
ENSCHIP00000025768 (view gene)
ENSTGUP00000000498 (view gene)
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
----------------------------------VVSKF---------------------
------------------------------------------------------------
--------REIQEEQV---SANVE--GQPNGD-TIPAEQAGPSEASGA-AGSCQS-DQIV
--QKIEEVLSGALGTELQCNPDKAQLLEIAKANAAAMCAKSGVPLPPSLMPLLSQ-KK-D
DKANQKSSRDTLKELTEKCKKIAQST--DDVIVNKPHVSDEE-EEERPFYNHPFKLSEPK
PIFFNLSTPSIKPAPPPQPKNQVSLSKEFPVSSGSQHRKKEADSVYGEWVPVDKNGKDDG
--KDDVFPKP-AIECVDITTAMNDRAVAQKRLNENSFDLEAM-CMLNRAQEQIDAWAQSN
SIPGQFTGSTGAQILSSDELTNSGPQAWIRKDQFLRAAPV
PF01585 // ENSGT00730000111141_0_Domain_1 (2621, 2664)
TGGMGAQLMRKMGWREGEGL
GKNKEGSVEPIMVDFKTDRKGLVENSGT00730000111141_0_Linker_1_2 (2664, 2688)
AVGEKXXRWVGPVP-GKFFLPGNHPF00035 // ENSGT00730000111141_0_Domain_2 (2688, 2755)
PVSALMEICNKRR
WTPPEFVV-EDSGPDHRKHFIFKVRVNGNEYRPTFASLNK-KHAKATAATAALQAMGLVP
KESMVNTTMFRSASHR
ENSANAP00000035842 (view gene)
ENSODEP00000011614 (view gene)
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
-------------------------------------------M
PF01585 // ENSGT00730000111141_0_Domain_1 (2625, 2651)
GAVLMRKMGWREGEGL
GKNKEGNKEPENSGT00730000111141_0_Linker_1_2 (2651, 2687)
IL-------------------RSGNFSAAMKDLSGKPF14709 // ENSGT00730000111141_0_Domain_2 (2687, 2731)
HPVSALMEICNKRR
WQPPEFLLVHDSGPDHRKHFLFRVLINGSAYQPNRLRFYINHFR----------------
----------------
ENSPVAP00000010544 (view gene)
MATNIEQIFRSFVVSKFREIQQELSSGRSEGQLNGETNTPIEGNQAGDA-AASARSLPNE
EIVQKIEEVLSGVLDTELRYKPDLKEAS-RKSRCVSVQTDPTD-EIPTKKSKKHKKHKNK
KKKKK----KEKEKKYKRQPEDAESKLKSHHDGNMDLESDSFLKFDSEPSAMAPEHPIRA
FGLSE------------------------TSESP-AVMLEPPIVSMEVSQPHTFESPKPA
TKTAELSV----ASTSVIAVQSEQSVAVTLEPSTTKILDSFATAPMPTT---TVVLQS-S
EPVVTMS--NQMKSVLKSLENTPPEPSKIV--LEPPVAKVLEPSETL-VITSETSTEVHP
EPSTSTTMDFPESSATEVLRLPEQPVEVPSEIADSSMTRSQELLELPKTTALELPESP-V
ASVMELPGPPATSMPELQGPPVTPVLELPGPSAT-----------PVPELPGPVSTPVPE
LLGPPATA--VSELPGPSVTSVPQLSQELPGLSAPSMGLEPPQEVPEPPVMAQ----ELP
GLPVVTAAVELPGQPAVTVAMELTEQPVTTTELEQPVGMTTXXXXXXXXXXXXXXXXXXX
XXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXX-XXX
XXXXXX-XXXXXXXX-XXXXXXXX-XXXLST--GLLPG-QPGAPELPGQPVA-VALEISV
-QSVMTTTELSTMTVSQSLEVPS-TTALE--SYNTVAQELPTTLVGETSITVGVDPLMAQ
ESHMLASNTMETHMLASNTMDSQMLASNTMDSQMLASNTMDSQMLATSSMDSQMLATSSM
DSQMLATSSMDSQMLATSSMDSQMLATSSMDSQMLATSS--MDSQMLATSSMDSQM----
------LATSSMDSQMLATSS--MDSQMLATSTMDSQMLATSTMDSQMLA--SGTMDSQM
LASGTMDAQMLASGTMDAQMLASSTQDSAMLGSKSPDPYRLAQDPYRLAQDPYRLGHDPY
RLGHDAYRLGQDPYRLGHDPYRLTPDPYRMSPRPYRIAPRSYRIAPRPYRLAPRPLMLAS
RRSMMMSYAAERSMMSSY--ERSMMSYERSMMSPMAERSMMSAYERSMMSAYERSMMSPM
AERSMMSAYERSMMSAYERSMMSPMADRSMMSMGADRSMMSSYSTADRS-MMSSYSAADR
SMMSSYTADRSMMSMTADSYTDSYTDTYTEAYMVPPLPPEEPPTMPPLPPEEPPMTPPLP
PEEPPEGPALPTEQSALTAENAWPTEVPALPPEESVSLPEPPVSQSEISEHSVVPANYSV
SASEPSLLA-SEAAMTVPEPP-LEPESSVTSTPTESAAIAEEHEIVPERPVTYMVSETPV
MSAEPTMLTSEPSVMSETAETFDSMRASEHV-P-SEVSMSLLDPTV-----------TIP
EPSQSTL---------------E-LSAM--AVSELPAVSVPELPAEAVVEPPAEA-VPEP
PVLAVPE----------------------PLAVTVPEPL----------ALVEPEPVTIP
VPVVSAV------DSAVPVLEPAVSVFQ-PNMIVSE-------P-SVSVQESTVSVLEPA
VTVSEQTQVISTEMVLESTPVMLESG------VVKGMNLLSG-DQNLAPEIGMQEFPLHS
DEE-PHTEGYLKNDTYESEHGVNIDHNVNNHLIAKEMEHSTVSAASTGAVGEIGGEKSLP
VNETKQCTVLDTCPSVSEADVGGT-LSSTGPF-ALESDT----VGTSKGIEFATASALSS
ISKYD-EVCLTTQDTEHDMVISSSPSGGSEADIEGPLPAKDIHLDLPSN-NFISN-DAEG
PLPIKESDQTLAGALSPKEN-S-EDKEVPVSTKEILSDTGFSANIDDINEADLVRPLLPK
DMERLTSLRAG-------IEGPLLASEVERDKS-AASPIVVSIPERASE-SSSEEKDDYE
IFVKVKDTHEKSKKNKNRDKGEKEKK----------------------------------
-----------------------------------------------RDSSLRSRSKRSK
SSEHKSRK-------RTSESRSRARKRSSKSKS-HRSQTRSRSRSRRRRRSSRSRSKSRG
RRSVSKEKRKRSPKHRSKSRERKRKRSS-SRDNRKTVRARSRTPSRRSRSHTPSRRRRSR
SGGR-RSFSISPSRRSRT-----PSRRSRTPSRRSRTPSRRS--------------RTPS
RRSRT--PSRRSRTPSRRRRSRSV--VRRRSFSISPVR-LRRSRT-PLRRRFSRSPLRRK
RSRSSE---RGRS-PKRLTDLNKAQLLEIAKANAAAMCAKAGVPLPPNLKPA-PP-PSIE
EKVAKKSGGATIEELTEKCKQIAQSKEDDDVIVNKPHVSDEE-EEEPPFYHHPFKLSEPK
PIFFNLNIAA---AKPTPPKSQVTLTKEFPVSSGSQHRKKEADSVYGEWVPVEKNGEENK
D-DDNVFSSNLPSEPVDISTAMSERALAQKRLSENAFDLEAM-SMLNRAQERIDAWAQLN
SIPGQFTGSTGVQVLTQEQLANTGAQAWIKKDQFLRAAPV
PF01585 // ENSGT00730000111141_0_Domain_1 (2621, 2665)
TGGMGAVLMRKMGWREGEGL
GKNKEGNKEPILVDFKTDRKGLVAENSGT00730000111141_0_Linker_1_2 (2665, 2688)
VGERAQKRSGNFSAAMKDLSGKHPF00035 // ENSGT00730000111141_0_Domain_2 (2688, 2724)
PVSALMEICNKRR
WQPPEFLLVHDSGPDHRKHFLFRVLRNGA-LTPNCMFFLN-RY-----------------
----------------
ENSTTRP00000004433 (view gene)
---------------------------RSEGQLNGETNTPIEGNQAGDA-AASARSLPNE
EIVQKIEEVLSGVLDTELRYKPDLKEAS-RKSRCVSVQTDPTD-EIPTKKSKIHKKHKNK
KKKKK----KEKEKKYKRQPEESESKPKSHHDGNVDLESDSFLKFDS--SSMALEHPVRA
FGLSD------------------------TSESP-ALVLEPPVVSMEVLEPHTLEILKPA
TKTAELSV----ASTSVVSVQSEQSVAVMLEPSTTKILDSFATAPVPTT---TVVLKS-S
EPVVTMSVECQVKPGLKSLESTPPEPSKIML-LEPPVAKVLEPSETL--VSSETPTQVHP
EPSTSTTMDFPESSATEVLRLPEQPVEVPSEIADSSMTRPQEMLELPKTTALELQESS-V
ASAMELPGPPATSMLELQGPPVTPVLELPGPSAT-----------PVPELPGPLSTPVPE
LLGPPATA--VPELPGPSVTSVPQLSQELPGLPAPSMGLEPPQEVPEPPVMAQ----ELP
GLPVVTAAVELPGQPVVTVAMETE-QPVXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXX
XXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXL-PGQ
LMAAGA-LEFSGQSG-AAGALELL-GQPLATGVLELPG-Q-GAPELPGQPVATVALEISV
-QSVVTTTELSTMTVSQSLEVPS-TTALE--SYNTVAQELPTTLVGETSVTVGVDPLMAQ
ESHMLASNTMETHMLASNTMDSQMLASNTMDSQMLASNTMDSQMLASSTMDSQMLATSSM
DSQMLATSSMDSQMLATSSMDSQMLATSSMDSQM------------LATSSMDSQMLATS
SMDSQMLATSSMDSQMLATST--MDSQMLATSSMDSQMLATSSMDSQMLA--SGTMDSQM
LASGTMDAQMLASGTMDAQMLASSTQDSAMLGSKSPDPYRLAQDPYRLAQDPYRLGHDPY
RLGHDAYRLGQDPYRLGHDPYRLTPDPYRMSPRPYRIAPRSYRIAPRPYRLAPRPLMLAS
RRSMMMSYAAERSMMSSY--ERSMMSYERSMMSPMAERSMMSAYERSMMSAYERSMMSPM
AERSMMSAYERSMMSAYERSMMSPMADRSMMSMGADRSMMSSYSAADRS-MMSSYSAADR
SMMSSYTADRSMMSMAADSYTDSYTDTYTEAYMVPPLPPEEPPTMPPLPPEEPPMTPPLP
PEEPPEGPALPTEQSALTAENTWPTEVPALPPEESVSLPEPPVSQSEISEPSAVLANYSV
SASEPSMLA-SEASMTVPEPP-LEPESLVTSTPVESAAVAEEHEIIPERPVTYMVSETPT
TSGEPTVLTSEPSVMSEAAETFDSMRASGHV-A-SEVSMSLLEAAV-----------PIP
EPSQSTVDLPAMAVS-------E-LPAV--AVPEQPAVAVPEQPAGTVSEPP--------
-------------------------------AVAVQDPP--AEAVTEPLALSEPEHITVP
VPVVSAL------EPTVPVLEPAVSVFQPNVIVSE---------PSICVQESTVTISEPP
VTLSEQTQVIPTEMVLESTPMILESN------VIKGMNLLSG-DQNLAPEINMQEIPMHS
DEE-PHAEGHLKNDSYETESGMNIDLNINNHLIAKEMEHNTVSAASTGAVGETGEEKILP
TNETKQCTVLDTCPTVSETDVGGT-LSSTGSL-ALEPDA----LGTGKGLEFATASAFSS
ISKYDVEVSLTTQDTEHDMVISTSPSGGSEADIEGPLPAKDIHPDIPANNNFVSK-DAEG
SLPIQESDQTLAVALSPKES-SGEDKEVPLPAKEILSDSGFSANIDDINEADLVRPLLPK
DMERLTSLRAG-------IEGPLLASEVERDKS-AASPVVISIPERASE-SSSEEKDDYE
IFVKVKDTHEKSKKNKNRDKGEKEKK----------------------------------
-----------------------------------------------RDSSLRSRSKRSK
SSEHKSRK-------RTSESRSRARKRSSKSKS-HRSQTRSRSRSRRRRRSSRSRSKSRG
RRSVSKEKRKRSPKHRSKSRERKRKRSS-SRDNRKTGRARSRTPSRRSRSHTPSRRRRSR
SGGR-RSFSISPSRRSRT-----PSRRSRTP----------------------------S
RRSRT--PSRRSRTPSRRRRSRSV--VRRRSFSISPVR-LRRSRT-PLRRRFSRSPIRRK
RSRSSE---RGRS-PKRLTDLNKAQLLEIAKANAAAMCAKAGVPLPPNLKPA-PP-PSIE
EKVAKKSGGATIEELTEKCKQIAQSKEDDDVIVNKPHVSDEE---RPPFYHH-FKLSEPK
PIFL---IAA---AKPTPPKSQVTLTKEFPVSSGSQHRKKEADSVYGEWVPVEKNGEENK
D-DDNVFSSSLPSEPVDISTAMSERALAQKRLSENAFDLEAM-SMLNRAQERIDAWAQLN
SIPGQFTGSTGVQVLTQEQLANTGAQAWIKKDQFLRAAPV
PF01585 // ENSGT00730000111141_0_Domain_1 (2621, 2665)
TGGMGAVLMRKMGWKEGEGL
GKNKEGNKEPILVDFKTDRKGLVAVGERAQKRSGNSAAM---DSGKHPVS-LMEICNKRR
WQPPEFLLVHDSGPDHRKHFLFRVLRNGA-LTPNCMFFLN-RY-----------------
----------------
ENSMOCP00000012900 (view gene)
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
--------------------------------------------------------MAAD
IEQVFRSFVVSKFREIQQ-----ELSSG--------------RNE-----------GQLN
GETNA--PIEGSQTGDPAAS----------------------------ARSLPNEEIVQK
IEEVLS---GVLD-TELRYKTDKAQLLEIAKANAAAMCAKAGVPLPPNLKPA-PP-PTIE
EKVAKKSGGATIEELTEKCKQIAQSKEDDDVIVNKPHVSDEE-EEEPPFYHHPFKLSEPK
PIFFNLNIAA---AKPTPPKSQVTLTKEFPVSSGSQHRKKEADSVYGEWVPVEKNGEENK
D-DDNVFSSSLPSEPVDISTAMSERALAQKRLSENAFDLEAM-SMLNRAQERIDAWAQLN
SIPGQFTGSTGVQVLTQEQLANTGAQAWIKKDQFLRAAPV
PF01585 // ENSGT00730000111141_0_Domain_1 (2621, 2664)
TGGMGAVLMRKMGWREGEGL
GKNKEGNKEPILVDFKTDRKGLVENSGT00730000111141_0_Linker_1_2 (2664, 2687)
AVGERAQKRSGNFSAAMKDLSGKPF14709 // ENSGT00730000111141_0_Domain_2 (2687, 2732)
HPVSALMEICNKRR
WQPPEFLLVHDSGPDHRKHFLFRVLRNGSPYQPNCMFFLN-RY-----------------
----------------
ENSCANP00000007967 (view gene)
ENSODEP00000005985 (view gene)
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
-------------------------------------------M
PF01585 // ENSGT00730000111141_0_Domain_1 (2625, 2664)
GAVLMRKMGWREGEGL
GKNKEGNKEPTLVDFKTDQKGLVENSGT00730000111141_0_Linker_1_2 (2664, 2687)
AVGERAQKRSGNFSAAMKDLSGKPF14709 // ENSGT00730000111141_0_Domain_2 (2687, 2739)
HPVSALMEICNKRR
WQPPEFLLLHDSVPDHRKHFLFRVLINGSAYQPSFCKIFFQEYIY---------------
----------------
ENSPPAP00000025487 (view gene)
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
-----------------------------------------MPGLTVLMRKM--------
----EGNKEPILVDFKTDRKGLVAVGERARKRSGNFSAAMKDLPGK
PF14709 // ENSGT00730000111141_0_Domain_2 (2687, 2732)
HPVSALMEICNKRR
WQPPEFLLVHDSGPDHRKRFLFRVLRSGSPYQPSCMFFLN-RD-----------------
----------------
ENSRROP00000037529 (view gene)
ENSCPOP00000032872 (view gene)
MAADIEQVFRSFVVSKFREIQQELSSGRSEGQLNGETNTPIEGNQAGD-AASSARSLPNE
EIVQKIEEVLSGVLDTELRYKPDLKEAS-RKSRCVSVQTDPTD-EIPSKKSKKHKKHKNK
KKKKK----KEKEKKYKRQPEESESKPKSHHDGNIDLESDSFLKFDSEPSAMALEHPIRA
FDLSE------------------------TSESS-AVVLEPPI-SMEIPDPHIPETLKPA
TKTAELSV----GSTSVISE---QSMAVMLEPSVTKIVDSFATTLVPTTVPATVVVKS-P
EPVATMSVEYQMKSALKSSEHTSLEPSHIML-AEPPVAKELEPSETH-VIALETSTEVYP
EPSISTTKDFPESSATEVLRLPEHSI--PSEIADPAMTRPQDLLELSKTSALELPVSS-V
ASALELPGPPATSMPELQGPPVTPMLELPGPSA-----------TPVSELSGPLSTPVPE
LPGPPATA--VPELPGPSVTPVPQLSQELPGPLAPSVGLEPPQEVPEPPVMAQ----ELP
GLPAVSATVELPGQPAITVAMELTEQPVTTTEFEQPVGMTAVEHPGHSEVTAATGLLGQP
EAAMVLELPGQPVATTALELSGQPSVTGVPELPGLSSATRALELSGQSV-AGALEL-PGQ
LMAAGA-LEFSGQSG-AAGALELL-GQPLATGVLELPG-QPGAPELPGQPVATVALEISV
-QSVVTTSELSTMTVSQSLEVPS-TTALE--SYNTVAQELPTTLVGETSVTVGVDPLMAQ
ESRMLASNTMETHMLASNTMDSQMLASNTMDSQMLASNSMDSQMLASSTMDSQMLASSTM
DSQMLATSSMDSQMLATSSMDSQMLATSSMDSQML------------ATSSMDSQMLATS
SMDSQMLATSSMDSQMLATSS--MDSQMLATSSMDSQMLATSSMDSQMLA--SGTMDSQM
LASGTMDAQMLASGTMDAQMLASSTQDSAMLDSKSPDPYRLAQDPYRLAQDAYRLGHDPY
RLGHDAYRLGQDPYRLGHDPYRLTPDPYRMSPRPYRIAPRSYRIAPRPYRLAPRPLMLAS
RRSMMMSYAAERSMMSSY--ERSMMSYERSMMSPMAERSMMSAYERSMMSAYERSMMSPM
AERSMMSAYERSMMSAYERSMMSPMADRSMMSMGADRSMMSSYSAADRS-MMSSYSAADR
SMMSSYTADRSMMSMAADSYTDSYTDTYTEAYMVPPLPPEEPPTMPPLPPEEPPMTPPLP
PEEPPEGSALPTEQSALTAENTWATEMPALPPEENVSHPEPPVGQSEISEPSAVPTNYSV
SGSESSMLV-SESVVTIPEPP-LEPESSVTSTPVESAVVA-EHEIVTERPVACVVSETAV
-MAEATLLTSEPPIVTETIETSESMKFER--HVASEASVSLLEAAV-----------TVP
EPAENIQELSSVAVPESPAMAVPESPAVAGLVPESPAVAVPQSPAVAFPESSTVAAPEP-
------------------------------PAVRKPEPPAV--AVLELPSVAEPEHIPVP
VSAVSAL------EPMVPVLEPAVTMLQSAVIVSE---------PCVSVQQYTVTVSEPA
VPVSEQTQVIPTEMALESAAVIAEPSVMS-SHVIRGMNSLYG-DTNLAPEICMQET-PLH
SGEEPHAEGHLKSDSYESEHGMNVDLNINNHFIAKTIES---------------------
------------------------------------------------------------
------------------------------------------------------------
---------------------------------------------SIINEADLVRPLLPK
DMERLTSLRAG-------IEGPLLPSEVERDKS-AASPVVISIPERASE-SSSEEKDDYE
IFVKVKDTHEKSKKNKNRDKGEKEKK----------------------------------
-----------------------------------------------RDSSLRSRSKRSK
SSEHKSRK-------RTSESRSRARKRSSKSKS-HRSQTRSRSRSRRRRRSSRSRSKSRG
RRSVSKEKRKRSPKHRSKSRERKRKRSS-SRDNRKTGRVRSRTPSRRSRSHTPSRRRRSR
SVGRRRSFSISPSRRSRT-----PSRRS--------------RTPSRRSRTPSRRS----
---RT--PSRRSRTPSRRRRSRSV--VRRRSFSISPVR-LRRSRT-PLRRRFSRSPIRRK
RSRSSE---R-----ERLTDLDKAQLLEIAKANAAAMCAKAGVPLPPNLKPA-PP-PTIE
EKVAKKSGGATIEELTEKCKQIAQSKEDDDVIVNKPHVSDEE-EEEPPFYHHPFKLSEPK
PIFFNLNIAA---AKPTPPKSQVTLTKEFPVSSGSQHRKKEADSVYGEWVPVEKNGEENK
D-DDNVFSSNLPSEPVDISTAMSERALAQKRLSENAFDLEAM-SMLNRAQERIDAWAQLN
SIPGQFTGSTGVQVLTQEQLANTGAQAWIKKDQFLRAAPV
PF01585 // ENSGT00730000111141_0_Domain_1 (2621, 2664)
TGGMGAVLMRKMGWREGEGL
GKNKEGNKEPILVDFKTDRKGLVENSGT00730000111141_0_Linker_1_2 (2664, 2687)
AVGERAQKRSGNFSAAMKDLSGKPF14709 // ENSGT00730000111141_0_Domain_2 (2687, 2756)
HPVSALMEICNKRR
WQPPEFLLVHDSGPDHRKHFLFRVLINGSAYQPSFASPNK-KHAKATAATVVLQAMGLVP
KDLMANATCFRSASRR
MGP_PahariEiJ_P0029650 (view gene)
ENSECAP00000015914 (view gene)
MATNIEQIFRSFVVSKFREIQQELSSGRSEGQLNGETNTPIEGNQAGDA-AASARSLPNE
EIVQKIEEVLSGVLDTELRYKPDLKEAS-RKSRCVSVQTDPTD-EIPSKKSKKHKKHKNK
KKKKK----KEKEKKYKRQPEESESKLKSHHDGNIDLESDSFLKFDSEPSAMALEHPVRA
FGLSE------------------------TSESS-AVVLEPPVVSMEVSEPHTLETLKPA
MKTAELSV----ASTSVISVQSEQSVAVTLEPSLTKIVDPFATAPVPTT---TVVLKT-S
EPVVTMSVEYQMKSVLKSLESTPPEPSKIMF-LEPPVAKVLEPSETL-AVSSETPTEVHP
EPSTSTTMDFPESSATEVLRLPEQPVEVPSEIADSSMTRPQELLELPKTTALELPESS-V
ASAMELPGPPATSMLELQGPPVTPVLELPGPSAT-----------PVPELPGPLSTPVPE
LLGPPATA--VPELPGPSVTSVPQLSQELPLPATILWGLEPPQEYPEPPVIGT---GSCQ
GLPAVTAAVELPGQPLPV-A-METEQPVTTTELEQPVGMTSMEHPGQPEVTTASGLLGQP
EAAMVLELPGQPVATTALELSGQPSVTGVPELPGLPSATRALELSGQPVATGALEL-PGQ
LMAAGA-LEFSGQSG-AAGALELL-GQPLATGVLELPG-QPGAPELPGQPVATVALEISV
-QSVVTTTELSTMTVSQSLEVPS-TTALE--SYNTVAQELPATLVGETSVTVGVDPLMAQ
ESHMLASNTMETHMLASNTMDSQMLASNTMDSQMLASNTMDSQMLASSTMDSQMLATSSM
DSQMLATSSMDSQMLATSSMDSQMLATSSMDSQMLATSS--MDSQMLATSSMDSQM----
------LATSSMDSQMLATST--MDSQMLATSTMDSQMLATSSMDSQMLA--SGTMDSQM
LASGTMDAQMLASGTMDAQMLASSSQDSAMLGSKSPDPYRLAQDPYRLAQDPYRLGHDPY
RLGHDAYRLGQDPYRLGHDPYRLTPDPYRMSPRPYRIAPRSYRIAPRPYRLAPRPLMLAS
RRSMMMSYAAERSMMSSY--ERSMMSYERSMMSPMAERSMMSAYERSMMSAYERSMMSPM
AERSMMSAYERSMMSAYERSMMSPMADRSMMSMGADRSMMSSYSAADRS-MMSSYSAADR
SMMSSYTADRSMMSMAADSYTDSYTDTYTEAYMVPPLPPEEPPTMPPLPPEEPPMTPPLP
PEEPPEGPALPTEQSTLTAENTWPTEVSALPPEESVSLPEPPVSQSEISEPSAVPANYSV
SASEPSVLA-SEATVTVPEPP-LEPESSVMSTPMESAVVAEEHEIVPERPVTYMVSETPV
MSAEPAVLTSEPSIMSETAETFDSVRASGHV-A-SEVSMSLLDPAV-----------TIP
EPSQSTL---------------E-LPAT--AVSELPAVAVPEPPAGTVPEPPAMA-VLEP
PAVAVPEP----------------------PADAVMEPL----------ALAETEHVTVP
VPVVSAL------EPTVPILETGVSVLQ-PNVIVSE-------P-SVSVQESAVTVSEPA
VTVSEQTQVIPAEMMLESTPMILESS------VIKGMNLLSG-DQNLAPEIGLQEIPMHS
DEE-PHAEGHLKNDSYESKHGINVDLNINNHLIAKEMEHNAVSAISTGAVGEIGEERILP
SSETKQCTVLDTCPIVSETDGGG-----TGPL-ALEPDA----LGTSKGIEFATASALSS
VSKYYVEVSLTTQDTEHDMVISSSPSGGSEPDIEGPLPAKDIHLDLPSNNNFISK-DAEG
PLPIKESDQTLAVALSPKES-SGEDKEVPLPTKEILSDSGFPANIDDINEADLVRPLLPK
DMERLTSLRAG-------IEGPLLASEVERDKS-AASPVVISIPERASE-SSSEEKDDYE
IVVKVKDTHEKSKKNKNRDKGEKEKK----------------------------------
-----------------------------------------------RDSSLRSRSKRSK
SSEHKSRK-------RTSESRSRARKRSSKSKS-HRSQTRSRSRSRRRRRSSRSRSKSRG
RRSVSKEKRKRSPKHRSKSRERKRKRSS-SRDNRKTVRARSRTPSRRSRSHTPSRRRRSR
SAGR-RSFSISPSRRSRT-----PSRRSRTPSRRSRTPSRRSR-------TPSRRSRTPS
RRSRT--PSRRSRTPSRRRRSRSV--VRRRSFSISPVR-LRRSRT-PLRRRFSRSPIRRK
RSRSSE---RGRS-PKRLTDLNKAQLLEIAKANAAAMCAKAGVPLPPNLKPA-PP-PTIE
EKVAKKSGGATIEELTEKCKQIAQSKEDDDVIVNKPHVSDEE-EEEPPFYHHPFKLSEPK
PIFFNLNIAA---AKPTPPKSQVTLTKEFPVSSGSQHRKKEADSVYGEWVPVEKNGEENK
D-DDNVFSSNLPSEPVDISTAMSERALAQKRLSENAFDLEAM-SMLNRAQERIDAWAQLN
SIPGQFTGSTGVQVLTQEQLANTGAQAWIKKDQFLRAAPV
PF01585 // ENSGT00730000111141_0_Domain_1 (2621, 2664)
TGGMGAVLMRKMGWREGEGL
GKNKEGNKEPILVDFKTDRKGLVAVGERAQKRSGNFSAAMKDLSGKHPVSALMEICNKRR
WQPPEFLLVHDSGPDHRKHFLFRVLRNGSPYQPNCMFFLN-RY-----------------
----------------
ENSMAUP00000002486 (view gene)
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
--------------------------------------------------------MAAD
IEQVFRSFVVSKFREIQQ-----ELSSG--------------RSE-----------GQLN
GETNA--PIEGSQAGDTAAS----------------------------ARSLPNEEIVQK
IEEVLS---GVLD-TELRYKPDKAQLLEIAKANAAAMCAKAGVPLPPNLKPA-PP-PTIE
EKVAKKSGGATIEELTEKCKQIAQSKEDDDVIVNKPHVSDEE-EEEPPFYHHPFKLSEPK
PIFFNLNIAA---AKPTPPKSQVTLTKEFPVSSGSQHRKKEADSVYGEWVPVEKNGEENK
D-DDNVFSSSLPSEPVDISTAMSERALAQKRLSENAFDLEAM-SMLNRAQERIDAWAQLN
SIPGQFTGSTGVQVLTQEQLANTGAQAWIKKDQFLRAAPV
PF01585 // ENSGT00730000111141_0_Domain_1 (2621, 2664)
TGGMGAVLMRKMGWREGEGL
GKNKEGNKEPILVDFKTDRKGLVENSGT00730000111141_0_Linker_1_2 (2664, 2687)
AVGERAQKRSGNFSAAMKDLSGKPF14709 // ENSGT00730000111141_0_Domain_2 (2687, 2732)
HPVSALMEICNKRR
WQPPEFLLVHDSGPDHRKHFLFRVLRNGSPYQPNCMFFLN-RY-----------------
----------------
ENSJJAP00000009701 (view gene)
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
--------------------------------------------------------MAAD
IEQVFRSFVVSKFREIQQ-----ELSSG--------------RSE-----------GQLN
GETNT--PIEGNQAGDAAAS----------------------------ARSLPNEEIVQK
IEEVLS---GVLD-TELRYKPDKAQLLEIAKANAAAMCAKAGVPLPPNLKPA-PP-PTIE
EKVAKKSGGATIEELTEKCKQIAQSKEDDDVIVNKPHVSDEE-EEEPPFYHHPFKLSEPK
PIFFNLNIAA---AKPTPPKSQVTLTKEFPVSSGSQHRKKEADSVYGEWVPVEKNGEENK
D-DDNVFSSSLPSEPVDISTAMSERALAQKRLSENAFDLEAM-SMLNRAQERIDAWAQLN
SIPGQFTGSTGVQVLTQEQLANTGAQAWIKKDQFLRAAPV
PF01585 // ENSGT00730000111141_0_Domain_1 (2621, 2664)
TGGMGAVLMRKMGWREGEGL
GKNKEGNKEPILVDFKTDRKGLVENSGT00730000111141_0_Linker_1_2 (2664, 2687)
AVGERAQKRSGNFSAAMKDLSGKPF14709 // ENSGT00730000111141_0_Domain_2 (2687, 2732)
HPVSALMEICNKRR
WQPPEFLLVHDSGPDHRKHFLFRVLRNGSPYQPNCMFFLN-RY-----------------
----------------
ENSCGRP00000020986 (view gene)
ENSMMUP00000007291 (view gene)
ENSMLUP00000002235 (view gene)
MATNIEQIFRSFVVSKFREIQQELSSGRSEGQLNGETNTPIEGNQAGDDAAASARSLPNE
EIVQKIEEVLSGVLDTELRCKPDLKEAS-RKSRCVSVQTDPTD-EVPTKKSKKHKKHKNK
KKKKK----KEKEKKYKRQPEESESKAKSHHEENMDLESDSFLKFDSEPSAMALEHPVRA
FGLSE------------------------TSETP-AVVLEPPVVSMEVSQPHTLETLKPA
TKSAELSL----ASTSEISVQSEQSVAVTLEPPMTKILDSFSTAPVSTT---TVVLKS-S
EPVVTMSMDYQMKS----LECTSPESSK-TI-LEPPVAKVLEPSETL-MVSSQTPIEAHP
EPSTS-TMDIAESSETEDLRLPEQPVEVPSETADSSMTRSQELLELPKPTALELPESS-V
ASAMQLPGPPATSMPELQGPPVTPGLELPGPSAT-----------PVPELPGPPSTPVPE
LLGPPATA--VSELPGPSVISVPQLSQELPGLPAPSMGLEPTQEVPEPPVMAQ-------
---------ELPGQPAVTVAMELTEQPATTTELEQPVGMTTMEHPGQPEVTTATTLLGQP
EAAMVLELPGQPVATTALELPGQPSVTGVPELPGLPSATRALELSGQPVATGALEL-PGQ
LMATGA-LEFAGQSG-AAGALELL-GQPLATGVLELPG-QPGAPELPGQPAATVALEISV
-QSVVT-TELSTMTVSQSLEVPS-TTALE--SYNTVAQELPTTLVGETSITVGVDPLMAQ
ESHMLASNTMETHMLASNTMDSQMLASNTMDSQMLASNTMDSQMLASSSMDSQMLASSSM
DSQMLASSSMDSQMLASSSMDSQMLASSSMDSQMLATSS--MDSQMLASSSMDSQM----
------LATSSMDSQMLATSS--MDSQMLATSTMDSQMLATSSMDSQMLA--SGTMDSQM
LASGTMDAQMLASGTMDAQMLASSTQDSSMLGSKSPDPYRLAQDPYRLAQDPYRLGHDPY
RLGHDAYRLGQDPYRLGHDPYRLTPDPYRMSPRPYRIAPRSYRAVPRPYRLAPRPLMLAS
RRSMMMSYAAERSMMSSY--ERSMMSYERSMMSPMAERSMMSAYERSMMSAYERSMMSPM
AERSMMSAYERSMMSAYERSMMT---DRS---MMADRSMMSSYSAADRS-MMSSYSAADR
TMMSSYTADRSMMSMAADSYTDSYTDTYTEAYMVPPLPPEEPPTMPPLPPEEPPMTPPLP
PEEPPEGTALPTEQSALTAENTWPTEMPPLPPEESVSLPEPPVSQSEISEPSVVPANYSL
SASEPSMLA-SEAAMTVPDPP-LEPESVVTSAPIESAAEAEEHE-VPERPGTYMVSETTM
MSAEPTMSTSEPSIMSETAETFDSMRAPGHV-A-SEVSVSLLEPAV-----------TIP
EPSQSAL---------------E-PPAE--AV--------PEPPAEAVPEPS--------
--AEVPEPLAM------TVPGPLAMAVLEPSPEAVPEPM----------ALVEPEHVTIP
VPVVSAP------EPPVPVLEPAVSVQ--PNIISE--------P--PVVQESTVTVAEPA
VTVSEQTQITPTEMALESASVMLKSG------VITGMNLIST-YQDLTPEIGMQEVSMHS
DEE-PHDEEHLKNDSYESEHGINRDLDINNHLIAKDMEHSTMSTASTGAVGEIGKEKILA
INETKQCTVLDTCPSVSEADVGGT-LSSAGPL-AVEPDT----LGTSKGIEFATVSALSS
VSKYDVEVSLTTQDTEHDMVISTSPSGGSEADIEGPLPAKDIHLDLPSN-NFISN-DTEG
PLPIKDSDQTLAVALSPKES-S-EDKEVPLPAKETVSESALPANIEDINEADLVRPLLPK
DMERLTSLRAG-------IEGPLLASEVERDKS-AASPVVMSIPERASE-SSSEEKDDYE
IFVKVKDTHEKSKKNKNRDKGEKEKK----------------------------------
-----------------------------------------------RDSSLRSRSKRSK
SSEHKSRK-------RTSESRSRARKRSSKSKS-RRSQTRSRSRSRRRRRSSRSRSKSRG
RRSVSKEKRKRSPKHRSKSRERKRKRSS-SRDNRKTVRARSRTPSRRSRSHTPSRRRRSR
SVGR-RSFSISPSRRSRT-----PSRRSRTPSRRSRTPSRRSR-------TPSRRSRTPS
RRSRT--PSRRSRTPSRRRRSRSV--VRRRSFSISPVR-LRRSRT-PLRRRFSRSPLRRK
RSRSSE---RGRS-PKRLTDLNKAQLLEIAKANAAAMCAKAGVPLPPNLKPA-PP-PTIE
EKVAKKSGGATIEELTEKCKQIAQSKEDDDVIVNKPHVSDEE-EEEPPFYHHPFKLSEPK
PIFFNLNIAA---AKPTPPKSQVTLTKEFPVSSGSQHRKKEADSVYGEWVPVDKNGEENK
D-DDNVFSSNLPSEPVDISTAMSERALAQKRLSENAFDLEAM-SMLNRAQERIDAWAQLN
SIPGQFTGSTGVQVLTQEQLANTGAQAWIKKDQFLRAAPV
PF01585 // ENSGT00730000111141_0_Domain_1 (2621, 2665)
TGGMGAVLMRKMGWREGEGL
GKNKEGNKEPILVDFKTDRKGLVAENSGT00730000111141_0_Linker_1_2 (2665, 2688)
VGERAQKRSGNFSAAMKDLSGKHPF00035 // ENSGT00730000111141_0_Domain_2 (2688, 2724)
PVSALMEICNKRR
WQPPEFLLIHDSGPDHRKHFLFRVLRNGSPYQPNCMFFLN-RY-----------------
----------------
ENSMFAP00000036158 (view gene)
ENSTSYP00000007689 (view gene)
MATNIEQIFRSFVVSKFREIQQELSSGRSEGQLNGETNTPIEGNQASDA-AASARNLPNE
EIVQKIEEVLSGVLDTELRHKPDLKEAS-RKSRCVSVQTDPTD-EIPTKKSKKHKKHKNK
KKKKK----KEKEKKYKRQPEESESKPKSHHDGNIDLESDSFLKLDSETSAMSLEHPTRA
FDLSE------------------------TSESS-AAVLEPSVVSMEVSEPHILETLKPA
TDTAELSV----APTSVVSEQSEESVTVMLEPSMTRILDSFATAPVPTT---TVVPKS-S
EPVVTMSVEYQMKSVLKPLENTSPESSKIML-VEPPVAKVLEPSETL-VVSSETPTEVYP
EPSTSTTMDFPESPATEAPRLPEQPVEVPSEIADSSMTRPQELLELPKTTALELPESS-V
ASAMELPGPPATSMPELQGPPVTPVLELPGPSAT-----------LVPELPGPLSTPVPE
LPGPPATA--VPELPGPSVTPVPQLSQELAGLPAPSMGLEPPQEVPEPPVMAQ----ELP
GLPVVTAAVELPEQPAVAVAMELTEQPVMTTELEQPVGMTTVDHPGHPEVTTATGLLGQP
EAAMVLELPGQPVATTALELQGQPSVTGVPELPGLPSATRALELSGQPVATGALEL-PGQ
LMATGA-LEFAGQSG-AAGALELL-GQPLATGVLELPG-QPGAPELPGQPVATVALEISV
-QSVVTTSELSTMTVSQSLEVPS-TTALE--SYNTVAQELPTTLVGETSVTVGVDPLMAQ
ESHMLASNTMETHMLASNTMDSQMLASNTMDSQMLASNTMDSQMLASSTMDSQMLATSTM
DSQMLATSSMDSQMLATSSMDSQMLATSSMDSQMLATSS--MDSQMLATSSMDSQM----
------LATSSMDSQMLATSS--MDSQMLATSTMDSQMLATSSMDSQMLA--SGTMDSQM
LASGTMDAQMLASGTMDAQMLASSTQDSAMLGSKSPDPYRLAQDPYRLAQDPYRLGHDPY
RLGHDAYRLGQDPYRLGHDPYRLTPDPYRMSPRPYRIAPRSYRIAPRPYRLAPRPLMLAS
RRSMMMSYAAERSMMSSY--ERSMMSYERSMMSPMAERSMMSAYERSMMSAYERSMMSPM
AERSMMSAYERSMMSAYERSMMSPMADRSMMSMGADRSMMSSYSAADRS-MMSSYSAADR
SMMSSYTADRSMMSMAADSYTDSYTDTYTEAYMVPPLPPEEPPTMPPLPPEEPPVTPPLP
PEEPPEGPALPNEQSALTAENTWPAEVPSLPSEESVSQPEPPVCQSEISEPSAMPVNYSM
SASDPSVLA-SEASVTVPEPP-LEPESSVTSTPVESAVVAEEHEIVPERAVTSVVSETPI
MSTEPTILTSEPPIMSETAETFDSMRVSGHT-ASSEVSISLLEPAV-----------TLP
EPAESIL---------------E-PLAM--AVPDPLAMTVPELPAAVP--------ELPL
PAVAVP--------------EIPAVAVPEPLVMAVPESP----------VMAEPEHVTVS
VPVVSAL------EPTVPVLEPTMSVFQ-PAMIVSE-------P-SVSVQESTVTISESA
VTVSEQTQVISTEMVIESTPMILESSVISSLHVMKRMNLTSG-DQNLVPEIGMPEIPLHS
GEE-PHADGHLKSDSYESEIGINIDLSVNNHLIAKEMEHNTMSVASSGSGGEIGEEKILP
VNETQQSTVLDTYPGVSEADVGGT-LT-SGPV-ALEPDA----MGTSKDIEFITASAFSS
VNRYDIEVSLTTQDTEHDMVISTSPSAGSEADIEGPLPAKDIHLDLPSNDNLVSK-DAEE
PLPMKDSDQTLAVALSPKES-SGEEKEVTLPTKEM-SDSGFPANIEDINEADLVRPLLPK
DMERLTNLRAG-------IEGSLLASEVERDKS-AASPVVI--TERASE-SSSEEKDDYE
IFVKVKDTHEKSKKNKNRDKGEKEKK----------------------------------
-----------------------------------------------RDSSLRSRSKRSK
SSEHKSRK-------RTSESRSRARKRSSKSKS-HRSQTRSRSRSRRRRRSSRSRSKSRG
RRSVSKEKRKRSPKHRSKSRERKRKRS-----------TRSRTPSRRSRSHTPSRRRRSR
SVGRRRSFSISPSRRSRT-----PSRRSRTPSRRSRTPSRRSR-------TPSRRSRTPS
RRSRT--PSRRSRTPSRRRRSRSV--VRRRSFSISPVR-LRRSRT-PLRRRFSRSPIRRK
RSRSSE---RGRS-PKRLTDLDKAQLLEIAKANAAAMCAKAGVPLPPNLKPA-PP-PSIE
EKAAKKSGGATIEELTEKCKQIAQSKEDDDVIVNKPHVSDEE-EEEPPFYHHPFKLSEPK
PIFFNLNIAA---AKPTPPKSQVTLTKEFPVSSGSQHRKKEADSVYGEWVPVEKNGEENK
D-DDNVFSSNLPSEPVDISTAMSERALAQKRLSENAFDLEAM-SMLNRAQERIDAWAQLN
SIPGQFTGSTGVQVLTQEQLANTGAQAWIKKDQFLRAAPV
PF01585 // ENSGT00730000111141_0_Domain_1 (2621, 2664)
TGGMGAVLMRKMGWREGEGL
GKNKEGNKEPILVDFKTDRKGLVENSGT00730000111141_0_Linker_1_2 (2664, 2687)
AVGERAQKRSGNFSAAMKDLSGKPF14709 // ENSGT00730000111141_0_Domain_2 (2687, 2732)
HPVSALMEICNKRR
WQPPEFLLVHDSGPDHRKHFLFRVLRNGSPYQPNCMFFLN-RY-----------------
----------------
ENSOCUP00000003354 (view gene)
--IKVEPFFELLGVSKFYKFKNRMTSGRSEGQLNGETNTPVEGNQAGD-AAASARNLPNE
EIVQKIEEVLSGVLDTELRCKPDLKEAS-RKSRCVSVQTDPTD-EVPTKKSKKH------
-------------KKYKRQPEESESKPKSHNDGNIDLESDSFLKFDSESSAMALE-PVRA
FGLSE------------------------TSESS-AVLLEPPLVSMEVSEPHILETLKPA
TKTVELSV----ASTSVISEQSEQPVAGMLEPSTSKILDSFATAPVPAT---AVVPKS-S
EPVVTMSVEYQ-KSVLKSLENASTESSKM--LVDPLGAKALEPPQTP-GIASEAPAEVHP
EPSTSTAMDFSESSATEVLSLPEQPVDVPSEMTESSVTRPQELMELPKTTALELPESS-V
ASALELPGPPAISMPELQGPPVTPVLGLPGPSA-----------TPVPELPGPLSTPVPE
LPGPPVTP--VPELPGPPVTPVPQLSQDLPGLPAPSMGLEPPQEVPEPPVMAQ----ELP
GLPAVSSAVELPGQPAVTVAMELTEQPVTTTELEHPVGMTTVEHPGHPEVTAATGLLGQP
EATMVLELPGQPEATTALELPGQPSVTGVPELPGLPSATRALELSGQPVATGALEL-PGQ
LMATGA-LEFSGQSG-AAGALELL-GQPLATGVLELPG-QPGAPELPGQPVATVALEISV
-QSVVTTSELSTMTVSQSLEVPS-TTALE--SYNTVAQELPTTLVGETSVTVGVDPLLAQ
DSHMLASNTMEAHMLASNTMESQMLASNTMDSQMLASNTMDSQMLASSTMDSQMLASSTM
DSQMLATSSMDSQMLATSTMDSQMLATSSMDSQML------------ATSSMDSQMLATS
SMDSQMLATSSMDSQMLATSS--MDSQMLATSSMDSQMLATSSMDSQMLA--SGTMDSQM
LASGTMDAQMLASGTMDAQMLASSTQDSAMLGSKSPDPYRLAQDPYRLAQDPYRLGHDPY
RLGHDAYRLGQDPYRLGHDPYRLTPDPYRMSPRPYRIAPRSYRIAPRPYRLAPRPLMLAS
RRSMMMSYAAERSMMSSY--ERSMMSYERSMMSPMAERSMMSAYERSMMSAYERSMMSPM
AERSMMSAYERSMMSAYERSMMSPMADRSMMSMGADRSMMSSYSAADRS-MMSSYSAADR
SMMSSYT-DRSMMSMAADSYTDTYTDTYTEAYMVPPLPPEEPPTMPPLPPEEPPMTPPLP
PEEPPEGPALPAEDSALAAENTWPTEVPALPTEGSVSQPEPPVSQSEISEPSAVPAHYSV
SASESSVLT-SEAVMTVPEPP-LEPESSVTSTPRESAGVEQ---EVPERPVTE---TPMM
-SVEPAVLTAEPPVVSETAETLDSMRTSA--HA-SELSVPLLEPAV-----------TIP
EPAENILELPAVAVSEAAAVAVQEPPAMVVAVPESPTT------VVAVPESPAVAVLEPP
-PVAE-------------------------------SEPQV--VAVDPPAVIEPEHVTIP
VS-VSAL------EPTVPIVDPAVSVLQPAMIISE---------PSLSVQESTVTISESA
VTISEQTEIISTEMVLESTPVILQSSVMSSSLVVKGMNLLSG-DQSFAPEIGMQKI-PLH
SSEEPRAEGHLKSDSYESECSINTDLNTNNHLIAKEMEHSTVSAADTGPVGEIGEEKILP
IKETEQCTVLDISSGVNETDARGT-LSSTGPL-AVEPDAV---E-TSKGTEYMTPSALGS
VNKYDVDVSLTTQDTEHDMVISTSPSGGSEADIEGPLPAKDIHLDLASG-NL-SK-DTEE
LLPVKESDQTLAVVVSPKES-SGEDKEVPLPTKEI-PDSGFPTNIEDINEADLVRPLLPK
DMERLTSLRAG-------IEGPLLASEAERDKS-AASPVVISIPERASE-SSSEEKDDYE
IFVKVKDTHEKSKK-KNRDKGEKEKK----------------------------------
-----------------------------------------------RDSSLRSRSKRSK
SSEHKSRK-------RTSESRSRARKRSSKSKS-HRSQTRSRSRSRRRRRSSRSRSKSRG
RRSVSKEKRKRSPKHRSKSRERKRKRSS-SRDNRKAVRARSRTPSRRSRSHTPSRRRRSR
SVGRRRSFSVSPSRRSRT-----PSRRS--------------RTPSRRSRTPSRRSRTPS
RRSRTPSPTRRSRTPSRRRRSRSV--VRRRSFSISPVR-LRRSRT-PLRRRFSRSPIRRK
RSRSSE---RGRS-PKRLTDLDKAQLLEIAKANAAAMCAKAGVPLPPNLKPA-PP-PTIE
EKVAKKSGGATIEELTEKCKQIAQSKEDDDVIVNKPHVSDEE-EEEPPFYHHPFKLSEPK
PIFFNLNIAA---AKPTPPKSQVTLTKEFPVSSGSQHRKKEADSVYGEWVPVEKNGEENK
D-DDNVFSSNLPSEPVDISTAMSERALAQKRLSENAFDLEAM-SMLNRAQERIDAWAQLN
SIPGQFTGSTGVQVLTQEQLANTGAQAWIKKDQFLRAAPV
PF01585 // ENSGT00730000111141_0_Domain_1 (2621, 2664)
TGGMGAVLMRKMGWREGEGL
GKNKEGNKEPILVDFKTDRKGLVAVGERAQKRSGNFSAAMKDLSGKHPVSALMEICNKRR
WQPPEFLLVHDSGPDHRKHFLFRVLRNGSPYQPNCMFFLN-RY-----------------
----------------
ENSSSCP00000056796 (view gene)
ENSNGAP00000000072 (view gene)
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
--------------------------------------------------------MAAD
IEQVFRSFVVSKFREIQQ-----ELSSG--------------RSE-----------GQLN
GETNT--TIEGNQAGDAAAS----------------------------ARSLPNEEIVQK
IEEVLS---GVLD-TELRYKSDKAQLLEIAKANAAAMCAKAGVPLPPNLKPA-PP-PTIE
EKVAKKSGGATIEELTEKCKQIAQSKEDDDIIVNKPHVSDEE-EEEPPFYHHPFKLSEPK
PIFFNLNIAA---AKPTPPKSQVTLTKEFPVSSGSQHRKKEADSVYGEWVPVEKNGEENK
D-DDNVFSSSLPSEPVDISTAMSERALAQKRLSENAFDLEAM-SMLNRAQERIDAWAQLN
SIPGQFTGSTGVQVLTQEQLANTGAQAWIKKDQFLRAAPV
PF01585 // ENSGT00730000111141_0_Domain_1 (2621, 2664)
TGGMGAVLMRKMGWREGEGL
GKNKEGNKEPILVDFKTDRKGLVENSGT00730000111141_0_Linker_1_2 (2664, 2687)
AVGERAQKRSGNFSAAMKDLSGKPF14709 // ENSGT00730000111141_0_Domain_2 (2687, 2732)
HPVSALMEICNKRR
WQPPEFLLVHDSGPDHRKHFLFRVLRNGSPYQPNCMFFLN-RY-----------------
----------------
ENSVPAP00000008517 (view gene)
--------------------------GRSEGHLYGETNTPVEGNQAG-DAAASARSLPNE
EIVQKIQ-VLSGVLDTELRYKPDLKEAS-RKSRCVSVQTDPTD-EIPTKKSKKHKKHKNK
KKKKK----KEKEKKYKRQSEESESKPKSHHDGNIDLESDSFLKFDSEPS-TALEHPVRV
FGLSD------------------------TSESP-AVVLEPPVVSMEVSEPHTLETLKPA
SKMAELSV----ASTSVVSVQSEQSVAVTLEPSMTKILDSFAAAPVPTT---TVAPKS-S
EPVVTVSVEYQMKSVLKSLESTPPEPSKIML--EPPVAKVLEPPETL-V-SSETPAEVYP
EPSTSTTTDFPESSATEVLRLPEQPVEVPSEIADSSMTRPQEXXXXXXXXXXXXXXXX-X
XXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXX----------XXXXXXXXXXXXXXXX
XXXXXXXX--XXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXX----XXX
XXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXX
XXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXX
XXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXX
XXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXX
XXXXXXXXXXXXXXXXXXTMDSQMLASNTMDSQMLASSTMDSQMLATS----------SM
DSQMLATSSMDSQMLATSSMDSQMLATSSMDSQML------------ATSSMDSQMLATS
SMDSQMLATSSMDSQMLATST--MDSQMLATSTMDSQMLATSSMDSQMLA--SGTMDSQM
LASGTMDAQMLASGTMDAQMLASSTQDSAMLGSKSPDPYRLAQDPYRLAQDPYRLGHDPY
RLGHDAYRLGQDPYRLGHDPYRLTPDPYRMSPRPYRIAPRSYRIAPRPYRLAPRPLMLAS
RRSMMMSYAAERSMMSSY--ERFMMSYERSMMSPMAERSMMSAYERSMMSAYERSMMSPM
AERSMMSAYERSMMSAYERSMMSPMADRSMMSMGADRSMMSSYSAADRS-MMSSYSAADR
SMMSSYTADRSMMSMAADSYTDSYTDTYTEAYMVPPLPPEEPPTMPPLPPEEPPMTPPLP
PEEPPEGPALSTEQSALTAENTWSTEVPALPPEESVSLPEPPVSQSEISEPSAVPANYSV
SASEPSVLA-SEAAVTVPEP--LEPEPSVTSTPVESAAVAEEHEIVPERPVTYLVSEIPI
-MSAPTVLTSXXXXXX--------XXXXXXXSVASEVSMSLLEPAV-----------PIP
EPSQSTLELPAMAVSEPPAVTVPEPPAVAIP-----------------------------
--------------------APPAVAAPEPPAMAAPEPLVV--AVPEPLAVAEPEHVTIP
VPVVSAL------EPAVPVLEPVVSVLQPNMIVSE---------PSISVQESTVAISEPA
VTISEQTQVISTDMVLESTPITLESS------VIKRMNLLSG-DENLAPETGMQEI-PMH
SDEEPHAEGHLKNDSCETGNGINIDLNIN--HLAKEMEHNTVSAVSTGAVGEIGEEKIVP
TNETEQCTVLDTCPSVSEADVGGT-LSSPGPL-ALEPDAM---G-TGKGFEFATASALSS
VSKYDIEVPLTTQDTEHDMVISTSPSGGSEADIEGPLPAKDIHLDLPSNNNFISK-DAEG
PFPIQESDQTLAVALSSKES-SGEDKELSLPPKEILPESGFSANIDDINEADLVRPLLPK
DMERLTSLRAG-------LEGSSLASEVERDKS-ATSPVVISIPERASE-SSSEEKDDYE
IFVKVKDTHEKSKKNKNRDKGEKEKK----------------------------------
-----------------------------------------------RDSSLRSRSKRSK
SSEHKSRK-------RTSESRSRARKRSSKSKS-HRSQTRSRSRSRRRRRSSRSRSKSRG
RRSVSKEKR-RSPKHRSKSRERKRKRSS-SRDNRKTGRARSRTPSRRSRSHTPSRRRRSR
SAGR-RSFSISPSRRSRT-----PSRRS--------------RTPSRRSRTPSRRSRTPS
RRSRT--PSRRSRTPSRRRRSRSV--ARRRSFSISPVR-LRRSRT-PLRRRFSRSPIRRK
RSRSSE---RGRS-PKRLTDLNKAQLLEIAKANAAAMCAKAGVPLPPNLKPA-PP-PTIE
EKVAKKSGGATIEELTEKCKQIAQSKEDDDVIVNKPHVSDEE-EEEPPFYHHPFKLSEPK
PIFFNLNIAA---AKPTPPKSQVTLTKEFPVSSGSQHRKKEADSVYGEWVPVEKNGEENK
D-DDNVFSSNLPSEPVDISTAMSERALAQKRLSENAFDLEAM-SMLNRT--EIDAWAQLN
SIPGQFTGSTGVQVLTQEQLANTGAQAWIKKDQFLRAAPV
PF01585 // ENSGT00730000111141_0_Domain_1 (2621, 2665)
TGGMGAVLMRKMGWREGEGL
GKNKEGNKEPILVDFKTDRKGLVAVGERAQKRSGNFSAAMKDLSSKHPS---LEICNKRR
WQP-EFLLVHDSGPDHRKHFLFRVLRNGA-LTPNCMFFLN-RY-----------------
----------------
ENSMUSP00000109671 (view gene)
ENSRBIP00000041914 (view gene)
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
--------------------------------------------------------MATN
IEQIFRSFVVSKFREIQQ-----ELSSGRN-------------------------EGQLN
GETNT--PIEGNQAGD------AA--A--------------------SARSLPNEEIVQK
IEEVLS---GVLD-TELRYKPDKAQLLEIAKANAAAMCAKAGVPLPPNLKPA-PP-PTIE
EKVAKKSGGATIEELTEKCKQIAQSKEDDDVIVNKPHVSDEE-EEEPPFYHHPFKLSEPK
PIFFNLNIAA---AKPTPPKSQVTLTKEFPVSSGSQHRKKEADSVYGEWVPVEKNGEENK
D-DDNVFSSNLPSEPVDISTAMSERALAQKRLSENAFDLEAM-SMLNRAQERIDAWAQLN
SIPGQFTGSTGVQVLTQEQLANTGAQAWIKKDQFLRAAPV
PF01585 // ENSGT00730000111141_0_Domain_1 (2621, 2664)
TGGMGAVLMRKMGWREGEGL
GKNKEGNKEPILVDFKTDRKGLVENSGT00730000111141_0_Linker_1_2 (2664, 2687)
AVGERAQKRSGNFSAAMKDLSGKPF14709 // ENSGT00730000111141_0_Domain_2 (2687, 2732)
HPVSALMEICNKRR
WQPPEFLLVHDSGPDHRKHFLFRVLRNGSPYQPNCMFFLN-RY-----------------
----------------
ENSAMEP00000015669 (view gene)
MATNIEQIFRSFVVSKFREIQQELSSGRSEGQLNGETNTPNEGNQAGDA-AASARSLPNE
EIVQKIEEVLSGVLDTELRYKPDLKEAS-RKSRCVSVQTDPTD-EIPTKKSKKHKKHKNK
KKKKK----KEKEKKYKRQPEESESKAKSHHDGNIDLESDLFLKFDSEPSAMALEHPIRA
FGLSE------------------------TSKSP-AVVLEPPVVSMEVSEPHTLETLKPA
TKTTELSV----ASTSVISVQSEQSVAVTLEPSMTKILDSFTTAPVPTT---TVVLKS-P
EPVVTMSMEYQMKPVLKSLEMTSPEQSKIM--LEPPVAKGLDSSEAL-VVSSEIPTEVHP
EPSTSTTMDFPESSATEGLRLPEQPVEVPSEIADSSVTRPQELLELPKTTALELPESS-V
ASVMELPGPPATSRPELQGPSVTPVLELPGPSAT-----------LLPELPGPLSTPVPE
LLGPPATA--VPELPGPSATSVPQLSQELPGLPAPSVGLEPPQEVPEPPVMAQ----ELP
GLPAVTAAVELPGQPAVTVAMELTEQPVTTSELEQPVGMTTVEHPGQPEVTTATGLLGQP
EAAMVLELPGQPVATTALELPGQPSVTGVPELPGLPSATRALELSGQPVATGALEL-PGQ
LMATGA-LEFSGQSG-AAGALELL-GQPLATGVLELPG-QPGAPELPGQPVATVALEISV
-QSVVT-TELSTMTVSQSLEVPS-TTALE--SYNTVAQELPTTLVGETSVTVGVDPLMAQ
ESHMLASNTMETHMLASNTMDSQMLASNTMDSQMLASSTMDSQMLA--------------
------------------------------------------------------------
------------------TSS--MDSQMLATSTMDSQMLATSSMDSQMLA--SGTMDSQM
LASGTMDAQMLASGTMDAQMLASSTQDSAMLGSKSPDPYRLAQDPYRLAQDPYRLGHDPY
RLGHDAYRLGQDPYRLGHDPYRLTPDPYRMSPRPYRIAPRSYRIAPRPYRLAPRPLMLAS
RRSMMMSYAAERSMMSSY--ERSMMSYERSMMSPMAERSMMSAYE---------------
----------RSMMSAYERSMMSPMADRSMMSMGADRSMMSSYSAADRS-MMSSYSAADR
SMMSSYTADRSMMSMAADSYTDSYTDTYTEAYMVPPLPPEEPPTMPPLPPEEPPMTPPLP
PEEPPEGPALPTEQSALTAENTWPTEVPALPPEESVSLSEPSVSQSEISEPSALPANYSV
--SDPSVLA-SEAAVTVPEP--LEPESLVTSTPVESAVVAEEHQVVPERAVTYVVSETPI
MSAEPTVLTSEPSVMSETAETFDSMKASGHV-A-SEASLSLLEPAA-----------TNP
EPSQSTL---------------E-LPAM--AVSELPAVAVPEPPTGTVAEPPAVA-VLET
SAVAIPDP----------------------AAVAVPETL----------ALAESEHVTIP
VPVVSAL------EPTVPVLEPAVSVPQ-PNAVVSE-------P-SVSLQESTVIISEPA
VTVSEQTQVIPTEMVLESTPMLLESN------VIKGVNLLSG-DQSLAPEIGIQEIPMHS
DDE-PHAEGHLKNDPYESEHGTNIDLNINNHLVAKDMEHNTVSAASTGAVVESGEGNILS
ISETKQCTVLDTCSSVSEADGGT--LSSAGPL-ALEPDA----MGTSKGIEFATESVLGS
VSNYDVEVSLTTQDTEHDMIISTSPSGGSEADIEGPLPAKDIHLDLPSN-NFITK-DVEG
PLSIKDCDQTLAVALSPKES-SGEDKEVPLPTKEIMSHSGFSANIDDINEADLVRPLLPK
DMERLTSLRAG-------IEGPLLASEVERDKS-AASPVVISIPERASE-SSSEEKDDYE
IFVKVKDTHEKSKKNKNRDKGEKEKK----------------------------------
-----------------------------------------------RDSSLRSRSKRSK
SSEHKSRK-------RTSESRSRARKRSSKSKS-HRSQTRSRSRSRRRRRSSRSRSKSRG
RRSVSKEKRKRSPKHRSKSRERKRKRSS-SRDNRKTVRARSRTPSRRSRSHTPSRRRRSR
SVGR-RSFSISPSRRSRT-----PS-----------------------------------
RRSRT--PSRRSRTPSRRRRSRSV--VRRRSFSISPVR-LRRSRT-PLRRRFSRSPIRRK
RSRSSE---RGRS-PKRLTDLNKAQLLEIAKANAAAMCAKAGVPLPPNLKPA-PP-PTIE
EKVAKKSGGATIEELTEKCKQIAQSKEDDDVIVNKPHVSDEE-EEEPPFYHHPFKLSEPK
PIFFNLNIAA---AKPTPPKSQVTLTKEFPVSSGSQHRKKEADSVYGEWVPVEKNGEENK
D-DDNVFSSNLPSEPVDISTAMSERALAQKRLSENAFDLEAM-SMLNRAQERIDAWAQLN
SIPGQFTGSTGVQVLTQEQLANTGAQAWIKKDQFLRAAPV
PF01585 // ENSGT00730000111141_0_Domain_1 (2621, 2665)
TGGMGAVLMRKMGWREGEGL
GKNKEGNKEPILVDFKTDRKGLVAENSGT00730000111141_0_Linker_1_2 (2665, 2688)
VGERAQKRSGNFSAAMKDLSGKHPF00035 // ENSGT00730000111141_0_Domain_2 (2688, 2724)
PVSALMEICNKRR
WQPPEFLLVHDSGPDHRKHFLFRVLRNGSPYQPNCMFFLN-RY-----------------
----------------
ENSTBEP00000002791 (view gene)
-ATNIEQVFRSFVXXXXXXXXXXXXXGRSEGQLNGETNTPVEGNQAGD-AAASARSLPNE
EIVQKIEEVLSGVLDTELRCKPDLKEAS-RKSRCVSVQTDPTD-EIPTKKSKKHKKHKNK
KKKKK----KEKEKKYKRQPEESESKPKSHHDGNMDLESDSFLKFDSEPSVMALEHPVRA
YGLSE------------------------TSESS-AVVLEPPVVPMEVSEPHILETPKPA
TKTAELSV----ASTS-MSEQSEQSVIVMLEPS-TKILDSFATAPVPTT---TVVLKS-S
EPIITMSTEYQMKSVLKSLESKSPEPSKMLV-EPPPVAKVLDSSENL-AATSDTPTEVHP
EPGTSTTMDFPESSATEVLRLPEQPVEVSSEVADSSMTRPQELLEQPKTTALELPESS-V
ASVVELPGPPATSMPELQGPPVTPVLELPGPSA-----------TTVPELSGPLSTPAPE
LSGPPATA--GPELPASTPVPQCQMTS---------APGLEPQEVPEPPVMAQ----ETQ
---------------PAVVAMELTEQPVTTTELEQPVGMTAVEHPGHPEVTTAAGLLGQP
EATMVLELPGQPVATTALELPGQPSVTGVPELPGLPSATRALELSGQPVATGALEL-PGQ
LMAAGA-LEFSGQSG-AAGALELL-GQPLATGVLELPG-QPGAPELPGQPVATVALEISV
-QSVVTTSELSTMTVSQSLEVPS-TTALE--SYNTVAQELPTTLVGETSVTVGVDPLMAQ
ESHMLASNTMETHMLASNTMDSQMLASNTMDSQMLASNTMDSQMLASSTMDSQMLASSTM
DSQMLATSSMDSQMLATSSMDSQMLATSSMDSQ---------------------------
---------------------------MLATSSMDSQMLATSSMDSQMLA--SGTMDSQM
LASGTMDAQMLASGTMDAQMLASSTQDSAMMGSKSPDPYRLAQDPYRLAQDPYRLGHDPY
RLGHDAYRLGQDPYRLGHDPYRLTPDPYRMSPRPYRIAPRSYRIAPRPYRLAPRPLMLAS
RRSMMMSYAAERSMMSSY--ERSMMSYERSMMSPMAERSMMSAYERSMMSAYERSMMSPM
AERSMMSAYERSMMSAYERSMMSPMADRSMMSMVADRSMMSSYSAADRA-MMSSYSAADR
SMMSSYT-DRSMMSMAADSYADSYTDTYTEAYMVPPLPPEEPPTMPPLPPEEPPMTPPLP
PEEPPEGPALPTEESAL-AENTWPTEMPALPPEESVSQPEPPMNQSDISEPSAVLANYSV
SASEPSVLA-SEAAMTAAETP-LEPETSFTSTPVESAVIAEEHEIVSERPVTCVLSETPV
MSTETTMLTSEPTVRSETADMFDSTRASG--HIASEVSVSLLDPAV-----------TTT
EPAESILEVPAVTVPEPLTT--LEPPAV--------------------------------
-------------------------VISESPAVTVSEPPAV--AVSEPPAMAEPEHVGIP
VP-VSAM------EPTVPILEPAVSVLQPSMIVSE---------PSVSVQESTVTVSEPA
VTISEQTQMAPTDMVLESAPMIVESSVMSSSHVIKGVNLLSV-DQNLVPEIGMQEI-PLH
SGEQPHAEGHLRSDSYESEHGINVXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXX
XXXXXXXXXXXXXXXXXXXXXXXX-XXXXXXXXXXXXXXX---XXXXXXXXXXXXXXXXX
XXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXX-XXXX
XXXXXXXXXXXXXXXXXXXX-XXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXX
XXXXXXXXXXX-------XXXXXXXXXXXXXXX-XXXXXXXXXXXXXXX-XXXXXXXXXX
XXXXXXXXXXXXXXXXXXXXXXXXXX----------------------------------
-----------------------------------------------XXXXXXXXXXXXX
XXXXXXXX-------XXXXXXXXXXXXXXXXXX-XXXXXXXXXXXXXXXXX-X-XXXXXX
XXXXXXXXXXXXXXXXXXXXXXXXXXXX-XXXXXXXXXXXXXXXXXXXXXXXXXXXXXSR
SVGRRRSFSISTSRRSRT-----PSRRS--------------RTPSRRSRTPSRRSRTPS
RRSRT--PSRRSRTPSRRRRSRSV--VRRRSFSISPVR-LRRSRT-PLRRRFSRSPIRRK
RSRSSE---RGRS-PKRLTDLDKAQLLEIAKANAAAMCAKAGVPLPPNLKPA-PP-PTIE
EKVAKKSGGATIEELTEKCKQIAQSKEDDDVIVNKPHVSDEE-EEEPPFYHHPFKLSEPK
PIFFNLNIAA---AKPTPPKSQVTLTKEFPVSSGSQHRKKEADSVYGEWVPVEKNGEENK
D-DDNVFSSSLPSEPVDISTAMSERALAQKRLSENAFDLEAM-SMLNRAQERXXXXXXXX
XXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXX
XXXXXXXXXXXXXXXXXXXXXLVAVGERAQKRSGNFSAAMKDLSGKH
PF00035 // ENSGT00730000111141_0_Domain_2 (2688, 2724)
PVSALMEICNKRR
WQPPEFLLVHDSGPDHRKHFLFRVLRNGA-LTPNCMFFLNRY------------------
----------------
ENSP00000348984 (view gene)
ENSPANP00000013073 (view gene)
ENSDORP00000022865 (view gene)
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
--------------------------------------------------------MAAD
IEQVFRSFVVSKFREIQQ-----ELSSG--------------RSE-----------GQLN
GETNT--PIEGNQAGDAAAS----------------------------ARSLPNEEIVQK
IEEVLS---GVLD-TELRYKPDKAQLLEIAKANAAAMCAKAGVPLPPNLKPA-PP-PTIE
EKVAKKSGGATIEELTEKCKQIAQSKEDDDVIVNKPHVSDEE-EEEPPFYHHPFKLSEPK
PIFFNLNIAA---AKPTPPKSQVTLTKEFPVSSGSQHRKKEADSVYGEWVPVEKNGEENK
D-DDNVFSSNLPSEPVDISTAMSERALAQKRLSENAFDLEAM-SMLNRAQERIDAWAQLN
SIPGQFTGSTGVQVLTQEQLANTGAQAWIKKDQFLRAAPV
PF01585 // ENSGT00730000111141_0_Domain_1 (2621, 2664)
TGGMGAVLMRKMGWREGEGL
GKNKEGNKEPILVDFKTDRKGLVENSGT00730000111141_0_Linker_1_2 (2664, 2687)
AVGERAQKRSGNFSAAMKDLSGKPF14709 // ENSGT00730000111141_0_Domain_2 (2687, 2732)
HPVSALMEICNKRR
WQPPEFLLVHDSGPDHRKHFLFRVLRNGSPYQPNCMFFLN-RY-----------------
----------------
ENSPPAP00000021001 (view gene)
ENSPSIP00000008930 (view gene)
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------FRSFIVLAF---------------------
------------------------------------------------------------
--------FRVVEEL---VSGKVE--GQQNGE-TIPAEQANTSDDTVTGAGR-LQNDQIV
--QKIEEVLSGALDTQLQCKSDKAQLLEIAKANAAAMCAKAGVPLPPSLLPIIAPAKK-E
EKVTQKSARDTIKELTEKCKKIAQST--DDVIVNKPHDSDEE-EEERPFYNHPFKLSEPK
PIFFNLNVSTTTPIVKPAPKNQVTLTKEFPVSSGSQHRKKEADSVYGEWVPVEK-NKEES
--KDDVFPKP-ALAQVDISTAMNERTMAQKRLTENTFDLEAM-CLLNRAQEQIDAWAQSN
SLPGQFTGSTGAQILSSDELSNSGPQAWIRKDQFLRAAPV
PF01585 // ENSGT00730000111141_0_Domain_1 (2621, 2664)
TGGMGAQLMRKMGWREGEGL
GKNKEGTVEPILVDFKTDRKGLVENSGT00730000111141_0_Linker_1_2 (2664, 2688)
AVGEKTQKRSGHFS-VMKDLSGKHPF00035 // ENSGT00730000111141_0_Domain_2 (2688, 2755)
PVSALMEICNKRR
WSPPVFVLVDDSGPDHRKHFIFKVMVNGHEYRPSFPSLNK-KHAKATAATAALQAMGLVP
KESMPNATIFRSASHR
ENSFALP00000002461 (view gene)
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
------------------------------------------------------------
----------------------------------VVSKF---------------------
------------------------------------------------------------
--------REIQEEQL---SAKVE--GQPNGD-TIPAEQAGPSEASGA-AGSCQS-EQIV
--QKIEEVLSGALGTELQCNPDKAQLLEIAKANAAAMCAKSGVPLPPSLMPLLSQ-KK-D
DKANQKSSRDTLKELTEKCKKIAQST--DDVIVNKPHVSDEE-EEERPFYNHPFKLSEPK
PIFFNLSTPSIKPAPPPQPKNQVSLSKEFPVSSGSQHRKKEADSVYGEWVPVDKNGKDDG
--KDDVFPKP-AIECVDITTAMNDRAVAQKRLNENSFDLEAM-CMLNRAQEQIDAWAQSN
SIPGQFTGSTGAQILSSDELTNSGPQAWIRKDQFLRAAPV
PF01585 // ENSGT00730000111141_0_Domain_1 (2621, 2664)
TGGMGAQLMRKMGWREGEGL
GKNKEGSVEPIMVDFKTDRKGLVENSGT00730000111141_0_Linker_1_2 (2664, 2688)
AVGEKAQKRSGHYV-WLCVCAGKHPF00035 // ENSGT00730000111141_0_Domain_2 (2688, 2755)
PVSALMEICNKRR
WTPPEFVLVDDSGPDHRKHFIFKVRVNGNEYRPTFASLNK-KHAKATAATAALQAMGLVP
KESMVNTTMFRSASHR
{"ENSHGLP00000014574": [["Domain_0", 2174, 2419, "PF17069"], ["Linker_0_1", 2419, 2621], ["Domain_1", 2621, 2664, "PF01585"], ["Linker_1_2", 2664, 2687], ["Domain_2", 2687, 2756, "PF14709"]], "ENSGALP00000053471": [["Domain_0", 2115, 2413, "PF17069"], ["Linker_0_1", 2413, 2621], ["Domain_1", 2621, 2664, "PF01585"], ["Linker_1_2", 2664, 2688], ["Domain_2", 2688, 2755, "PF00035"]], "ENSMGAP00000015381": [["Domain_1", 2621, 2665, "PF01585"], ["Linker_1_2", 2665, 2688], ["Domain_2", 2688, 2756, "PF00035"]], "ENSFDAP00000011600": [["Domain_0", 2172, 2420, "PF17069"], ["Linker_0_1", 2420, 2621], ["Domain_1", 2621, 2664, "PF01585"], ["Linker_1_2", 2664, 2687], ["Domain_2", 2687, 2756, "PF14709"]], "ENSSARP00000011196": [["Domain_1", 2621, 2665, "PF01585"], ["Linker_1_2", 2665, 2688], ["Domain_2", 2688, 2724, "PF00035"]], "ENSCJAP00000054117": [["Domain_0", 2177, 2419, "PF17069"], ["Linker_0_1", 2419, 2621], ["Domain_1", 2621, 2664, "PF01585"], ["Linker_1_2", 2664, 2687], ["Domain_2", 2687, 2756, "PF14709"]], "ENSEEUP00000002160": [["Domain_1", 2621, 2665, "PF01585"], ["Linker_1_2", 2665, 2688], ["Domain_2", 2688, 2724, "PF00035"]], "ENSMICP00000030960": [["Domain_0", 2181, 2420, "PF17069"], ["Linker_0_1", 2420, 2621], ["Domain_1", 2621, 2664, "PF01585"], ["Linker_1_2", 2664, 2687], ["Domain_2", 2687, 2732, "PF14709"]], "ENSMPUP00000001279": [["Domain_1", 2621, 2664, "PF01585"], ["Linker_1_2", 2664, 2688], ["Domain_2", 2688, 2755, "PF00035"]], "ENSPCOP00000008444": [["Domain_0", 2187, 2420, "PF17069"], ["Linker_0_1", 2420, 2621], ["Domain_1", 2621, 2664, "PF01585"], ["Linker_1_2", 2664, 2687], ["Domain_2", 2687, 2732, "PF14709"]], "ENSMNEP00000015129": [["Domain_0", 2187, 2419, "PF17069"], ["Linker_0_1", 2419, 2621], ["Domain_1", 2621, 2664, "PF01585"], ["Linker_1_2", 2664, 2687], ["Domain_2", 2687, 2732, "PF14709"]], "ENSBTAP00000011175": [["Domain_1", 2621, 2664, "PF01585"], ["Linker_1_2", 2664, 2688], ["Domain_2", 2688, 2755, "PF00035"]], "ENSOPRP00000004675": [["Domain_1", 2621, 2665, "PF01585"], ["Linker_1_2", 2665, 2688], ["Domain_2", 2688, 2724, "PF00035"]], "ENSPTRP00000086465": [["Domain_1", 2628, 2664, "PF01585"], ["Linker_1_2", 2664, 2687], ["Domain_2", 2687, 2732, "PF14709"]], "ENSHGLP00100020224": [["Domain_1", 2621, 2664, "PF01585"], ["Linker_1_2", 2664, 2687], ["Domain_2", 2687, 2756, "PF14709"]], "ENSCAFP00000040685": [["Domain_1", 2621, 2664, "PF01585"], ["Linker_1_2", 2664, 2686], ["Domain_2", 2686, 2732, "PF14709"]], "ENSOARP00000013974": [["Domain_1", 2621, 2664, "PF01585"], ["Linker_1_2", 2664, 2688], ["Domain_2", 2688, 2755, "PF00035"]], "ENSSTOP00000016835": [["Domain_0", 2170, 2419, "PF17069"], ["Linker_0_1", 2419, 2621], ["Domain_1", 2621, 2664, "PF01585"], ["Linker_1_2", 2664, 2687], ["Domain_2", 2687, 2756, "PF14709"]], "ENSCAPP00000017326": [["Domain_1", 2628, 2661, "PF01585"]], "ENSCAFP00000013423": [["Domain_1", 2621, 2664, "PF01585"], ["Linker_1_2", 2664, 2686], ["Domain_2", 2686, 2732, "PF14709"]], "ENSLAFP00000013694": [["Domain_1", 2621, 2665, "PF01585"], ["Linker_1_2", 2665, 2688], ["Domain_2", 2688, 2724, "PF00035"]], "ENSCLAP00000017154": [["Domain_1", 2621, 2664, "PF01585"], ["Linker_1_2", 2664, 2687], ["Domain_2", 2687, 2756, "PF14709"]], "ENSODEP00000004818": [["Domain_1", 2621, 2664, "PF01585"], ["Linker_1_2", 2664, 2687], ["Domain_2", 2687, 2732, "PF14709"]], "ENSCCAP00000024911": [["Domain_0", 2089, 2419, "PF17069"], ["Linker_0_1", 2419, 2621], ["Domain_1", 2621, 2664, "PF01585"], ["Linker_1_2", 2664, 2687], ["Domain_2", 2687, 2732, "PF14709"]], "ENSSBOP00000007974": [["Domain_0", 2171, 2419, "PF17069"], ["Linker_0_1", 2419, 2621], ["Domain_1", 2621, 2664, "PF01585"], ["Linker_1_2", 2664, 2687], ["Domain_2", 2687, 2732, "PF14709"]], "ENSXETP00000030802": [["Domain_1", 2621, 2665, "PF01585"], ["Linker_1_2", 2665, 2688], ["Domain_2", 2688, 2756, "PF00035"]], "MGP_SPRETEiJ_P0044192": [["Domain_0", 2187, 2419, "PF17069"], ["Linker_0_1", 2419, 2621], ["Domain_1", 2621, 2664, "PF01585"], ["Linker_1_2", 2664, 2687], ["Domain_2", 2687, 2756, "PF14709"]], "ENSETEP00000015781": [["Domain_1", 2621, 2660, "PF01585"]], "ENSNLEP00000004882": [["Domain_0", 2187, 2419, "PF17069"], ["Linker_0_1", 2419, 2621], ["Domain_1", 2621, 2664, "PF01585"], ["Linker_1_2", 2664, 2687], ["Domain_2", 2687, 2732, "PF14709"]], "ENSPTRP00000053981": [["Domain_0", 2187, 2419, "PF17069"], ["Linker_0_1", 2419, 2621], ["Domain_1", 2621, 2664, "PF01585"], ["Linker_1_2", 2664, 2687], ["Domain_2", 2687, 2732, "PF14709"]], "ENSCATP00000011175": [["Domain_0", 2187, 2419, "PF17069"], ["Linker_0_1", 2419, 2621], ["Domain_1", 2621, 2664, "PF01585"], ["Linker_1_2", 2664, 2687], ["Domain_2", 2687, 2732, "PF14709"]], "ENSPTRP00000072700": [["Domain_1", 2621, 2664, "PF01585"], ["Linker_1_2", 2664, 2687], ["Domain_2", 2687, 2732, "PF14709"]], "ENSMODP00000037779": [["Domain_1", 2621, 2664, "PF01585"]], "ENSPPYP00000012703": [["Domain_1", 2621, 2664, "PF01585"], ["Linker_1_2", 2664, 2688], ["Domain_2", 2688, 2755, "PF00035"]], "ENSPCAP00000009555": [["Domain_1", 2621, 2660, "PF01585"]], "ENSMLEP00000021708": [["Domain_0", 2165, 2419, "PF17069"], ["Linker_0_1", 2419, 2621], ["Domain_1", 2621, 2664, "PF01585"], ["Linker_1_2", 2664, 2687], ["Domain_2", 2687, 2732, "PF14709"]], "ENSGGOP00000010230": [["Domain_0", 2177, 2419, "PF17069"], ["Linker_0_1", 2419, 2621], ["Domain_1", 2621, 2664, "PF01585"], ["Linker_1_2", 2664, 2687], ["Domain_2", 2687, 2732, "PF14709"]], "ENSFCAP00000027852": [["Domain_1", 2621, 2664, "PF01585"], ["Linker_1_2", 2664, 2687], ["Domain_2", 2687, 2732, "PF14709"]], "ENSCSAP00000006065": [["Domain_1", 2621, 2664, "PF01585"], ["Linker_1_2", 2664, 2688], ["Domain_2", 2688, 2755, "PF00035"]], "ENSPEMP00000028357": [["Domain_1", 2621, 2664, "PF01585"], ["Linker_1_2", 2664, 2687], ["Domain_2", 2687, 2732, "PF14709"]], "ENSRNOP00000002769": [["Domain_0", 2187, 2419, "PF17069"], ["Linker_0_1", 2419, 2621], ["Domain_1", 2621, 2664, "PF01585"], ["Linker_1_2", 2664, 2687], ["Domain_2", 2687, 2732, "PF14709"]], "MGP_CAROLIEiJ_P0042291": [["Domain_0", 2178, 2419, "PF17069"], ["Linker_0_1", 2419, 2621], ["Domain_1", 2621, 2664, "PF01585"], ["Linker_1_2", 2664, 2687], ["Domain_2", 2687, 2756, "PF14709"]], "ENSOGAP00000016216": [["Domain_1", 2621, 2660, "PF01585"]], "ENSCGRP00001015682": [["Domain_0", 2187, 2419, "PF17069"], ["Linker_0_1", 2419, 2621], ["Domain_1", 2621, 2664, "PF01585"], ["Linker_1_2", 2664, 2687], ["Domain_2", 2687, 2756, "PF14709"]], "ENSDNOP00000022750": [["Domain_1", 2621, 2664, "PF01585"], ["Linker_1_2", 2664, 2688], ["Domain_2", 2688, 2755, "PF00035"]], "ENSCHIP00000025768": [["Domain_0", 2187, 2419, "PF17069"], ["Linker_0_1", 2419, 2621], ["Domain_1", 2621, 2664, "PF01585"], ["Linker_1_2", 2664, 2687], ["Domain_2", 2687, 2732, "PF14709"]], "ENSTGUP00000000498": [["Domain_1", 2621, 2664, "PF01585"], ["Linker_1_2", 2664, 2688], ["Domain_2", 2688, 2755, "PF00035"]], "ENSANAP00000035842": [["Domain_0", 2177, 2419, "PF17069"], ["Linker_0_1", 2419, 2621], ["Domain_1", 2621, 2664, "PF01585"], ["Linker_1_2", 2664, 2687], ["Domain_2", 2687, 2732, "PF14709"]], "ENSODEP00000011614": [["Domain_1", 2625, 2651, "PF01585"], ["Linker_1_2", 2651, 2687], ["Domain_2", 2687, 2731, "PF14709"]], "ENSPVAP00000010544": [["Domain_1", 2621, 2665, "PF01585"], ["Linker_1_2", 2665, 2688], ["Domain_2", 2688, 2724, "PF00035"]], "ENSTTRP00000004433": [["Domain_1", 2621, 2665, "PF01585"]], "ENSMOCP00000012900": [["Domain_1", 2621, 2664, "PF01585"], ["Linker_1_2", 2664, 2687], ["Domain_2", 2687, 2732, "PF14709"]], "ENSCANP00000007967": [["Domain_0", 2170, 2419, "PF17069"], ["Linker_0_1", 2419, 2621], ["Domain_1", 2621, 2664, "PF01585"], ["Linker_1_2", 2664, 2687], ["Domain_2", 2687, 2732, "PF14709"]], "ENSODEP00000005985": [["Domain_1", 2625, 2664, "PF01585"], ["Linker_1_2", 2664, 2687], ["Domain_2", 2687, 2739, "PF14709"]], "ENSPPAP00000025487": [["Domain_2", 2687, 2732, "PF14709"]], "ENSRROP00000037529": [["Domain_0", 2178, 2419, "PF17069"], ["Linker_0_1", 2419, 2621], ["Domain_1", 2621, 2664, "PF01585"], ["Linker_1_2", 2664, 2687], ["Domain_2", 2687, 2732, "PF14709"]], "ENSCPOP00000032872": [["Domain_1", 2621, 2664, "PF01585"], ["Linker_1_2", 2664, 2687], ["Domain_2", 2687, 2756, "PF14709"]], "MGP_PahariEiJ_P0029650": [["Domain_0", 2187, 2419, "PF17069"], ["Linker_0_1", 2419, 2621], ["Domain_1", 2621, 2664, "PF01585"], ["Linker_1_2", 2664, 2687], ["Domain_2", 2687, 2756, "PF14709"]], "ENSECAP00000015914": [["Domain_1", 2621, 2664, "PF01585"]], "ENSMAUP00000002486": [["Domain_1", 2621, 2664, "PF01585"], ["Linker_1_2", 2664, 2687], ["Domain_2", 2687, 2732, "PF14709"]], "ENSJJAP00000009701": [["Domain_1", 2621, 2664, "PF01585"], ["Linker_1_2", 2664, 2687], ["Domain_2", 2687, 2732, "PF14709"]], "ENSCGRP00000020986": [["Domain_0", 2174, 2419, "PF17069"], ["Linker_0_1", 2419, 2621], ["Domain_1", 2621, 2664, "PF01585"], ["Linker_1_2", 2664, 2687], ["Domain_2", 2687, 2756, "PF14709"]], "ENSMMUP00000007291": [["Domain_0", 2178, 2420, "PF17069"], ["Linker_0_1", 2420, 2621], ["Domain_1", 2621, 2664, "PF01585"], ["Linker_1_2", 2664, 2687], ["Domain_2", 2687, 2756, "PF14709"]], "ENSMLUP00000002235": [["Domain_1", 2621, 2665, "PF01585"], ["Linker_1_2", 2665, 2688], ["Domain_2", 2688, 2724, "PF00035"]], "ENSMFAP00000036158": [["Domain_0", 2187, 2419, "PF17069"], ["Linker_0_1", 2419, 2621], ["Domain_1", 2621, 2664, "PF01585"], ["Linker_1_2", 2664, 2687], ["Domain_2", 2687, 2732, "PF14709"]], "ENSTSYP00000007689": [["Domain_1", 2621, 2664, "PF01585"], ["Linker_1_2", 2664, 2687], ["Domain_2", 2687, 2732, "PF14709"]], "ENSOCUP00000003354": [["Domain_1", 2621, 2664, "PF01585"]], "ENSSSCP00000056796": [["Domain_0", 2187, 2419, "PF17069"], ["Linker_0_1", 2419, 2621], ["Domain_1", 2621, 2664, "PF01585"], ["Linker_1_2", 2664, 2687], ["Domain_2", 2687, 2732, "PF14709"]], "ENSNGAP00000000072": [["Domain_1", 2621, 2664, "PF01585"], ["Linker_1_2", 2664, 2687], ["Domain_2", 2687, 2732, "PF14709"]], "ENSVPAP00000008517": [["Domain_1", 2621, 2665, "PF01585"]], "ENSMUSP00000109671": [["Domain_0", 2171, 2419, "PF17069"], ["Linker_0_1", 2419, 2621], ["Domain_1", 2621, 2664, "PF01585"], ["Linker_1_2", 2664, 2687], ["Domain_2", 2687, 2732, "PF14709"]], "ENSRBIP00000041914": [["Domain_1", 2621, 2664, "PF01585"], ["Linker_1_2", 2664, 2687], ["Domain_2", 2687, 2732, "PF14709"]], "ENSAMEP00000015669": [["Domain_1", 2621, 2665, "PF01585"], ["Linker_1_2", 2665, 2688], ["Domain_2", 2688, 2724, "PF00035"]], "ENSTBEP00000002791": [["Domain_2", 2688, 2724, "PF00035"]], "ENSP00000348984": [["Domain_0", 2187, 2419, "PF17069"], ["Linker_0_1", 2419, 2621], ["Domain_1", 2621, 2664, "PF01585"], ["Linker_1_2", 2664, 2687], ["Domain_2", 2687, 2730, "PF14709"]], "ENSPANP00000013073": [["Domain_0", 2187, 2419, "PF17069"], ["Linker_0_1", 2419, 2621], ["Domain_1", 2621, 2664, "PF01585"], ["Linker_1_2", 2664, 2687], ["Domain_2", 2687, 2732, "PF14709"]], "ENSDORP00000022865": [["Domain_1", 2621, 2664, "PF01585"], ["Linker_1_2", 2664, 2687], ["Domain_2", 2687, 2732, "PF14709"]], "ENSPPAP00000021001": [["Domain_0", 2173, 2419, "PF17069"], ["Linker_0_1", 2419, 2621], ["Domain_1", 2621, 2664, "PF01585"], ["Linker_1_2", 2664, 2687], ["Domain_2", 2687, 2732, "PF14709"]], "ENSPSIP00000008930": [["Domain_1", 2621, 2664, "PF01585"], ["Linker_1_2", 2664, 2688], ["Domain_2", 2688, 2755, "PF00035"]], "ENSFALP00000002461": [["Domain_1", 2621, 2664, "PF01585"], ["Linker_1_2", 2664, 2688], ["Domain_2", 2688, 2755, "PF00035"]]}

